Object masks from prompts with SAM, OpenVINO, and FiftyOne¶
Note: This notebook is adapted from the SAM OpenVINO notebook.
Segmentation - identifying which image pixels belong to an object - is a core task in computer vision and is used in a broad array of applications, from analyzing scientific imagery to editing photos. But creating an accurate segmentation model for specific tasks typically requires highly specialized work by technical experts with access to AI training infrastructure and large volumes of carefully annotated in-domain data. Reducing the need for task-specific modeling expertise, training compute, and custom data annotation for image segmentation is the main goal of the Segment Anything project.
The Segment Anything Model (SAM) predicts object masks given prompts that indicate the desired object. SAM has learned a general notion of what objects are, and it can generate masks for any object in any image or any video, even including objects and image types that it had not encountered during training. SAM is general enough to cover a broad set of use cases and can be used out of the box on new image “domains” (e.g. underwater photos, MRI or cell microscopy) without requiring additional training (a capability often referred to as zero-shot transfer). This notebook shows an example of how to convert and use Segment Anything Model in OpenVINO format, allowing it to run on a variety of platforms that support an OpenVINO.
Background¶
Previously, to solve any kind of segmentation problem, there were two classes of approaches. The first, interactive segmentation, allowed for segmenting any class of object but required a person to guide the method by iterative refining a mask. The second, automatic segmentation, allowed for segmentation of specific object categories defined ahead of time (e.g., cats or chairs) but required substantial amounts of manually annotated objects to train (e.g., thousands or even tens of thousands of examples of segmented cats), along with the compute resources and technical expertise to train the segmentation model. Neither approach provided a general, fully automatic approach to segmentation.
Segment Anything Model is a generalization of these two classes of approaches. It is a single model that can easily perform both interactive segmentation and automatic segmentation. The Segment Anything Model (SAM) produces high quality object masks from input prompts such as points or boxes, and it can be used to generate masks for all objects in an image. It has been trained on a dataset of 11 million images and 1.1 billion masks, and has strong zero-shot performance on a variety of segmentation tasks. The model consists of 3 parts:
- Image Encoder - Vision Transformer model (VIT) pretrained using Masked Auto Encoders approach (MAE) for encoding image to embedding space. The image encoder runs once per image and can be applied prior to prompting the model.
- Prompt Encoder - Encoder for segmentation condition. As a condition can be used:
- points - set of points related to object which should be segmented. Prompt encoder converts points to embedding using positional encoding.
- boxes - bounding box where object for segmentation is located. Similar to points, coordinates of bounding box encoded via positional encoding.
- segmentation mask - provided by user segmentation mask is embedded using convolutions and summed element-wise with the image embedding.
- text - encoded by CLIP model text representation
- Mask Decoder - The mask decoder efficiently maps the image embedding, prompt embeddings, and an output token to a mask.
The diagram below demonstrates the process of mask generation using SAM:
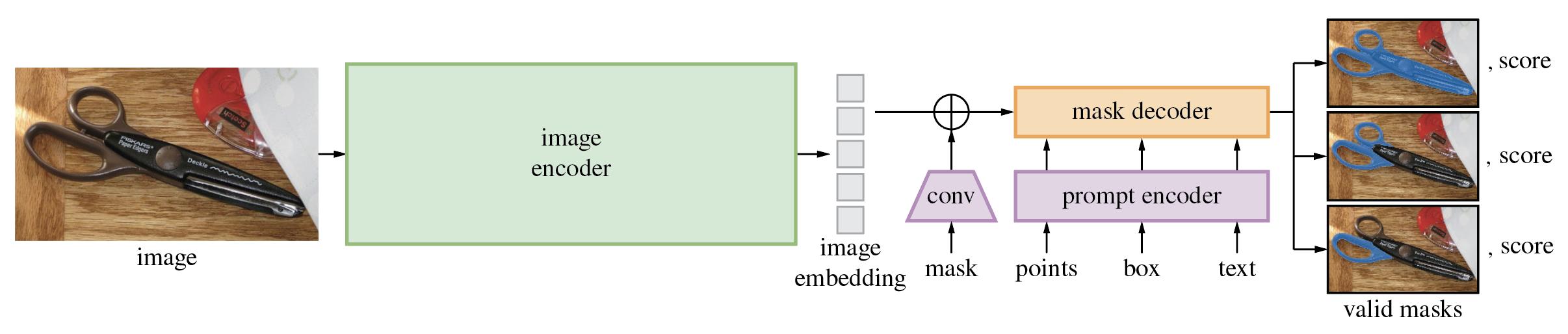
The model first converts the image into an image embedding that allows high quality masks to be efficiently produced from a prompt. The model returns multiple masks which fit to the provided prompt and its score. The provided masks can be overlapped areas as it shown on diagram, it is useful for complicated cases when prompt can be interpreted in different manner, e.g. segment whole object or only its specific part or when provided point at the intersection of multiple objects. The model’s promptable interface allows it to be used in flexible ways that make a wide range of segmentation tasks possible simply by engineering the right prompt for the model (clicks, boxes, text, and so on).
More details about approach can be found in the paper, original repo and Meta AI blog post
Prerequisites¶
In addition to OpenVINO and Meta AI's Segment Anything library, we will use the open source library FiftyOne for visualizing and evaluating our Segment Anything Model predictions.
!pip install -q "segment_anything" "fiftyone" "openvino"
Convert model to OpenVINO Intermediate Representation¶
Download model checkpoint and create PyTorch model¶
There are several Segment Anything Model checkpoints available for downloading
In this tutorial we will use model based on vit_b, but the demonstrated approach is very general and applicable to other SAM models.
Set the model url, path for saving checkpoint and model type below to a SAM model checkpoint, then load the model using sam_model_registry.
import sys
sys.path.append("../utils")
from notebook_utils import download_file
checkpoint = "sam_vit_b_01ec64.pth"
model_url = "https://dl.fbaipublicfiles.com/segment_anything/sam_vit_b_01ec64.pth"
model_type = "vit_b"
download_file(model_url)
from segment_anything import sam_model_registry
sam = sam_model_registry[model_type](checkpoint=checkpoint)
As we already discussed, Image Encoder part can be used once per image, then changing prompt, prompt encoder and mask decoder can be run multiple times to retrieve different objects from the same image. Taking into account this fact, we split model on 2 independent parts: image_encoder and mask_predictor (combination of Prompt Encoder and Mask Decoder).
Image Encoder¶
Image Encoder input is tensor with shape 1x3x1024x1024 in NCHW format, contains image for segmentation.
Image Encoder output is image embeddings, tensor with shape 1x256x64x64
import warnings
from pathlib import Path
import torch
from openvino.tools import mo
from openvino.runtime import serialize, Core
core = Core()
ov_encoder_path = Path("sam_image_encoder.xml")
if not ov_encoder_path.exists():
onnx_encoder_path = ov_encoder_path.with_suffix(".onnx")
if not onnx_encoder_path.exists():
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=torch.jit.TracerWarning)
warnings.filterwarnings("ignore", category=UserWarning)
torch.onnx.export(
sam.image_encoder, torch.zeros(1, 3, 1024, 1024), onnx_encoder_path
)
ov_encoder_model = mo.convert_model(onnx_encoder_path, compress_to_fp16=True)
serialize(ov_encoder_model, str(ov_encoder_path))
else:
ov_encoder_model = core.read_model(ov_encoder_path)
ov_encoder = core.compile_model(ov_encoder_model)
Mask predictor¶
This notebook expects the model was exported with the parameter return_single_mask=True. It means that model will only return the best mask, instead of returning multiple masks. For high resolution images this can improve runtime when upscaling masks is expensive.
Combined prompt encoder and mask decoder model has following list of inputs:
image_embeddings: The image embedding fromimage_encoder. Has a batch index of length 1.point_coords: Coordinates of sparse input prompts, corresponding to both point inputs and box inputs. Boxes are encoded using two points, one for the top-left corner and one for the bottom-right corner. Coordinates must already be transformed to long-side 1024. Has a batch index of length 1.point_labels: Labels for the sparse input prompts. 0 is a negative input point, 1 is a positive input point, 2 is a top-left box corner, 3 is a bottom-right box corner, and -1 is a padding point. *If there is no box input, a single padding point with label -1 and coordinates (0.0, 0.0) should be concatenated.
Model outputs:
masks- predicted masks resized to original image size, to obtain a binary mask, should be compared withthreshold(usually equal 0.0).iou_predictions- intersection over union predictionslow_res_masks- predicted masks before postprocessing, can be used as mask input for model.
from typing import Tuple
class SamONNXModel(torch.nn.Module):
def __init__(
self,
model,
return_single_mask: bool,
use_stability_score: bool = False,
return_extra_metrics: bool = False,
) -> None:
super().__init__()
self.mask_decoder = model.mask_decoder
self.model = model
self.img_size = model.image_encoder.img_size
self.return_single_mask = return_single_mask
self.use_stability_score = use_stability_score
self.stability_score_offset = 1.0
self.return_extra_metrics = return_extra_metrics
def _embed_points(
self, point_coords: torch.Tensor, point_labels: torch.Tensor
) -> torch.Tensor:
point_coords = point_coords + 0.5
point_coords = point_coords / self.img_size
point_embedding = self.model.prompt_encoder.pe_layer._pe_encoding(point_coords)
point_labels = point_labels.unsqueeze(-1).expand_as(point_embedding)
point_embedding = point_embedding * (point_labels != -1)
point_embedding = (
point_embedding
+ self.model.prompt_encoder.not_a_point_embed.weight * (point_labels == -1)
)
for i in range(self.model.prompt_encoder.num_point_embeddings):
point_embedding = (
point_embedding
+ self.model.prompt_encoder.point_embeddings[i].weight
* (point_labels == i)
)
return point_embedding
def t_embed_masks(self, input_mask: torch.Tensor) -> torch.Tensor:
mask_embedding = self.model.prompt_encoder.mask_downscaling(input_mask)
return mask_embedding
def mask_postprocessing(self, masks: torch.Tensor) -> torch.Tensor:
masks = torch.nn.functional.interpolate(
masks,
size=(self.img_size, self.img_size),
mode="bilinear",
align_corners=False,
)
return masks
def select_masks(
self, masks: torch.Tensor, iou_preds: torch.Tensor, num_points: int
) -> Tuple[torch.Tensor, torch.Tensor]:
# Determine if we should return the multiclick mask or not from the number of points.
# The reweighting is used to avoid control flow.
score_reweight = torch.tensor(
[[1000] + [0] * (self.model.mask_decoder.num_mask_tokens - 1)]
).to(iou_preds.device)
score = iou_preds + (num_points - 2.5) * score_reweight
best_idx = torch.argmax(score, dim=1)
masks = masks[torch.arange(masks.shape[0]), best_idx, :, :].unsqueeze(1)
iou_preds = iou_preds[torch.arange(masks.shape[0]), best_idx].unsqueeze(1)
return masks, iou_preds
@torch.no_grad()
def forward(
self,
image_embeddings: torch.Tensor,
point_coords: torch.Tensor,
point_labels: torch.Tensor,
mask_input: torch.Tensor = None,
):
sparse_embedding = self._embed_points(point_coords, point_labels)
if mask_input is None:
dense_embedding = self.model.prompt_encoder.no_mask_embed.weight.reshape(
1, -1, 1, 1
).expand(point_coords.shape[0], -1, image_embeddings.shape[0], 64)
else:
dense_embedding = self._embed_masks(mask_input)
masks, scores = self.model.mask_decoder.predict_masks(
image_embeddings=image_embeddings,
image_pe=self.model.prompt_encoder.get_dense_pe(),
sparse_prompt_embeddings=sparse_embedding,
dense_prompt_embeddings=dense_embedding,
)
if self.use_stability_score:
scores = calculate_stability_score(
masks, self.model.mask_threshold, self.stability_score_offset
)
if self.return_single_mask:
masks, scores = self.select_masks(masks, scores, point_coords.shape[1])
upscaled_masks = self.mask_postprocessing(masks)
if self.return_extra_metrics:
stability_scores = calculate_stability_score(
upscaled_masks, self.model.mask_threshold, self.stability_score_offset
)
areas = (upscaled_masks > self.model.mask_threshold).sum(-1).sum(-1)
return upscaled_masks, scores, stability_scores, areas, masks
return upscaled_masks, scores
ov_model_path = Path("sam_mask_predictor.xml")
if not ov_model_path.exists():
onnx_model_path = ov_model_path.with_suffix(".onnx")
if not onnx_model_path.exists():
onnx_model = SamONNXModel(sam, return_single_mask=True)
dynamic_axes = {
"point_coords": {0: "batch_size", 1: "num_points"},
"point_labels": {0: "batch_size", 1: "num_points"},
}
embed_dim = sam.prompt_encoder.embed_dim
embed_size = sam.prompt_encoder.image_embedding_size
dummy_inputs = {
"image_embeddings": torch.randn(
1, embed_dim, *embed_size, dtype=torch.float
),
"point_coords": torch.randint(
low=0, high=1024, size=(1, 5, 2), dtype=torch.float
),
"point_labels": torch.randint(
low=0, high=4, size=(1, 5), dtype=torch.float
),
}
output_names = ["masks", "iou_predictions"]
with warnings.catch_warnings():
warnings.filterwarnings("ignore", category=torch.jit.TracerWarning)
warnings.filterwarnings("ignore", category=UserWarning)
torch.onnx.export(
onnx_model,
tuple(dummy_inputs.values()),
onnx_model_path,
input_names=list(dummy_inputs.keys()),
output_names=output_names,
dynamic_axes=dynamic_axes,
)
ov_model = mo.convert_model(onnx_model_path, compress_to_fp16=True)
serialize(ov_model, str(ov_model_path))
else:
ov_model = core.read_model(ov_model_path)
ov_predictor = core.compile_model(ov_model)
Preparing an image for OpenVINO SAM segmentation¶
Before we can use the OpenVINO model, we need to convert the input data to the correct format. Using an example image, this section details
- Downloading the image
- Loading the image into FiftyOne for visualization
- Preprocessing for OpenVINO
- Image encoding
Download an image¶
To start, we'll use the same image as Facebook Research uses in their Segment Anything Model demo notebooks:
import numpy as np
import cv2
import fiftyone as fo
download_file(
"https://raw.githubusercontent.com/facebookresearch/segment-anything/main/notebooks/images/truck.jpg"
)
filepath = "truck.jpg"
image = cv2.imread(filepath)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
Load the image into FiftyOne¶
Create a FiftyOne Dataset containing this image:
dataset = fo.Dataset(name="openvino_sam", persistent=True, overwrite=True)
dataset.add_sample(fo.Sample(filepath=filepath))
dataset.compute_metadata()
sample = dataset.first()
Computing metadata... 100% |█████████████████████| 1/1 [788.9ms elapsed, 0s remaining, 1.3 samples/s]
Above, we are using overwrite=True to overwrite the existing dataset, which will allow you to run this cell multiple times without throwing an error. Alternatively, if you don't want to overwrite any datasets, you can create a new dataset without a name via dataset = fo.Dataset(persistent=True).
Once we've created the dataset, we can visualize it in the FiftyOne App:
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

We will also define a few utility functions to help us convert between FiftyOne and OpenVINO SAM formats. abs_to_rel() and rel_to_abs() convert between absolute and relative pixel coordinates; convert_sam_to_fo_box() and convert_fo_to_sam_box() converts a box output by OpenVINO SAM to a format that can be used by FiftyOne, and vice versa; and convert_label() converts the integer labels output by OpenVINO SAM to strings that can be used by FiftyOne:
def abs_to_rel(abs_coords, sample):
rel_coords = np.copy(abs_coords).astype("float")
rel_coords[:, 0] /= sample.metadata.width
rel_coords[:, 1] /= sample.metadata.height
return rel_coords
def rel_to_abs(rel_coords, sample):
abs_coords = np.copy(rel_coords)
abs_coords[:, 0] *= sample.metadata.width
abs_coords[:, 1] *= sample.metadata.height
return abs_coords.astype("int")
def convert_sam_to_fo_box(box, sample):
## convert bounding box from SAM format with absolute coordinates
## [<top-left-x>, <top-left-y>, <bottom-right-x>, <bottom-right-y>]
## to FiftyOne bounding box format with relative coordinates
## [<top-left-x>, <top-left-y>, <width>, <height>]
w, h = sample.metadata.width, sample.metadata.height
fo_box = np.copy(box).astype("float")
fo_box[0] /= w
fo_box[2] /= w
fo_box[1] /= h
fo_box[3] /= h
fo_box[2] -= fo_box[0]
fo_box[3] -= fo_box[1]
return fo_box
def convert_fo_to_sam_box(box, sample):
## convert bounding box format from FiftyOne with relative coordinates
## [<top-left-x>, <top-left-y>, <width>, <height>]
## to SAM absolute coordinates
## [<top-left-x>, <top-left-y>, <bottom-right-x>, <bottom-right-y>]
w, h = sample.metadata.width, sample.metadata.height
sam_box = np.copy(box)
sam_box[0] *= w
sam_box[2] *= w
sam_box[1] *= h
sam_box[3] *= h
sam_box[2] += sam_box[0]
sam_box[3] += sam_box[1]
return sam_box.astype("int")
def convert_label(input_label):
return str(int(input_label))
Preprocessing utilities¶
To prepare iinput for Image Encoder we should:
- Convert BGR image to RGB
- Resize image saving aspect ratio where longest size equal to Image Encoder input size - 1024.
- Normalize image subtract mean values (123.675, 116.28, 103.53) and divide by std (58.395, 57.12, 57.375)
- transpose HWC data layout to CHW and add batch dimension.
- add zero padding to input tensor by height or width (depends on aspect ratio) according Image Encoder expected input shape.
These steps are applicable to all available models
from copy import deepcopy
from typing import Tuple
from torchvision.transforms.functional import resize, to_pil_image
class ResizeLongestSide:
"""
Resizes images to longest side 'target_length', as well as provides
methods for resizing coordinates and boxes. Provides methods for
transforming numpy arrays.
"""
def __init__(self, target_length: int) -> None:
self.target_length = target_length
def apply_image(self, image: np.ndarray) -> np.ndarray:
"""
Expects a numpy array with shape HxWxC in uint8 format.
"""
target_size = self.get_preprocess_shape(
image.shape[0], image.shape[1], self.target_length
)
return np.array(resize(to_pil_image(image), target_size))
def apply_coords(
self, coords: np.ndarray, original_size: Tuple[int, ...]
) -> np.ndarray:
"""
Expects a numpy array of length 2 in the final dimension. Requires the
original image size in (H, W) format.
"""
old_h, old_w = original_size
new_h, new_w = self.get_preprocess_shape(
original_size[0], original_size[1], self.target_length
)
coords = deepcopy(coords).astype(float)
coords[..., 0] = coords[..., 0] * (new_w / old_w)
coords[..., 1] = coords[..., 1] * (new_h / old_h)
return coords
def apply_boxes(
self, boxes: np.ndarray, original_size: Tuple[int, ...]
) -> np.ndarray:
"""
Expects a numpy array shape Bx4. Requires the original image size
in (H, W) format.
"""
boxes = self.apply_coords(boxes.reshape(-1, 2, 2), original_size)
return boxes.reshape(-1, 4)
@staticmethod
def get_preprocess_shape(
oldh: int, oldw: int, long_side_length: int
) -> Tuple[int, int]:
"""
Compute the output size given input size and target long side length.
"""
scale = long_side_length * 1.0 / max(oldh, oldw)
newh, neww = oldh * scale, oldw * scale
neww = int(neww + 0.5)
newh = int(newh + 0.5)
return (newh, neww)
resizer = ResizeLongestSide(1024)
def preprocess_image(image: np.ndarray):
resized_image = resizer.apply_image(image)
resized_image = (resized_image.astype(np.float32) - [123.675, 116.28, 103.53]) / [
58.395,
57.12,
57.375,
]
resized_image = np.expand_dims(
np.transpose(resized_image, (2, 0, 1)).astype(np.float32), 0
)
# Pad
h, w = resized_image.shape[-2:]
padh = 1024 - h
padw = 1024 - w
x = np.pad(resized_image, ((0, 0), (0, 0), (0, padh), (0, padw)))
return x
def postprocess_masks(masks: np.ndarray, orig_size):
size_before_pad = resizer.get_preprocess_shape(
orig_size[0], orig_size[1], masks.shape[-1]
)
masks = masks[..., : int(size_before_pad[0]), : int(size_before_pad[1])]
masks = torch.nn.functional.interpolate(
torch.from_numpy(masks), size=orig_size, mode="bilinear", align_corners=False
).numpy()
return masks
Image encoding¶
To start work with image, we should preprocess it and obtain image embeddings using ov_encoder. We will use the same image for all experiments, so it is possible to generate image embedding once and then reuse them.
preprocessed_image = preprocess_image(image)
encoding_results = ov_encoder(preprocessed_image)
image_embeddings = encoding_results[ov_encoder.output(0)]
Save the image embeddings on the sample:
sample["image_embeddings"] = image_embeddings
sample.save()
Next, we define a function which will take in the sample (with the image embeddings) and a prompt, and return the predicted mask:
def generate_mask(sample, point_coords, point_labels, box_coords=None, box_labels=None):
image = cv2.imread(sample.filepath)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image_embeddings = sample["image_embeddings"]
if box_coords is None:
box_coords = np.array([[0.0, 0.0]])
box_labels = np.array([-1])
else:
box_coords = box_coords.reshape(2, 2)
if point_coords is None or len(point_coords) == 0:
point_coords = np.array([[0.0, 0.0]])
point_labels = np.array([0])
coords = np.concatenate([point_coords, box_coords], axis=0)[None, :, :]
labels = np.concatenate([point_labels, box_labels], axis=0)[None, :].astype(
np.float32
)
coords = resizer.apply_coords(coords, image.shape[:2]).astype(np.float32)
inputs = {
"image_embeddings": image_embeddings,
"point_coords": coords,
"point_labels": labels,
}
results = ov_predictor(inputs)
masks = results[ov_predictor.output(0)]
masks = postprocess_masks(masks, image.shape[:-1])
masks = masks > 0.0
return masks[0, 0, :, :].astype(np.uint8)
This function adds a batch index, concatenates a padding point, and transforms it to input tensor coordinate system. It then packages the inputs to run in the mask predictor. Finally, it predicts a mask and thresholds it to get binary mask (0 - no object, 1 - object).
Run OpenVINO SAM with prompts¶
Now we are ready to run OpenVINO SAM with various prompts for mask generation.
Run OpenVINO SAM with single-point prompt¶
First, we select a single point. Starting from the point's location in pixels,
we convert it to relative coordinates and then define a Keypoint label in FiftyOne for this point:
input_point = np.array([[500, 375]])
input_label = np.array([1])
sample["window_point"] = fo.Keypoint(
label=convert_label(input_label[0]), points=abs_to_rel(input_point, sample)
)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

Hovering over the point in the FiftyOne App, we can see the name of the Keypoint label we created, "window_point", as well as its label: "1".
Now it is easy to generate a mask for this point:
window_mask = generate_mask(sample, input_point, input_label)
And then we can add the mask to the sample and visualize it in the FiftyOne App:
sample["window_mask"] = fo.Segmentation(mask=window_mask)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

Run OpenVINO SAM with multi-point prompt¶
Now let's provide additional points covering a larger object area.
input_point = np.array([[500, 375], [1125, 625], [575, 750], [1405, 575]])
input_label = np.array([1, 1, 1, 1])
To see what this prompt for model looks like on this image, we can represent these points with a FiftyOne Keypoints label:
input_point_fo = abs_to_rel(input_point, sample)
sample["car_points"] = fo.Keypoints(
keypoints=[
fo.Keypoint(label=convert_label(il), points=[tuple(ip)])
for il, ip in zip(input_label, input_point_fo)
]
)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

We can generate a mask as in the previous example:
car_mask = generate_mask(sample, input_point, input_label)
sample["car_mask"] = fo.Segmentation(mask=car_mask)
sample.save()
And then once again visualize the mask in the FiftyOne App:
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

Great! Looks like now, the predicted mask cover whole truck.
Run OpenVINO SAM with a box and negative point label¶
In this final prompting example, we define input prompt using bounding box and point inside it. The bounding box represented as set of points of its left upper corner and right lower corner. Label 0 for a point means that this point should be excluded from mask.
input_point = np.array([[575, 750]])
input_label = np.array([0])
input_box = np.array([425, 600, 700, 875])
box_labels = np.array([2, 3])
We will represent the point as a Keypoint label, and the box as a Detections label:
sample["tire_point"] = fo.Keypoint(
label=convert_label(input_label[0]), points=abs_to_rel(input_point, sample)
)
sample.save()
sample["tire_box"] = fo.Detection(
label="tire",
bounding_box=convert_sam_to_fo_box(input_box, sample),
)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

This time, we pass in box_coords and box_labels arguments to generate_mask(). There is no padding point since the input includes a box input.
tire_mask = generate_mask(
sample, input_point, input_label, box_coords=input_box, box_labels=box_labels
)
We can visualize the mask in the FiftyOne App with a Detection label that has a non-trivial mask field:
x0, y0, x1, y1 = input_box
mask_trimmed = np.array(tire_mask[y0 : y1 + 1, x0 : x1 + 1])
sample["tire_mask"] = fo.Detection(
label="tire",
bounding_box=convert_sam_to_fo_box(input_box, sample),
mask=mask_trimmed,
)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

Run OpenVINO SAM in automatic mask generation mode¶
Since SAM can efficiently process prompts, masks for the entire image can be generated by sampling a large number of prompts over an image.
The automatic_mask_generation() function implements this capability.
It works by sampling single-point input prompts in a grid over the image, from each of which SAM can predict multiple masks. Then, masks are filtered for quality and deduplicated using non-maximal suppression. Additional options allow for further improvement of mask quality and quantity, such as running prediction on multiple crops of the image or postprocessing masks to remove small disconnected regions and holes.
from segment_anything.utils.amg import (
MaskData,
generate_crop_boxes,
uncrop_boxes_xyxy,
uncrop_masks,
uncrop_points,
calculate_stability_score,
rle_to_mask,
batched_mask_to_box,
mask_to_rle_pytorch,
is_box_near_crop_edge,
batch_iterator,
remove_small_regions,
build_all_layer_point_grids,
box_xyxy_to_xywh,
area_from_rle,
)
from torchvision.ops.boxes import batched_nms, box_area
from typing import Tuple, List, Dict, Any
def process_batch(
image_embedding: np.ndarray,
points: np.ndarray,
im_size: Tuple[int, ...],
crop_box: List[int],
orig_size: Tuple[int, ...],
iou_thresh,
mask_threshold,
stability_score_offset,
stability_score_thresh,
) -> MaskData:
orig_h, orig_w = orig_size
# Run model on this batch
transformed_points = resizer.apply_coords(points, im_size)
in_points = transformed_points
in_labels = np.ones(in_points.shape[0], dtype=int)
inputs = {
"image_embeddings": image_embedding,
"point_coords": in_points[:, None, :],
"point_labels": in_labels[:, None],
}
res = ov_predictor(inputs)
masks = postprocess_masks(res[ov_predictor.output(0)], orig_size)
masks = torch.from_numpy(masks)
iou_preds = torch.from_numpy(res[ov_predictor.output(1)])
# Serialize predictions and store in MaskData
data = MaskData(
masks=masks.flatten(0, 1),
iou_preds=iou_preds.flatten(0, 1),
points=torch.as_tensor(points.repeat(masks.shape[1], axis=0)),
)
del masks
# Filter by predicted IoU
if iou_thresh > 0.0:
keep_mask = data["iou_preds"] > iou_thresh
data.filter(keep_mask)
# Calculate stability score
data["stability_score"] = calculate_stability_score(
data["masks"], mask_threshold, stability_score_offset
)
if stability_score_thresh > 0.0:
keep_mask = data["stability_score"] >= stability_score_thresh
data.filter(keep_mask)
# Threshold masks and calculate boxes
data["masks"] = data["masks"] > mask_threshold
data["boxes"] = batched_mask_to_box(data["masks"])
# Filter boxes that touch crop boundaries
keep_mask = ~is_box_near_crop_edge(data["boxes"], crop_box, [0, 0, orig_w, orig_h])
if not torch.all(keep_mask):
data.filter(keep_mask)
# Compress to RLE
data["masks"] = uncrop_masks(data["masks"], crop_box, orig_h, orig_w)
data["rles"] = mask_to_rle_pytorch(data["masks"])
del data["masks"]
return data
def process_crop(
image: np.ndarray,
point_grids,
crop_box: List[int],
crop_layer_idx: int,
orig_size: Tuple[int, ...],
box_nms_thresh: float = 0.7,
mask_threshold: float = 0.0,
points_per_batch: int = 64,
pred_iou_thresh: float = 0.88,
stability_score_thresh: float = 0.95,
stability_score_offset: float = 1.0,
) -> MaskData:
# Crop the image and calculate embeddings
x0, y0, x1, y1 = crop_box
cropped_im = image[y0:y1, x0:x1, :]
cropped_im_size = cropped_im.shape[:2]
preprocessed_cropped_im = preprocess_image(cropped_im)
crop_embeddings = ov_encoder(preprocessed_cropped_im)[ov_encoder.output(0)]
# Get points for this crop
points_scale = np.array(cropped_im_size)[None, ::-1]
points_for_image = point_grids[crop_layer_idx] * points_scale
# Generate masks for this crop in batches
data = MaskData()
for (points,) in batch_iterator(points_per_batch, points_for_image):
batch_data = process_batch(
crop_embeddings,
points,
cropped_im_size,
crop_box,
orig_size,
pred_iou_thresh,
mask_threshold,
stability_score_offset,
stability_score_thresh,
)
data.cat(batch_data)
del batch_data
# Remove duplicates within this crop.
keep_by_nms = batched_nms(
data["boxes"].float(),
data["iou_preds"],
torch.zeros(len(data["boxes"])), # categories
iou_threshold=box_nms_thresh,
)
data.filter(keep_by_nms)
# Return to the original image frame
data["boxes"] = uncrop_boxes_xyxy(data["boxes"], crop_box)
data["points"] = uncrop_points(data["points"], crop_box)
data["crop_boxes"] = torch.tensor([crop_box for _ in range(len(data["rles"]))])
return data
def generate_masks(
image: np.ndarray, point_grids, crop_n_layers, crop_overlap_ratio, crop_nms_thresh
) -> MaskData:
orig_size = image.shape[:2]
crop_boxes, layer_idxs = generate_crop_boxes(
orig_size, crop_n_layers, crop_overlap_ratio
)
# Iterate over image crops
data = MaskData()
for crop_box, layer_idx in zip(crop_boxes, layer_idxs):
crop_data = process_crop(image, point_grids, crop_box, layer_idx, orig_size)
data.cat(crop_data)
# Remove duplicate masks between crops
if len(crop_boxes) > 1:
# Prefer masks from smaller crops
scores = 1 / box_area(data["crop_boxes"])
scores = scores.to(data["boxes"].device)
keep_by_nms = batched_nms(
data["boxes"].float(),
scores,
torch.zeros(len(data["boxes"])), # categories
iou_threshold=crop_nms_thresh,
)
data.filter(keep_by_nms)
data.to_numpy()
return data
def postprocess_small_regions(
mask_data: MaskData, min_area: int, nms_thresh: float
) -> MaskData:
"""
Removes small disconnected regions and holes in masks, then reruns
box NMS to remove any new duplicates.
Edits mask_data in place.
Requires open-cv as a dependency.
"""
if len(mask_data["rles"]) == 0:
return mask_data
# Filter small disconnected regions and holes
new_masks = []
scores = []
for rle in mask_data["rles"]:
mask = rle_to_mask(rle)
mask, changed = remove_small_regions(mask, min_area, mode="holes")
unchanged = not changed
mask, changed = remove_small_regions(mask, min_area, mode="islands")
unchanged = unchanged and not changed
new_masks.append(torch.as_tensor(mask).unsqueeze(0))
# Give score=0 to changed masks and score=1 to unchanged masks
# so NMS will prefer ones that didn't need postprocessing
scores.append(float(unchanged))
# Recalculate boxes and remove any new duplicates
masks = torch.cat(new_masks, dim=0)
boxes = batched_mask_to_box(masks)
keep_by_nms = batched_nms(
boxes.float(),
torch.as_tensor(scores),
torch.zeros(len(boxes)), # categories
iou_threshold=nms_thresh,
)
# Only recalculate RLEs for masks that have changed
for i_mask in keep_by_nms:
if scores[i_mask] == 0.0:
mask_torch = masks[i_mask].unsqueeze(0)
mask_data["rles"][i_mask] = mask_to_rle_pytorch(mask_torch)[0]
# update res directly
mask_data["boxes"][i_mask] = boxes[i_mask]
mask_data.filter(keep_by_nms)
return mask_data
There are several tunable parameters in automatic mask generation that control how densely points are sampled and what the thresholds are for removing low quality or duplicate masks. Additionally, generation can be automatically run on crops of the image to get improved performance on smaller objects, and post-processing can remove stray pixels and holes
def automatic_mask_generation(
image: np.ndarray,
min_mask_region_area: int = 0,
points_per_side: int = 32,
crop_n_layers: int = 0,
crop_n_points_downscale_factor: int = 1,
crop_overlap_ratio: float = 512 / 1500,
box_nms_thresh: float = 0.7,
crop_nms_thresh: float = 0.7,
) -> List[Dict[str, Any]]:
"""
Generates masks for the given image.
Arguments:
image (np.ndarray): The image to generate masks for, in HWC uint8 format.
Returns:
list(dict(str, any)): A list over records for masks. Each record is
a dict containing the following keys:
segmentation (dict(str, any) or np.ndarray): The mask. If
output_mode='binary_mask', is an array of shape HW. Otherwise,
is a dictionary containing the RLE.
bbox (list(float)): The box around the mask, in XYWH format.
area (int): The area in pixels of the mask.
predicted_iou (float): The model's own prediction of the mask's
quality. This is filtered by the pred_iou_thresh parameter.
point_coords (list(list(float))): The point coordinates input
to the model to generate this mask.
stability_score (float): A measure of the mask's quality. This
is filtered on using the stability_score_thresh parameter.
crop_box (list(float)): The crop of the image used to generate
the mask, given in XYWH format.
"""
point_grids = build_all_layer_point_grids(
points_per_side,
crop_n_layers,
crop_n_points_downscale_factor,
)
mask_data = generate_masks(
image, point_grids, crop_n_layers, crop_overlap_ratio, crop_nms_thresh
)
# Filter small disconnected regions and holes in masks
if min_mask_region_area > 0:
mask_data = postprocess_small_regions(
mask_data,
min_mask_region_area,
max(box_nms_thresh, crop_nms_thresh),
)
mask_data["segmentations"] = [rle_to_mask(rle) for rle in mask_data["rles"]]
# Write mask records
curr_anns = []
for idx in range(len(mask_data["segmentations"])):
ann = {
"segmentation": mask_data["segmentations"][idx],
"area": area_from_rle(mask_data["rles"][idx]),
"bbox": box_xyxy_to_xywh(mask_data["boxes"][idx]).tolist(),
"predicted_iou": mask_data["iou_preds"][idx].item(),
"point_coords": [mask_data["points"][idx].tolist()],
"stability_score": mask_data["stability_score"][idx].item(),
"crop_box": box_xyxy_to_xywh(mask_data["crop_boxes"][idx]).tolist(),
}
curr_anns.append(ann)
return curr_anns
prediction = automatic_mask_generation(image)
automatic_mask_generation returns a list over masks, where each mask is a dictionary containing various data about the mask. These keys are:
segmentation: the maskarea: the area of the mask in pixelsbbox: the boundary box of the mask in XYWH formatpredicted_iou: the model's own prediction for the quality of the maskpoint_coords: the sampled input point that generated this maskstability_score: an additional measure of mask qualitycrop_box: the crop of the image used to generate this mask in XYWH format
print(f"Number of detected masks: {len(prediction)}")
print(f"Annotation keys: {prediction[0].keys()}")
Number of detected masks: 48 Annotation keys: dict_keys(['segmentation', 'area', 'bbox', 'predicted_iou', 'point_coords', 'stability_score', 'crop_box'])
Now we can add these automatically generated masks to the sample and visualize them in the FiftyOne App. We will use a randomly chosen color for each mask:
def add_auto_mask(auto_mask, col, full_mask):
mask = auto_mask["segmentation"].astype(np.uint8)
bbox = auto_mask["bbox"]
x0, y0, x1, y1 = bbox
mask[y0 : y1 + 1, x0 : x1 + 1] *= col
full_mask += mask
return full_mask
full_mask = np.zeros(image.shape[:-1])
for am in sorted(prediction, key=(lambda x: x["area"]), reverse=True):
col = np.random.randint(0, 100)
full_mask = add_auto_mask(am, col, full_mask)
sample["autoseg"] = fo.Segmentation(mask=full_mask)
sample.save()
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.

Curate OpenVINO SAM predictions across an entire dataset¶
With FiftyOne, extending any of these example workflows to a full dataset is easy. Doing so will allow us to visually and programmatically understand the quality of the model's predictions across the entire dataset, or any subset which may be of interest.
For the sake of brevity (this is already a long notebook!), we will just show how to do this for one type of segmentation: namely, we will generate instance segmentations for object detection bounding boxes in a subset of the MS COCO dataset. However, the same process can be applied to any of the other segmentation types, or to any other dataset.
To start, we will load a subset of the MS COCO dataset. In FiftyOne, we can load the dataset (or a subset thereof) from the FiftyOne Dataset Zoo. For example, we can load 100 samples from the validation split of the MS COCO 2017 dataset with the following command:
import fiftyone.zoo as foz
dataset = foz.load_zoo_dataset(
"coco-2017",
split="validation",
max_samples=100,
)
Downloading split 'validation' to '/scratch/user/jacob/zoo_datasets/coco-2017/validation' if necessary Found annotations at '/scratch/user/jacob/zoo_datasets/coco-2017/raw/instances_val2017.json' Sufficient images already downloaded Existing download of split 'validation' is sufficient Loading 'coco-2017' split 'validation' 100% |█████████████████| 100/100 [405.8ms elapsed, 0s remaining, 248.7 samples/s] Dataset 'coco-2017-validation-100' created
We van visualize the dataset in the FiftyOne App:
session = fo.launch_app(dataset, auto=False)
Session launched. Run `session.show()` to open the App in a cell output.
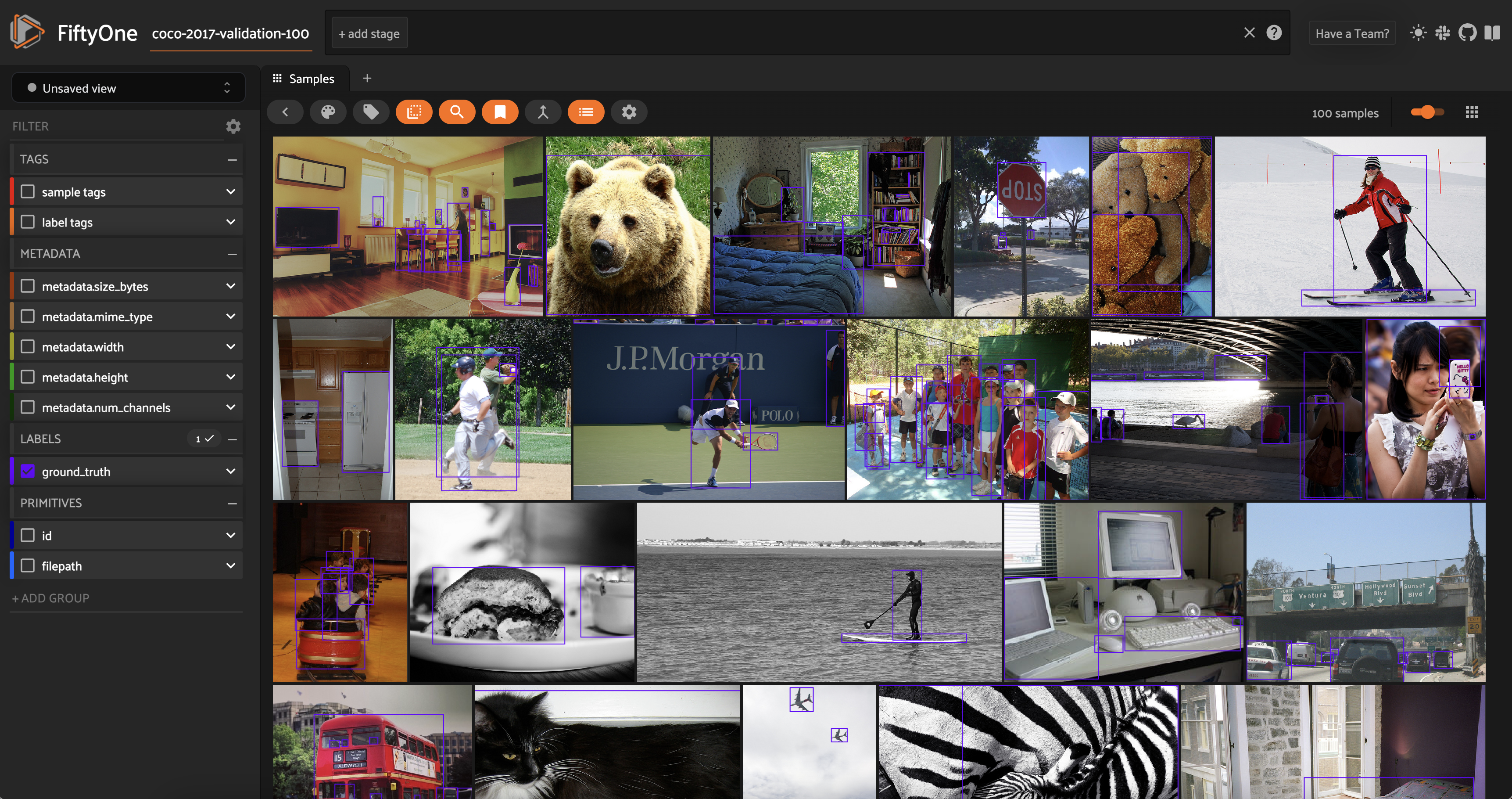
We can see that the dataset contains a variety of images with different types of objects, and that the objects are annotated with bounding boxes. The label field is called ground_truth and contains a Detections label.
When downloading or loading in a new dataset, you also typically need to compute metadata for this dataset as we did for the single-sample example above, so we have easy access to the image width and height in pixels. In this case, the image width and height are already present in the dataset!
dataset.first().metadata
<ImageMetadata: {
'size_bytes': None,
'mime_type': None,
'width': 640,
'height': 426,
'num_channels': None,
}>
We will then iterate through the samples in the dataset, processing the images and adding the image embeddings to each sample:
def generate_image_embeddings(dataset):
for sample in dataset.iter_samples(autosave=True, progress=True):
image = cv2.imread(sample.filepath)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
preprocessed_image = preprocess_image(image)
encoding_results = ov_encoder(preprocessed_image)
image_embeddings = encoding_results[ov_encoder.output(0)]
sample["image_embeddings"] = image_embeddings
dataset.add_sample_field("image_embeddings", fo.ArrayField)
generate_image_embeddings(dataset)
100% |█████████████████| 100/100 [3.0m elapsed, 0s remaining, 0.6 samples/s]
Because we already have the object detection bounding boxes, we can use them as input prompts for SAM. Instead of creating a new label field for the masks, we can just add them to the existing label field in the mask field of each Detection label. We will rename the ground_truth label field to sam to reflect this:
dataset.rename_sample_field("ground_truth", "sam")
We will use the output of the generate_mask() function we defined earlier to generate a mask for each bounding box in an image. We will then truncate these masks to the regions of the image that are covered by the bounding box, and add them to the mask field of the corresponding Detection label.
First, we will define a helper function that will add a mask to a Detection label:
def add_SAM_mask_to_detection(detection, mask, sample):
y0, x0, y1, x1 = convert_fo_to_sam_box(detection.bounding_box, sample)
mask_trimmed = mask[x0 : x1 + 1, y0 : y1 + 1]
detection["mask"] = np.array(mask_trimmed)
return detection
We can then loop over the detections in each sample, generate a mask for each detection, and add it to the mask field of the Detection label. We will do so with the add_SAM_instance_masks_to_sample(). We give the function a field_name argument so that we can use it to add masks to any label field in the dataset, but in this case we will use it to add masks to the sam label field:
def add_SAM_instance_masks_to_sample(sample, field_name = "sam"):
if sample[field_name] is None:
return
dets = sample[field_name].detections
image_embedding = sample['image_embeddings']
input_point = np.array([[0.0, 0.0]])
input_label = np.array([0])
box_labels = np.array([2, 3])
new_dets = []
for det in dets:
fo_box = det.bounding_box
sam_box = convert_fo_to_sam_box(fo_box, sample)
mask = generate_mask(
sample,
input_point,
input_label,
box_coords=sam_box,
box_labels=box_labels,
)
new_dets.append(add_SAM_mask_to_detection(det, mask, sample))
sample[field_name] = fo.Detections(detections=new_dets)
def add_SAM_instance_masks_to_sample(sample, field_name="sam"):
if sample[field_name] is None:
return
dets = sample[field_name].detections
input_point = None
input_label = None
box_labels = np.array([2, 3])
new_dets = []
for det in dets:
fo_box = det.bounding_box
sam_box = convert_fo_to_sam_box(fo_box, sample)
mask = generate_mask(
sample, input_point, input_label, box_coords=sam_box, box_labels=box_labels
)
new_dets.append(add_SAM_mask_to_detection(det, mask, sample))
sample[field_name] = fo.Detections(detections=new_dets)
Finally, we will loop through all of the samples in our dataset and add the SAM masks to the sam label field:
def add_SAM_instance_segmentation_masks(dataset, field_name="sam"):
for sample in dataset.iter_samples(autosave=True, progress = True):
add_SAM_instance_masks_to_sample(sample, field_name=field_name)
Let's run this routine on our dataset and visualize the results in the FiftyOne App, coloring masks by the class of the object they are associated with:
add_SAM_instance_segmentation_masks(dataset)
100% |█████████████████| 100/100 [58.8s elapsed, 0s remaining, 1.4 samples/s]
session = fo.launch_app(dataset, auto = False)
Session launched. Run `session.show()` to open the App in a cell output.

It's that simple! I encourage you to try this - and the other segmentation types - out on your own dataset, and see how SAM performs.