# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
inputFolder = '../input/'
for root, directories, filenames in os.walk(inputFolder):
for filename in filenames: print(os.path.join(root,filename))
../input/rivers/The-River-Sieve-catchment-and-the-location-of-the-different-gauges.png ../input/rivers/Schematic-representation-of-the-proposed-methodology-1-Subdivision-of-the-three.png ../input/rivers/Figure Surface watergroundwater interactions.jpg ../input/rivers/capodacqua-in-val-canneto.jpg ../input/rivers/The-Petrignano-dAssisi-plain-Grey-areas-alluvial-deposits-White-areas-lacustrine-and.png ../input/rivers/ChiascioBastiaUmbra.JPG
from urllib import request
from itertools import product
import pickle
#import tslearn
import scipy.stats as ss
%matplotlib inline
import matplotlib.pyplot as plt
from IPython.display import HTML
import seaborn as sns
!pip install pastas
from IPython.display import Image, display
%config InlineBackend.figure_format = 'retina';
from IPython.core.display import display, HTML
display(HTML("<style>.container {width:70% !important;}</style>"))
imC= Image(url="http://vanoproy.be/css/ChiascioBastiaUmbra.JPG",width=500 ) # filename="/kaggle/input/rivers/ChiascioBastiaUmbra.JPG",
imGWI= Image(url="http://vanoproy.be/python/figs/Figure Surface watergroundwater interactions.jpg", width=600)
#imPUA= Image(filename="/kaggle/input/unconfined/pumped-unconfined-aquifer.jpg", width=600)
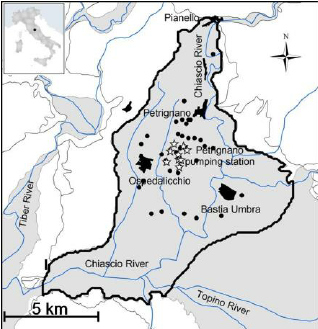
Petrignano Aquifer¶
Published studies on this topic:¶
- On the variables to be considered in assessing the impact of climatechange to alluvial aquifers: a case study in central Italy. (2013), E. Romano, S. Camici, L. Brocca, T. Moramarco, F. Pica, E. Preziosi
- The Sustainable Pumping Rate Concept: Lessons from a Case Study in Central Italy. (2009) Emanuele Romano, Elisabetta Preziosi, Italian National Research Council.
petrignano=pd.read_csv(r"http://vanoproy.be/css/Aquifer_Petrignano.csv", sep=",", engine='python', encoding="UTF-8")# "
petrignano#.head(10)
| Date | Rainfall_Bastia_Umbra | Depth_to_Groundwater_P24 | Depth_to_Groundwater_P25 | Temperature_Bastia_Umbra | Temperature_Petrignano | Volume_C10_Petrignano | Hydrometry_Fiume_Chiascio_Petrignano | |
|---|---|---|---|---|---|---|---|---|
| 0 | 14/03/2006 | NaN | -22.48 | -22.18 | NaN | NaN | NaN | NaN |
| 1 | 15/03/2006 | NaN | -22.38 | -22.14 | NaN | NaN | NaN | NaN |
| 2 | 16/03/2006 | NaN | -22.25 | -22.04 | NaN | NaN | NaN | NaN |
| 3 | 17/03/2006 | NaN | -22.38 | -22.04 | NaN | NaN | NaN | NaN |
| 4 | 18/03/2006 | NaN | -22.60 | -22.04 | NaN | NaN | NaN | NaN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 5218 | 26/06/2020 | 0.0 | -25.68 | -25.07 | 25.7 | 24.5 | -29930.688 | 2.5 |
| 5219 | 27/06/2020 | 0.0 | -25.80 | -25.11 | 26.2 | 25.0 | -31332.960 | 2.4 |
| 5220 | 28/06/2020 | 0.0 | -25.80 | -25.19 | 26.9 | 25.7 | -32120.928 | 2.4 |
| 5221 | 29/06/2020 | 0.0 | -25.78 | -25.18 | 26.9 | 26.0 | -30602.880 | 2.4 |
| 5222 | 30/06/2020 | 0.0 | -25.91 | -25.25 | 27.3 | 26.5 | -31878.144 | 2.4 |
5223 rows × 8 columns
petrignano["Date"] = pd.to_datetime( petrignano.Date, format='%d/%m/%Y' ) # euro dates
petrignano= petrignano.set_index("Date", inplace=False)
petrignano.head(6)
| Rainfall_Bastia_Umbra | Depth_to_Groundwater_P24 | Depth_to_Groundwater_P25 | Temperature_Bastia_Umbra | Temperature_Petrignano | Volume_C10_Petrignano | Hydrometry_Fiume_Chiascio_Petrignano | |
|---|---|---|---|---|---|---|---|
| Date | |||||||
| 2006-03-14 | NaN | -22.48 | -22.18 | NaN | NaN | NaN | NaN |
| 2006-03-15 | NaN | -22.38 | -22.14 | NaN | NaN | NaN | NaN |
| 2006-03-16 | NaN | -22.25 | -22.04 | NaN | NaN | NaN | NaN |
| 2006-03-17 | NaN | -22.38 | -22.04 | NaN | NaN | NaN | NaN |
| 2006-03-18 | NaN | -22.60 | -22.04 | NaN | NaN | NaN | NaN |
| 2006-03-19 | NaN | -22.35 | -21.95 | NaN | NaN | NaN | NaN |
petrignano.info()
dfp =petrignano.rename(columns={'Rainfall_Bastia_Umbra':'Rain_Bastia','Depth_to_Groundwater_P24':'Depth_to_P24','Depth_to_Groundwater_P25':'Depth_to_P25',
'Temperature_Bastia_Umbra':'Temp_Bastia', 'Temperature_Petrignano':'Temp_Petrig',"Volume_C10_Petrignano":"Volume_C10",
"Hydrometry_Fiume_Chiascio_Petrignano": "Hydrometry"})
dfp
| Rain_Bastia | Depth_to_P24 | Depth_to_P25 | Temp_Bastia | Temp_Petrig | Volume_C10 | Hydrometry | |
|---|---|---|---|---|---|---|---|
| Date | |||||||
| 2006-03-14 | NaN | -22.48 | -22.18 | NaN | NaN | NaN | NaN |
| 2006-03-15 | NaN | -22.38 | -22.14 | NaN | NaN | NaN | NaN |
| 2006-03-16 | NaN | -22.25 | -22.04 | NaN | NaN | NaN | NaN |
| 2006-03-17 | NaN | -22.38 | -22.04 | NaN | NaN | NaN | NaN |
| 2006-03-18 | NaN | -22.60 | -22.04 | NaN | NaN | NaN | NaN |
| ... | ... | ... | ... | ... | ... | ... | ... |
| 2020-06-26 | 0.0 | -25.68 | -25.07 | 25.7 | 24.5 | -29930.688 | 2.5 |
| 2020-06-27 | 0.0 | -25.80 | -25.11 | 26.2 | 25.0 | -31332.960 | 2.4 |
| 2020-06-28 | 0.0 | -25.80 | -25.19 | 26.9 | 25.7 | -32120.928 | 2.4 |
| 2020-06-29 | 0.0 | -25.78 | -25.18 | 26.9 | 26.0 | -30602.880 | 2.4 |
| 2020-06-30 | 0.0 | -25.91 | -25.25 | 27.3 | 26.5 | -31878.144 | 2.4 |
5223 rows × 7 columns
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(4,1, figsize=(21, 9), sharex=True) #, squeeze=False
sns.lineplot(x="Date", y= petrignano["Rainfall_Bastia_Umbra"],data=petrignano, ax=ax[0])
sns.lineplot(x="Date", y= petrignano["Depth_to_Groundwater_P24"], data=petrignano, ax=ax[1]) # ax=ax
sns.lineplot(x="Date", y= petrignano["Depth_to_Groundwater_P25"], data=petrignano, ax=ax[2]) # ax=ax
sns.lineplot(x="Date", y= petrignano["Hydrometry_Fiume_Chiascio_Petrignano"], data=petrignano, ax=ax[3]) # ax=ax
plt.xticks(rotation=90); plt.grid(b=True,) #axe.setxaxis(rotation=90)
plt.xlim(pd.to_datetime( "2010-01-01" ),pd.to_datetime("2020-02-01" ) );
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(4,1, figsize=(21, 9), sharex=True) #, squeeze=False
sns.lineplot(x="Date", y= petrignano["Rainfall_Bastia_Umbra"],data=petrignano, ax=ax[0])
sns.lineplot(x="Date", y= petrignano["Depth_to_Groundwater_P25"], data=petrignano, ax=ax[1]) # ax=ax
sns.lineplot(x="Date", y= petrignano["Temperature_Petrignano"], data=petrignano, ax=ax[2]) # ax=ax
sns.lineplot(x="Date", y= petrignano["Hydrometry_Fiume_Chiascio_Petrignano"], data=petrignano, ax=ax[3]) # ax=ax
plt.xticks(rotation=90); plt.grid(b=True,) #axe.setxaxis(rotation=90)
plt.xlim(pd.to_datetime( "2012-01-01" ),pd.to_datetime("2015-02-01" ) );
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(4,1, figsize=(21, 9), sharex=True) #, squeeze=False
sns.lineplot(x="Date", y= petrignano["Rainfall_Bastia_Umbra"],data=petrignano, ax=ax[0])
sns.lineplot(x="Date", y= petrignano["Depth_to_Groundwater_P24"], data=petrignano, ax=ax[1]); plt.ylim(-30,-25) # ax=ax
sns.lineplot(x="Date", y= petrignano["Temperature_Petrignano"], data=petrignano, ax=ax[2]); plt.ylim(0,4)# ax=ax
sns.lineplot(x="Date", y= petrignano["Hydrometry_Fiume_Chiascio_Petrignano"], data=petrignano, ax=ax[3]);plt.ylim(0,4) # ax=ax
plt.xticks(rotation=90); plt.grid(b=True,) #axe.setxaxis(rotation=90)
plt.xlim(pd.to_datetime( "2018-11-01" ),pd.to_datetime("2019-04-01" ) );
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(2,1, figsize=(21, 9), sharex=True) #, squeeze=False
sns.lineplot(x="Date", y= dfp["Depth_to_P24"],data=dfp, ax=ax[0])
sns.lineplot(x="Date", y= dfp["Depth_to_P25"],data=dfp, ax=ax[0],color="forestgreen") # ax=ax
sns.lineplot(x="Date", y= dfp["Volume_C10"], data=dfp, ax=ax[1] ,color="olivedrab") # ax=ax
plt.xticks(rotation=90); plt.grid(b=True,) #
plt.xlim(pd.to_datetime( "2006-09-19" ),pd.to_datetime("2007-09-01" ) );
Data refurbishment¶
We'll interpolate the missing values in the height data, and turn negative pumped volumes positive:
dfp.Depth_to_P24 =dfp.Depth_to_P24.interpolate(method="linear", limit_direction='forward')
dfp.Depth_to_P25 =dfp.Depth_to_P25.interpolate(method="linear", limit_direction='forward')
dfp.Volume_C10= -dfp.Volume_C10
Convertion of the volume of pumped water, expressed in cubic meters (mc) per day, to m³/s.
dfp["Debiet"]= dfp.Volume_C10/(3600*24)
dfp.describe()
| Rain_Bastia | Depth_to_P24 | Depth_to_P25 | Temp_Bastia | Temp_Petrig | Volume_C10 | Hydrometry | Debiet | |
|---|---|---|---|---|---|---|---|---|
| count | 4199.000000 | 5223.000000 | 5223.000000 | 4199.000000 | 4199.000000 | 5025.000000 | 4199.000000 | 5025.000000 |
| mean | 1.556633 | -26.279945 | -25.696110 | 15.030293 | 13.739081 | 29043.296726 | 2.372517 | 0.336149 |
| std | 5.217923 | 3.314307 | 3.222391 | 7.794871 | 7.701369 | 4751.864371 | 0.589088 | 0.054998 |
| min | 0.000000 | -34.470000 | -33.710000 | -3.700000 | -4.200000 | -0.000000 | 0.000000 | -0.000000 |
| 25% | 0.000000 | -28.265000 | -27.630000 | 8.800000 | 7.700000 | 26218.080000 | 2.100000 | 0.303450 |
| 50% | 0.000000 | -26.020000 | -25.540000 | 14.700000 | 13.500000 | 28689.120000 | 2.400000 | 0.332050 |
| 75% | 0.100000 | -23.840000 | -23.430000 | 21.400000 | 20.000000 | 31678.560000 | 2.700000 | 0.366650 |
| max | 67.300000 | -19.660000 | -19.100000 | 33.000000 | 31.100000 | 45544.896000 | 4.100000 | 0.527140 |
During the first years P25's level is initially higher during the pumping, so it must be further away from well than P24. But the average pump rate lowers around 2014-2015, so the aquifer can get the opportunity to recharge partially.
print("P24min",dfp.Depth_to_P24.min(),"P25min", dfp.Depth_to_P25.min(),"P24max", dfp.Depth_to_P24.max(),"P25max",dfp.Depth_to_P25.max(),
round(dfp.Depth_to_P24.max()-dfp.Depth_to_P24.min(),2),dfp.Depth_to_P25.max()-dfp.Depth_to_P25.min())
P24min -34.47 P25min -33.71 P24max -19.66 P25max -19.1 14.81 14.61
I suppose that the original base of the suction cone is higher than the max. height of the provided depths. Later I'll use other heights based on groundlevel altitude above sealevel.
dfp["Hth_to_P24"]= dfp.Depth_to_P24.max()-dfp.Depth_to_P24
dfp["Hth_to_P25"]= dfp.Depth_to_P25.max()-dfp.Depth_to_P25;
dfp["Hth_to_P25"].plot();
Note that 'Hth' is the heigth of the suction cone at a well, above the estimated "original" water table base level. In the plot you can notice that the average diff. in relative height shifts to the other side after 2013.
depthdiff= dfp.Hth_to_P24 -dfp.Hth_to_P25; sns.lineplot(data=depthdiff );
Somewhere in a published study I read that in the center of the aquifer the thickness is about 80 meters. I also saw in a published study that there has been an piezometric level of 170 meters in the aquifer center anno 2004. The location has an altitude of 200 m above sealevel.
dfp["CHt_P24"]= 80+ dfp.Depth_to_P24 # cone height at p24
dfp["CHt_P25"]= 80 +dfp.Depth_to_P25;
dfp["CHt_P25"].plot();
We converted the water table level another time to be able to use the Dupuit formula, and a sea level heigth.
dfpsl=pd.DataFrame()
dfpsl["head24"]= 200+dfp.Depth_to_P24 ;
dfpsl["head25"]= 200+dfp.Depth_to_P25;
dfpsl.plot( figsize=(19, 4));
dfp["head24"]= 200+dfp.Depth_to_P24 ;
dfp["head25"]= 200+dfp.Depth_to_P25;
This is an image or a cross section of a pumped unconfined aquifer.
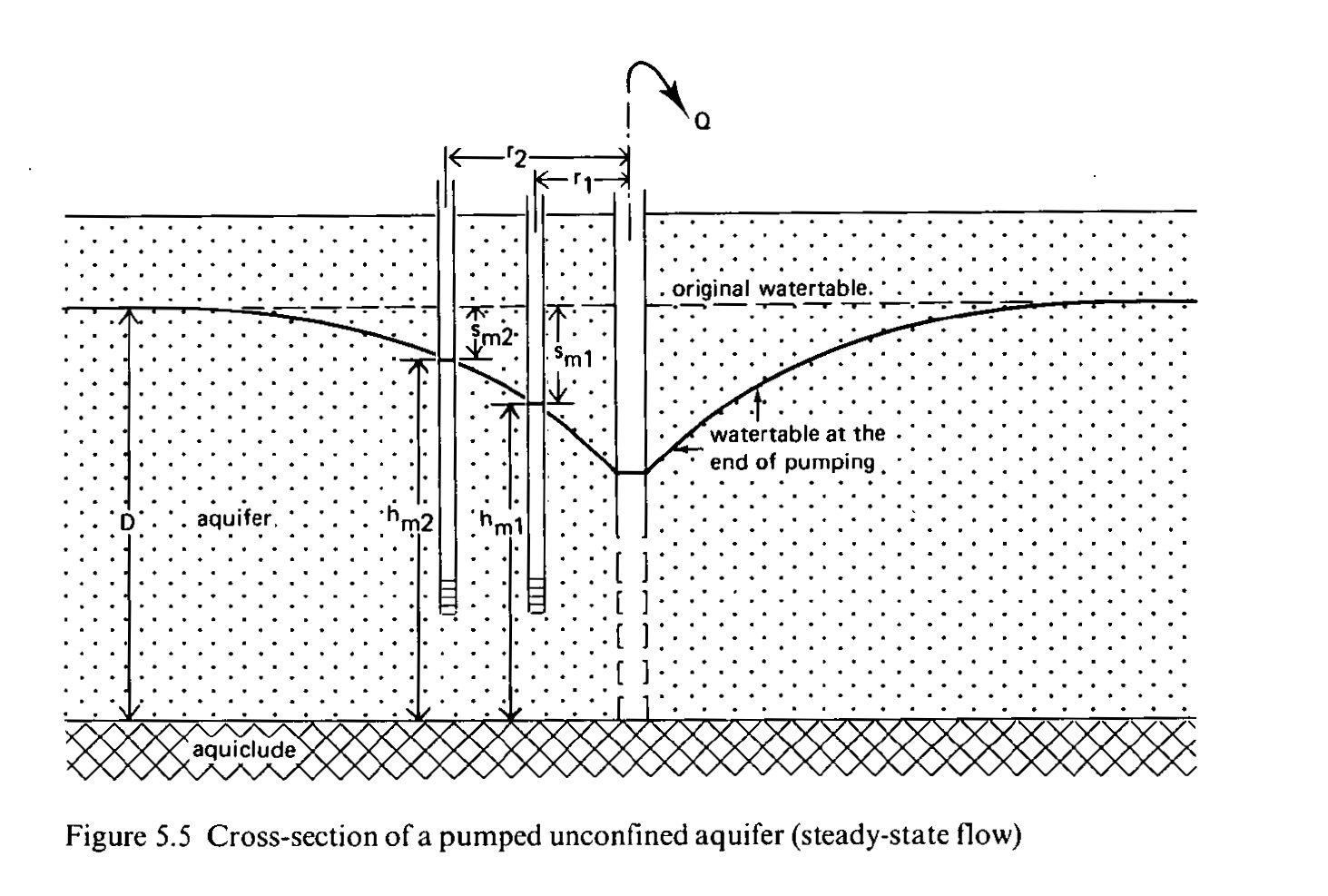
The next formula, based on the Dupuit formula, should result in a constant, but in this case it isn't, as there are fluctuations in the density of the soil. Also, there is competition from other pumping activities, and when they are excessive or nearby executed, they can lower the shape of the main well suction cone. The formula does not work near the well due to seepage.
In case the piezometer most far away from the well, P25, is located between P24 and the river, then water from the river can have positive or negative effects. But from the plots I deduct that the river water takes a month or more to arrive at the well site, so the effect on the slope is mild compared to others.
dfp["Hth2diff"]= (dfp.CHt_P25.rolling(7,min_periods=5).sum().fillna(method='bfill', limit=7) **2 -dfp.CHt_P24.rolling(7,min_periods=5).sum().fillna(method='bfill', limit=7)**2)/ (0.01+dfp.Volume_C10.rolling(7).sum().fillna(method='bfill', limit=7))
#dfp["Hth2diff"]= dfp["Hth2diff"].replace(np.nan,0) # why is this here, bit early
#dfp["Hth2diff"]= dfp.where( lambda x: dfp[dfp.Volume_C10==0], lambda x: 0)
dfp["Hth2diff_S1"]= (dfp.CHt_P25.rolling(7,min_periods=3).sum().fillna(method='bfill', limit=7).shift(1)**2 -dfp.CHt_P24.rolling(7,min_periods=3).sum().fillna(method='bfill', limit=7)**2)/(0.01+dfp.Volume_C10)
dfp["Hth2diff_S11"]= (dfp.CHt_P25.rolling(7,min_periods=3).sum().fillna(method='bfill', limit=7).shift(1)**2 -dfp.CHt_P24.rolling(7,min_periods=3).sum().fillna(method='bfill', limit=7).shift(1)**2)/(0.01+dfp.Volume_C10)
dfp["Hth2diff_S21"]= (dfp.CHt_P25.shift(2)**2 -dfp.CHt_P24.shift(1)**2)/(0.01+dfp.Volume_C10)
a plot with time shifts to get a better view on the fluctuations of the water levels.
sns.set_style("whitegrid", {'grid.linestyle': '--'}); fig, ax = plt.subplots(1,1, figsize=(23, 5), sharex=True)
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S1"], data=dfp) ;#
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S11"], data=dfp ); # 2014-04-13 0.011207
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S21"], data=dfp );
sns.lineplot(x=dfp.index, y=dfp["Hth2diff"], data=dfp,ax=ax ); plt.ylim( .0,.03)
#sns.lineplot(x=dfp.index, y=dfp["Volume_C10"], data=dfp,ax=ax[1] ); plt.ylim( 15000,40000)
plt.xlim(pd.to_datetime( "2012-07-02" ),pd.to_datetime("2014-04-01" ) ); # "2010-04-01"
sns.set_style("whitegrid", {'grid.linestyle': '--'}); fig, ax = plt.subplots(1,1, figsize=(23, 5), sharex=True)
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S1"], data=dfp) ;#
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S11"], data=dfp ); # 2014-04-13 0.011207
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff_S21"], data=dfp );
#sns.lineplot(x=dfp.index, y=dfp["Hth2diff"], data=dfp,ax=ax[0] ); plt.ylim( .0,.03)
sns.lineplot(x=dfp.index, y=dfp["Volume_C10"].rolling(90).sum() , data=dfp,ax=ax );
sns.lineplot(x=dfp.index, y=dfp["Volume_C10"], data=dfp,ax=ax ); plt.ylim( 20000,40000)
plt.xlim(pd.to_datetime( "2012-07-02" ),pd.to_datetime("2014-04-01" ) ); # "2010-04-01"
dfp["Hth2diff"]["2009-10-02":"2009-10-22"]
Date 2009-10-02 0.014837 2009-10-03 0.014863 2009-10-04 0.014794 2009-10-05 0.014341 2009-10-06 0.014232 2009-10-07 0.014128 2009-10-08 0.014168 2009-10-09 0.014004 2009-10-10 0.013911 2009-10-11 0.013761 2009-10-12 0.014191 2009-10-13 0.014145 2009-10-14 0.014043 2009-10-15 0.013934 2009-10-16 0.013846 2009-10-17 0.013673 2009-10-18 0.013800 2009-10-19 0.013541 2009-10-20 0.013652 2009-10-21 0.013670 2009-10-22 0.013503 Name: Hth2diff, dtype: float64
dfp["Hth2diff"].describe()
count 5027.000000 mean 99.537152 std 4994.655440 min -0.004075 25% 0.011687 50% 0.014767 75% 0.018150 max 262023.170000 Name: Hth2diff, dtype: float64
kwadrdf= dfp[["Volume_C10","Hth2diff_S11","Hth2diff_S21","Hth2diff_S1", "Hth2diff"]]
sns.heatmap( kwadrdf.corr(), annot=True );
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(1,1, figsize=(19, 5), sharex=True);
sns.lineplot(data=dfp.head24, ax=ax) ;
sns.lineplot(data=dfp.head25, ax=ax); plt.ylim( 165,180);
ax2= plt.twinx()
sns.lineplot( data=dfp.Volume_C10, ax=ax2,color="navy")
#plt.xlim(pd.to_datetime( "2006-05-01" ),pd.to_datetime("2010-07-01" ) );
plt.xlim(pd.to_datetime( "2008-11-30"),pd.to_datetime("2010-12-15" ) ); plt.ylim( 20000,45000);
Dates which are lacking pumping data: 9/05/2007, 30/08/2007, 17/04/2008, 9/06/2011 and 10/06/2011, 20/12/2011; 12/07/2019 and 14/07/2019 No pumping periods: 2019-06-28: 2019-07-05], 2019-08-01: 2019-08-05]
import datetime
datetime.datetime(2006,9,25,0,0)
datetime.datetime(2006, 9, 25, 0, 0)
Rainfall¶
print("Estimated drainage area of the aquifer:", 11.5*12.5/2, "km²")
Estimated drainage area of the aquifer: 71.875 km²
Conversion of rainfall units: mm/d to m³/d, for better comparison with pump abstraction values. Also the creation of an indicator for the month to be able to distinguish the months.
The actual drainage area for the source is not given, and here is an estimation based on info in a study of the geology of the alluvial plane.
pd.set_option('mode.chained_assignment',None)
dfp["Month"] =dfp.index.month
dfp["Rain_M³"] =dfp.Rain_Bastia* 72*1000.0*1000/ 1000 # 27km²
dfp["Rain_Log"] = np.log1p(dfp.Rain_Bastia) # Rain_M³
print("Estimated effective infiltration by the rain to aquifer:", round(250/(365*24*3600 ),8), "L/sec or ", round(250/365,2),"L/day")
effect_infil= 250/365* 72*1000.0; print( effect_infil, "m³")
Estimated effective infiltration by the rain to aquifer: 7.93e-06 L/sec or 0.68 L/day 49315.068493150684 m³
dfp["Rainfall_3"] =dfp.Rain_Bastia.rolling(3).sum().fillna(method='bfill', limit=3)
dfp["Rainfall_7"] =dfp.Rain_Bastia.rolling(7).sum().fillna(method='bfill', limit=7)
dfp["Rainfall_122"] =dfp.Rain_Bastia.rolling(122).sum().fillna(method='bfill', limit=122)
dfp["Rainfall_30"] =dfp.Rain_Bastia.rolling(30).sum().fillna(method='bfill', limit=30)
Temperature¶
We have obtained 2 temperature sets: Temp_Bastia and Temp_Petrig.
The temperature data for Petrignano in 2015 are flatlining from 2015-04-25 til 2015-09-21, 0's, and the weeks before that the device seems to be no longer correctly calibrated. What shall we do with such a series?
I'll also create a column to simulate a minimum temperature.
fig, ax = plt.subplots(1,1, figsize=(19, 6), sharex=True)
sns.lineplot(data=dfp.Temp_Bastia) ;
sns.lineplot(data=dfp.Temp_Petrig);
plt.xlim(pd.to_datetime( "2014-02-01" ),pd.to_datetime("2015-05-01" ) ); plt.ylim(-0.5,30);
We perform a temperature correction for the Bastia data in the mentioned time period, in order to fill up the Petrignano data and use it.
dfp.Temp_Petrig["2015-04-25":"2015-09-21"] = dfp.Temp_Bastia["2015-04-25":"2015-09-21"]-0.75
dfp["TmnStdB"] =dfp.Temp_Bastia- dfp.Temp_Bastia.std(skipna=True)
dfp["TmnStdP"] =dfp.Temp_Petrig- dfp.Temp_Petrig.std(skipna=True)
Add some temperature rolling means that have a sort of minimum temp. behaviour.
dfp["TmnStdP_4"] =dfp.TmnStdP.rolling(4).mean().fillna(method='bfill', limit=3)
dfp["TP_7"] =dfp.Temp_Petrig.rolling(7).mean().fillna(method='bfill', limit=7)
On 2020-06-15 there is a nan in the pumped volume, which is the only exception. The aquifer might no longer be able to provide the amount of water...
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(1,1, figsize=(19, 4.5), sharex=True);
sns.lineplot(data=dfp.head24, ax=ax) ;
sns.lineplot(data=dfp.head25, ax=ax); plt.ylim( 173,176);
ax2= plt.twinx()
sns.lineplot( data=dfp.Volume_C10, ax=ax2,color="navy")
plt.xlim(pd.to_datetime( "2020-01-30"),pd.to_datetime("2020-06-30" ) ); plt.ylim( 15000,35000);
fig, ax = plt.subplots(figsize=(17,5)) sns.lineplot(x= y_test.index,y= y_test.head24, data=y_test) #CHt_P25 Hth_to_P25 sns.lineplot(x= y_test.index,y= y_test.head25, data=y_test)
sns.lineplot(x= y_test.index,y= y_pred[:,0], data=y_pred) sns.lineplot(x= y_test.index,y= y_pred[:,1], data=y_pred);
Piezometer 24 has a nan at 2006-12-14, and from then on til 2007-04-28, it stays off indicating values about 25 cm higher. I'll subtract it
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(1,1, figsize=(19, 5), sharex=True);
sns.lineplot(data=dfp.head24, ax=ax) ;
sns.lineplot(data=dfp.head25, ax=ax); plt.ylim( 165,180);
ax2= plt.twinx()
sns.lineplot( data=dfp.Volume_C10, ax=ax2,color="navy")
plt.xlim(pd.to_datetime( "2008-11-30"),pd.to_datetime("2010-12-15" ) ); plt.ylim( 20000,45000);
No pumping¶
- Dates which are lacking pumping data: 9/05/2007, 30/08/2007, 17/04/2008, 9/06/2011 and 10/06/2011, 20/12/2011; 12/07/2019 and 14/07/2019
- No pumping periods: 2019-06-28: 2019-07-05], 2019-08-01: 2019-08-05]
Rainfall drainage area¶
print("Estimated drainage area of the aquifer:", 11.5*12.5/2, "km²", " study based area: 72-75 km²")
Estimated drainage area of the aquifer: 71.875 km² study based area: 72-75 km²
Conversion of rainfall and discharge units: mm/d to m³/d, and creation of an indicator for the month to be able to distinguish the months or seasons.
The actual drainage area for the source is not given, and here is an estimation based on info in a study of the geology of the alluvial plane.
pd.set_option('mode.chained_assignment',None)
dfp["Month"] =dfp.index.month
dfp["Rain_M³"] =dfp.Rain_Bastia* 721000.01000/ 1000 # 72km²
dfp["Rain_Log"] = np.log1p(dfp.Rain_Bastia) # Rain_M³
print("Estimated effective infiltration by the rain to aquifer:", round(250/(365243600 ),8), "L/sec or ", round(250/365,2),"L/day") dfp["Rainfall_3"] =dfp.Rain_Bastia.rolling(3).sum().fillna(method='bfill') dfp["Rainfall_7"] =dfp.Rain_Bastia.rolling(7).sum().fillna(method='bfill') dfp["Rainfall_122"] =dfp.Rain_Bastia.rolling(122).sum().fillna(method='bfill') dfp["Rainfall_30"] =dfp.Rain_Bastia.rolling(30).sum().fillna(method='bfill')
Discharge¶
Discharge, outtake, or pump volume, abstractions... are synonyms. There are some days when the volume is 0, which is caused by human intervention for reason of aquifer exhaustion, maintenance, safety inspection... These interruptions are exogenous in my opinion. I'm not sure what to do with them: drop, interpolate, ...
dfp.loc["Jun 2020"].Volume_C10= dfp.loc["Jun 2020"].Volume_C10.interpolate( limit_direction="forward")
dfp["Vol_124"]=dfp.Volume_C10.rolling(124).sum().fillna(method='bfill', limit=123) #
dfp["Vol_22"] =dfp.Volume_C10.rolling(22).sum().fillna(method='bfill', limit=22)
dfp["Vol_33"] =dfp.Volume_C10.rolling(30).sum().fillna(method='bfill', limit=30)
dfp["Vol_47"] =dfp.Volume_C10.rolling(47).sum().fillna(method='bfill', limit=47)
dfp.loc["2015-04-25":"2015-09-21"]
River Chiascio¶
The discharge average of the river is 20 m³/s. Let's find out the average of the water level over several years.
imC
riverday= 20*3600*24; riveryr= 20*3600*24*365
print(f"Discharge average is {riverday:5,.0f} m³/d, {riveryr:5,.0f} m³/year")
dfp['doy'] = dfp.index.dayofyear
dfp['Year'] = dfp.index.year
dfp['Week'] = dfp.index.isocalendar().week
Discharge average is 1,728,000 m³/d, 630,720,000 m³/year
The river water level is 0 between 25/04/2015 and 21/09/2015. This could be a meteo station error or outfall. Shall I use nan?
No, I'll compute the average of river level for this period over several adjacent years.
piv = pd.pivot_table(dfp, index=['doy'],columns=['Year'], values=['Hydrometry'])
piv.plot( figsize=(19.8, 4)); # .groupby("Year")
I gonna make the average of timeseries data of adjacent years river level series to replace the missing data.
rivergap=pd.concat([dfp["2014-04-25":"2014-09-21"], dfp["2016-04-25":"2016-09-21"],dfp["2017-04-25":"2017-09-21"],dfp["2018-04-25":"2018-09-21"] ],ignore_index=True); rivergap
| Rain_Bastia | Depth_to_P24 | Depth_to_P25 | Temp_Bastia | Temp_Petrig | Volume_C10 | Hydrometry | Debiet | Hth_to_P24 | Hth_to_P25 | ... | TmnStdP | TmnStdP_4 | TP_7 | Vol_124 | Vol_22 | Vol_33 | Vol_47 | doy | Year | Week | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 2.5 | -20.09 | -19.25 | 16.5 | 15.7 | 27427.680 | 2.8 | 0.31745 | 0.43 | 0.15 | ... | 8.242949 | 8.442949 | 14.400000 | 3615595.488 | 607848.192 | 844175.520 | 1344589.632 | 115 | 2014 | 17 |
| 1 | 0.1 | -19.99 | -19.24 | 17.0 | 16.2 | 26551.584 | 2.8 | 0.30731 | 0.33 | 0.14 | ... | 8.742949 | 8.242949 | 15.214286 | 3615242.112 | 606611.808 | 843095.520 | 1336940.640 | 116 | 2014 | 17 |
| 2 | 7.8 | -19.71 | -19.13 | 14.3 | 13.3 | 24432.192 | 2.7 | 0.28278 | 0.05 | 0.03 | ... | 5.842949 | 7.542949 | 15.485714 | 3611040.480 | 602445.600 | 839655.072 | 1333412.928 | 117 | 2014 | 17 |
| 3 | 1.8 | -19.80 | -19.16 | 12.3 | 11.6 | 24573.024 | 2.7 | 0.28441 | 0.14 | 0.06 | ... | 4.142949 | 6.742949 | 14.957143 | 3609717.696 | 598459.104 | 833086.080 | 1328408.640 | 118 | 2014 | 18 |
| 4 | 0.1 | -19.91 | -19.19 | 12.8 | 11.9 | 25180.416 | 2.7 | 0.29144 | 0.25 | 0.09 | ... | 4.442949 | 5.792949 | 14.228571 | 3611186.496 | 595231.200 | 828808.416 | 1322829.792 | 119 | 2014 | 18 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 595 | 7.6 | -25.74 | -25.18 | 23.0 | 22.2 | 31367.520 | 2.3 | 0.36305 | 6.08 | 6.08 | ... | 14.742949 | 15.992949 | 23.185714 | 3717809.280 | 682622.208 | 934350.336 | 1444591.584 | 260 | 2018 | 38 |
| 596 | 13.0 | -25.78 | -25.25 | 22.3 | 21.7 | 29403.648 | 2.4 | 0.34032 | 6.12 | 6.15 | ... | 14.242949 | 15.542949 | 23.014286 | 3720457.440 | 683049.888 | 934301.952 | 1440867.744 | 261 | 2018 | 38 |
| 597 | 0.2 | -25.90 | -25.28 | 21.9 | 21.8 | 29782.944 | 2.3 | 0.34471 | 6.24 | 6.18 | ... | 14.342949 | 15.092949 | 22.700000 | 3723190.272 | 683335.872 | 932164.416 | 1444997.664 | 262 | 2018 | 38 |
| 598 | 0.0 | -25.65 | -25.20 | 22.5 | 22.5 | 27522.720 | 2.3 | 0.31855 | 5.99 | 6.10 | ... | 15.042949 | 14.592949 | 22.828571 | 3722259.744 | 680270.400 | 926876.736 | 1438293.024 | 263 | 2018 | 38 |
| 599 | 0.0 | -25.70 | -25.19 | 23.3 | 23.2 | 30092.256 | 2.2 | 0.34829 | 6.04 | 6.09 | ... | 15.742949 | 14.842949 | 22.785714 | 3725691.552 | 678843.936 | 924373.728 | 1438006.176 | 264 | 2018 | 38 |
600 rows × 34 columns
riverHydro=rivergap.loc[:,["Hydrometry","doy"]];
riverHydroavg=riverHydro.groupby("doy").mean() # .Hydrometry axis=1
riverHydro.groupby("doy").mean().plot();
dfp.groupby("doy")["Hydrometry"].mean().plot();
and we update the original with the averages of the adjacent years for the period which has missing data.
riverHydroavg= riverHydroavg[:-1]
riverHydroavg.index = dfp["2015-04-25":"2015-09-21"].index
dfp.update(riverHydroavg)
Collected information and considerations¶
This is information that I retrieved from publicated studies. I also added my own considerations.
- I wanna make an indicator or weighted factor, based on the timely water budget of aquifer state and water level of the stream.
- How far is the river away from the measure pits?
- About 1900 meters, which means this river water can take a while to flow through the soil to the suction coil.
- Can I find out which measure pits is closest to the stream Chiascio?
- P25 had the heighest level during the intensive pumping times.
- The influence of the river level on the difference in cone height vs. the relative head of the closest measurement.
Some author wrote that the aquifer hasn't been in a steady-state since 2001.
The possible recharge of the aquifer, and the difference between surface water and groundwater levels, are influenced by pumping rate. But, there is also well pumping by agricultural and private actors happening, esp. when it's warm and dry (during May-September).
This is a dynamic system, which consists of 4 drivers: recharge by rain, aquifer storage, pump outtake and drain to river. The latter can be positive or negative.
The variation in recharge influences the aquifer storage and the drain to river, and the variation in abstraction influences storage and drain to river. These variations have different time lags.
On top of that there are 2 scenario's: the northern part of the river can feed the aquifer, while the southern part of the stream role depends on the balance btn. groundwater level and the amount of abstractions.
imGWI
dfp["diff_surface_gwater"] =((dfp.Hydrometry.rolling(128).mean(min_periods=1))*(dfp.head25**2-dfp.head24**2))/ -(dfp.head24/(0.00000000001+dfp.head24.max()-dfp.head24.min()))#**2 #.rolling(3).sum()
dfp["2009":"2020"]["diff_surface_gwater"].plot(); # -np.log1p( .fillna('bfill')
sns.set_style("whitegrid", {'grid.linestyle': '--'}) fig, ax = plt.subplots(1,1, figsize=(19, 4.5), sharex=True); sns.lineplot(data=dfp.Rain_Bastia.rolling(3).mean(), ax=ax) ; sns.lineplot(data=dfp.diff_surface_gwater.rolling(3).mean(), ax=ax); #plt.ylim( 0,8); ax2= plt.twinx() sns.lineplot( data=dfp.Volume_C10.rolling(3).mean(), ax=ax2,color="navy"); plt.ylim( 20000,35000); plt.xlim(pd.to_datetime( "2010-01-30"),pd.to_datetime("2010-12-30" ) );
dfp["Hyd_39"] =dfp.Hydrometry.rolling(39).sum().fillna(method='bfill', limit=39) # level to m³ ? or
dfp["Hyd_128"] =dfp.Hydrometry.rolling(128).sum().fillna(method='bfill', limit=128)
#dfp["HydStd_8"]=dfp.Hydrometry.rolling(8).std().fillna(method='bfill')
dfp["2009":"2020"].Hydrometry.plot( lw=0.5); # level to m³ ?
Minimum and max. ground water levels over the years¶
From 2008 to 2012 there has been a period of exhaustion of the aquifer storage, which led to more and more lower headings for the water levels. Since 2013-2014 the pumping rate has been somewhat reduced in order to give the aquifer time to replenish.
dfp['Year'] = dfp.index.year
Min_year = dfp["2008-01-01":"2020-04-01"].groupby('Year')[["Hth_to_P24","Hth_to_P25"]].min();
MinVol_year = dfp["2008-01-01":"2020-04-01"].groupby('Year')[["Volume_C10"]].mean();
Max_year = dfp["2008-01-01":"2020-04-01"].groupby('Year')[["Hth_to_P24","Hth_to_P25"]].min();
MaxVol_year = dfp["2008-01-01":"2020-04-01"].groupby('Year')[["Volume_C10"]].max();
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(2,1, figsize=(10.7, 6), sharex=True) #
plt.grid()
sns.lineplot(data=Min_year, label="Min height year",ax=ax[0] )# x=Rain_Bastia_month.index, y= "Rain_Bastia",
sns.lineplot(data=Max_year, label="Max height year",ax=ax[0] )
#ax[0].axis(xmin=0 ,xmax=15 ) # xlim( 0,) x=Volume_C10_month.index, y= "Volume_C10",
sns.lineplot(data=MinVol_year, label="Avg vol year",color="coral", ax=ax[1]) # ax=ax
sns.lineplot(data=MaxVol_year, label="Max vol",color="coral", ax=ax[1])
#sns.lineplot(x=dfp.index, y= "RainNO_2", data=dfp, label="Rain NE", ax=ax[2]) #
plt.xticks(rotation=90);
#plt.xlim(pd.to_datetime( "2010-01-01" ),pd.to_datetime("2019-12-31" ) );
Monthly rainfall and pump volume¶
Rain_Bastia_month = dfp["2009-01-01":"2020-04-01"].groupby('Month')[["Rain_Bastia"]].sum();
Volume_C10_month = dfp["2009-01-01":"2020-04-01"].groupby('Month')[["Volume_C10"]].mean();
Volume_sum_month = dfp["2009-01-01":"2020-04-01"].groupby('Month')[["Volume_C10"]].sum();
Hydro_month = dfp["2009-01-01":"2020-04-01"].groupby('Month')[["Hydrometry"]].sum();
Volume_C10_month
| Volume_C10 | |
|---|---|
| Month | |
| 1 | 28521.204387 |
| 2 | 28203.098336 |
| 3 | 27683.219613 |
| 4 | 27834.916350 |
| 5 | 27676.240891 |
| 6 | 28236.006982 |
| 7 | 29313.153501 |
| 8 | 29050.016094 |
| 9 | 29954.149527 |
| 10 | 30072.384000 |
| 11 | 29789.395200 |
| 12 | 28688.220528 |
sns.set_style("whitegrid", {'grid.linestyle': '--'})
fig, ax = plt.subplots(2,1, figsize=(17, 5), sharex=True) # plt.grid()
sns.lineplot(x=Rain_Bastia_month.index, y= "Rain_Bastia", data=Rain_Bastia_month, label="Rain_Bastia_month",ax=ax[0] )
#ax[0].axis(xmin=0 ,xmax=15 ) # xlim( 0,)
sns.lineplot(x=Volume_C10_month.index, y= "Volume_C10", data=Volume_C10_month, label="Volume_C10_month",color="coral", ax=ax[1]) # ax=ax
#sns.lineplot(x=dfp.index, y= "RainNO_2", data=dfp, label="Rain NE", ax=ax[2]) #
plt.xticks(rotation=90); #axe.setxaxis(rotation=90)
#plt.xlim(pd.to_datetime( "2010-01-01" ),pd.to_datetime("2019-12-31" ) );
Autocorrelation¶
The change in pump regime is visible in this period: notice the dips: 8, 25, and peaks: 14, 33, 47.
import statsmodels.api as sm #figsize=(10, 3)
sm.graphics.tsa.plot_acf(dfp["2018-01-01":"2020-04-01"].Volume_C10.values, lags=80, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm #figsize=(10, 3)
sm.graphics.tsa.plot_acf(dfp["2015-01-01":"2020-04-01"].Volume_C10.values, lags=80, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm
sm.graphics.tsa.plot_acf(dfp["2010-04-01":"2015-04-01"].Volume_C10.values, lags=80, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm
sm.graphics.tsa.plot_acf(dfp["2010-04-01":"2020-04-01"].Volume_C10.values, lags=80, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm
sm.graphics.tsa.plot_acf(dfp["2010-04-01":"2020-04-01"].Hydrometry.values, lags=360, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm
sm.graphics.tsa.plot_acf(dfp["2010-01-01":"2020-04-01"].head25.values, lags=80, color='indigo',markersize=3) #.squeeze()
plt.show()
import statsmodels.api as sm
sm.graphics.tsa.plot_acf(dfp["2010-01-01":"2020-04-01"].Rain_Bastia.values, lags=360, color='indigo',markersize=3) #.squeeze()
plt.show()
Seasonal decompose¶
from sklearn.feature_selection import RFE
from sklearn.ensemble import ExtraTreesRegressor
from statsmodels.tsa.seasonal import seasonal_decompose
dfpseason = dfp[['Rain_Bastia', 'Depth_to_P24', 'Depth_to_P25',
'Temp_Petrig', 'Volume_C10', 'Hydrometry', 'head24', 'head25', 'Hth2diff',
'Month', 'Rainfall_3', 'Rainfall_7', 'Rainfall_122', 'Rainfall_30',
'TmnStdP', 'TmnStdP_4', 'TP_7', 'Vol_124', 'Vol_22',
'Vol_33', 'Vol_47', 'doy', 'Year', 'Hyd_39', 'Hyd_128']] # 'diff_surface_gwater',
dfpseason['head25nextday'] = dfpseason['head25'].shift(-1)
dfpseason = dfpseason.dropna()
dfpseason = dfpseason.reset_index()
decompose_columns = ['Rain_Bastia', 'Temp_Petrig', 'head25','Volume_C10',"Hydrometry"]
for column in decompose_columns:
decompose_data = seasonal_decompose(dfpseason[column], freq=52, model='additive', extrapolate_trend='freq')# period n days! not weeks
dfpseason[f"{column}_Trend"] = decompose_data.trend
dfpseason[f"{column}_Seasonal"] = decompose_data.seasonal
f, ax = plt.subplots(nrows=12, ncols=1, figsize=(19, 20))
f.tight_layout()
f.subplots_adjust(hspace=0.5)
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia, data=dfpseason, ax=ax[0], color='black')
ax[0].set_title('Rainfall', fontsize=10)
ax[0].set_ylabel(ylabel=' ')
ax[0].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia_Trend, ax=ax[1], color='black')
ax[1].set_title('Rainfall_Trend', fontsize=10)
ax[1].set_ylabel(ylabel=' ')
ax[1].set_xlabel(xlabel=' ');
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia_Seasonal, data=dfpseason, ax=ax[2], color='black')
ax[2].set_title('Rainfall_Seasonal', fontsize=10)
ax[2].set_ylabel(ylabel=' ')
ax[2].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Temp_Petrig, ax=ax[3], color='black')
ax[3].set_title('Temperature_Petrig', fontsize=10)
ax[3].set_ylabel(ylabel=' ')
ax[3].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Temp_Petrig_Trend, ax=ax[4], color='black')
ax[4].set_title('Temperature_Petrig_Trend', fontsize=10)
ax[4].set_ylabel(ylabel=' ')
ax[4].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Temp_Petrig_Seasonal, ax=ax[5], color='black')
ax[5].set_title('Temperature_Petrig_Seasonal', fontsize=10)
ax[5].set_ylabel(ylabel=' ')
ax[5].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry, ax=ax[6], color='black')
ax[6].set_title('Hydrometry', fontsize=10)
ax[6].set_ylabel(ylabel=' ')
ax[6].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry_Trend, ax=ax[7], color='black')
ax[7].set_title('Hydrometry_Trend', fontsize=10)
ax[7].set_ylabel(ylabel=' ')
ax[7].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry_Seasonal, ax=ax[8], color='black')
ax[8].set_title('Hydrometry_Season', fontsize=10)
ax[8].set_ylabel(ylabel=' ')
ax[8].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10, ax=ax[9], color='black')
ax[6].set_title('Volume_C10', fontsize=10)
ax[6].set_ylabel(ylabel=' ')
ax[6].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10_Trend, ax=ax[10], color='black')
ax[7].set_title('Volume_C10y_Trend', fontsize=10)
ax[7].set_ylabel(ylabel=' ')
ax[7].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10_Seasonal, ax=ax[11], color='black')
ax[8].set_title('Volume_C10_Season', fontsize=10)
ax[8].set_ylabel(ylabel=' ')
ax[8].set_xlabel(xlabel=' ')
plt.show()
period of 180 days¶
from statsmodels.tsa.seasonal import seasonal_decompose
dfpseason = dfp[['Rain_Bastia', 'Depth_to_P24', 'Depth_to_P25',
'Temp_Petrig', 'Volume_C10', 'Hydrometry', 'head24', 'head25', 'Hth2diff',
'Month', 'Rainfall_3', 'Rainfall_7', 'Rainfall_122', 'Rainfall_30',
'TmnStdP', 'TmnStdP_4', 'TP_7', 'Vol_124', 'Vol_22',
'Vol_33', 'Vol_47', 'doy', 'Year', 'Hyd_39', 'Hyd_128']] #'diff_surface_gwater',
dfpseason['head25nextday'] = dfpseason['head25'].shift(-1)
dfpseason = dfpseason.dropna()
dfpseason = dfpseason.reset_index()
decompose_columns = ['Rain_Bastia', 'Temp_Petrig', 'head25','Volume_C10',"Hydrometry"]
for column in decompose_columns:
decompose_data = seasonal_decompose(dfpseason[column], period=180, model='additive', extrapolate_trend='freq')# period n days! not weeks
dfpseason[f"{column}_Trend"] = decompose_data.trend
dfpseason[f"{column}_Seasonal"] = decompose_data.seasonal
f, ax = plt.subplots(nrows=12, ncols=1, figsize=(19, 20))
f.tight_layout()
f.subplots_adjust(hspace=0.5)
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia, data=dfpseason, ax=ax[0], color='black')
ax[0].set_title('Rainfall', fontsize=10)
ax[0].set_ylabel(ylabel=' ')
ax[0].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia_Trend, ax=ax[1], color='black')
ax[1].set_title('Rainfall_Trend', fontsize=10)
ax[1].set_ylabel(ylabel=' ')
ax[1].set_xlabel(xlabel=' ');
sns.lineplot(x=dfpseason.Date, y=dfpseason.Rain_Bastia_Seasonal, data=dfpseason, ax=ax[2], color='black')
ax[2].set_title('Rainfall_Seasonal', fontsize=10)
ax[2].set_ylabel(ylabel=' ')
ax[2].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.head25, ax=ax[3], color='black')
ax[3].set_title('head25', fontsize=10)
ax[3].set_ylabel(ylabel=' ')
ax[3].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.head25_Trend, ax=ax[4], color='black')
ax[4].set_title('head25_Trend', fontsize=10)
ax[4].set_ylabel(ylabel=' ')
ax[4].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.head25_Seasonal, ax=ax[5], color='black')
ax[5].set_title('head25_Seasonal', fontsize=10)
ax[5].set_ylabel(ylabel=' ')
ax[5].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry, ax=ax[6], color='black')
ax[6].set_title('Hydrometry', fontsize=10)
ax[6].set_ylabel(ylabel=' ')
ax[6].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry_Trend, ax=ax[7], color='black')
ax[7].set_title('Hydrometry_Trend', fontsize=10)
ax[7].set_ylabel(ylabel=' ')
ax[7].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Hydrometry_Seasonal, ax=ax[8], color='black')
ax[8].set_title('Hydrometry_Seasonal', fontsize=10)
ax[8].set_ylabel(ylabel=' ')
ax[8].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10, ax=ax[9], color='black')
ax[6].set_title('Volume_C10', fontsize=10)
ax[6].set_ylabel(ylabel=' ')
ax[6].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10_Trend, ax=ax[10], color='black')
ax[7].set_title('Volume_C10y_Trend', fontsize=10)
ax[7].set_ylabel(ylabel=' ')
ax[7].set_xlabel(xlabel=' ')
sns.lineplot(x=dfpseason.Date, y=dfpseason.Volume_C10_Seasonal, ax=ax[11], color='black')
ax[8].set_title('Volume_C10_Seasonal', fontsize=10)
ax[8].set_ylabel(ylabel=' ')
ax[8].set_xlabel(xlabel=' ')
plt.show()
<ipython-input-55-a263334b80b4>:15: FutureWarning: the 'freq'' keyword is deprecated, use 'period' instead decompose_data = seasonal_decompose(dfpseason[column], freq=180, model='additive', extrapolate_trend='freq')# period n days! not weeks
Hydrological models¶
Pastas seems a nice framework for hydrological models, but it is necessary to use the correct unit for certain features. Also finding some usefull evaporation data for Italy was an obstacle.
!pip install pastas
import pastas as ps
###### Import groundwater time series and squeeze to Series object
gwdata = dfp.head25 #pd.read_csv('Aquifer_Petrignano.csv', parse_dates=['date'], index_col='date', squeeze=True)
print('The data type of the oseries is: %s' % type(gwdata))
###### Plot the observed groundwater levels
gwdata.plot(style='.', figsize=(17, 4), markersize=2.4)
plt.ylabel('Head [m]');
plt.xlabel('Time [years]');
The data type of the oseries is: <class 'pandas.core.series.Series'>
# Import observed precipitation series
#precip = dfp.Rain_Bastia #Rain_Prepignano #pd.read_csv('../data/rain_nb1.csv', parse_dates=['date'], index_col='date', squeeze=True)
#print('The data type of the precip series is: %s' % type(precip))
# Import observed evaporation series
#evap = pd.read_csv(r'../input/nb18_evap.csv', parse_dates=['date'], index_col='date', squeeze=True)
#evap = evap.loc["2008":"2021"]
#print('The data type of the evap series is: %s' % type(evap))
# Calculate the recharge to the groundwater
#recharge = precip - evap
# Plot the time series of the precipitation and evaporation
#plt.figure()
#recharge.plot(label='Recharge', figsize=(17, 4))
#plt.xlabel('Time [years]')
#plt.ylabel('Recharge (m/year)');
Create a time series model¶
In this code block the actual time series model is created. First, an instance of the Model class is created (named ml here). Second, the different components of the time series model are created and added to the model. The imported time series are automatically checked for missing values and other inconsistencies. The keyword argument fillnan can be used to determine how missing values are handled. If any nan-values are found this will be reported by pastas.
Here we remove most of the 0's from the volume series. Perhaps it is better to remove them all.
from pandas import Timestamp
strikedaystodump =[Timestamp('2011-06-10 00:00:00'), Timestamp('2019-06-29 00:00:00'),
Timestamp('2019-06-30 00:00:00'), Timestamp('2019-07-01 00:00:00'),
Timestamp('2019-07-02 00:00:00'), #Timestamp('2019-07-03 00:00:00'),
Timestamp('2019-07-04 00:00:00'), Timestamp('2019-07-05 00:00:00'),
Timestamp('2019-07-14 00:00:00'), Timestamp('2019-07-21 00:00:00'),
Timestamp('2019-08-01 00:00:00'), Timestamp('2019-08-05 00:00:00'),
Timestamp('2019-08-14 00:00:00'), Timestamp('2019-08-16 00:00:00'),
Timestamp('2019-08-19 00:00:00'), Timestamp('2019-08-21 00:00:00')]
dfp=dfp[ ~dfp.index.isin(strikedaystodump)]
Single target regression for Petrignano¶
First I'll try the RandomForestRegressor, to predict 1 of the 2 headings/cone heights based on the pure - or some transformation(s)- of the other heading/cone height.
I should find a factor which brings in the weight of rainfall (and evaporation), river level and pump discharge, based on the waterbalance...
after that ... Keras or XGBoost.
There are some days when the volume is 0, which is caused by human intervention I believe. These records are unpure, and distort the series, so I'll only keep the first day, and the rest will be dropped.
# Just take dates with decent data...train the model, and predict the 2020 values
dfp08= dfp["2009-01-01":"2020-06-14"]
dfp08.replace([np.inf, -np.inf], np.nan, inplace=True)
dfp08= dfp08.dropna() # inplace=True
Depth_to_P24= dfp08.loc[:,["head25"]]# "CHt_P25"
Petri= dfp08.drop(columns= ["Hth_to_P24","Hth_to_P25","Depth_to_P24","Depth_to_P25","CHt_P24","CHt_P25","head24","head25"], axis=1)
Petri.head() # ,"depthdiff","depth2diff","Hthdiff", "Hth2diff","T_Delta" #
| Rain_Bastia | Temp_Bastia | Temp_Petrig | Volume_C10 | Hydrometry | Debiet | Hth2diff | Hth2diff_S1 | Hth2diff_S11 | Hth2diff_S21 | ... | TP_7 | Vol_124 | Vol_22 | Vol_33 | Vol_47 | doy | Year | Week | Hyd_39 | Hyd_128 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | |||||||||||||||||||||
| 2009-01-01 | 0.0 | 5.2 | 4.9 | 24530.688 | 2.4 | 0.28392 | 0.021735 | 0.153637 | 0.167255 | 0.003378 | ... | 1.285714 | 4085576.064 | 626293.728 | 861848.640 | 1404765.504 | 1 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-02 | 0.0 | 2.3 | 2.5 | 28785.888 | 2.5 | 0.33317 | 0.021625 | 0.135752 | 0.141561 | 0.002117 | ... | 1.285714 | 4080887.136 | 627008.256 | 861249.888 | 1402965.792 | 2 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-03 | 0.0 | 4.4 | 3.9 | 25766.208 | 2.4 | 0.29822 | 0.021607 | 0.148980 | 0.162234 | 0.003345 | ... | 1.285714 | 4070869.920 | 621679.104 | 857704.032 | 1395009.216 | 3 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-04 | 0.0 | 0.8 | 0.8 | 27919.296 | 2.4 | 0.32314 | 0.021496 | 0.140069 | 0.147994 | 0.002985 | ... | 1.285714 | 4063772.160 | 613121.184 | 857570.976 | 1392793.056 | 4 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-05 | 0.0 | -1.9 | -2.1 | 29854.656 | 2.3 | 0.34554 | 0.021474 | 0.132703 | 0.139222 | 0.002730 | ... | 1.285714 | 4057692.192 | 610816.032 | 855811.872 | 1391827.968 | 5 | 2009 | 2 | 94.9 | 312.5 |
5 rows × 28 columns
print(Petri.shape, Depth_to_P24.shape )
(4168, 28) (4168, 1)
Petri.iloc[:,0:15].head(10)
| Rain_Bastia | Temp_Bastia | Temp_Petrig | Volume_C10 | Hydrometry | Debiet | Hth2diff | Hth2diff_S1 | Hth2diff_S11 | Hth2diff_S21 | Month | Rainfall_3 | Rainfall_7 | Rainfall_122 | Rainfall_30 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | |||||||||||||||
| 2009-01-01 | 0.0 | 5.2 | 4.9 | 24530.688 | 2.4 | 0.28392 | 0.021735 | 0.153637 | 0.167255 | 0.003378 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-02 | 0.0 | 2.3 | 2.5 | 28785.888 | 2.5 | 0.33317 | 0.021625 | 0.135752 | 0.141561 | 0.002117 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-03 | 0.0 | 4.4 | 3.9 | 25766.208 | 2.4 | 0.29822 | 0.021607 | 0.148980 | 0.162234 | 0.003345 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-04 | 0.0 | 0.8 | 0.8 | 27919.296 | 2.4 | 0.32314 | 0.021496 | 0.140069 | 0.147994 | 0.002985 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-05 | 0.0 | -1.9 | -2.1 | 29854.656 | 2.3 | 0.34554 | 0.021474 | 0.132703 | 0.139222 | 0.002730 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-06 | 0.0 | -0.7 | -0.7 | 29124.576 | 2.3 | 0.33709 | 0.021617 | 0.136174 | 0.144477 | 0.002964 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-07 | 0.0 | 1.5 | -0.3 | 31173.120 | 2.3 | 0.36080 | 0.021595 | 0.128451 | 0.135785 | 0.002741 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-08 | 0.0 | 4.3 | 6.6 | 30232.224 | 2.4 | 0.34991 | 0.021233 | 0.137938 | 0.140831 | 0.002922 | 1 | 0.0 | 0.0 | 70.8 | 6.3 |
| 2009-01-09 | 0.9 | 4.9 | 4.8 | 30597.696 | 2.3 | 0.35414 | 0.020892 | 0.135710 | 0.140771 | 0.002764 | 1 | 0.9 | 0.9 | 70.8 | 6.3 |
| 2009-01-10 | 0.0 | 1.9 | 4.2 | 31337.280 | 2.3 | 0.36270 | 0.020318 | 0.131932 | 0.136447 | 0.002669 | 1 | 0.9 | 0.9 | 70.8 | 6.3 |
we drop some features that proved not to progress the model after several train and test runs. at random several combinations were used.
Petri= Petri.drop(columns=["Temp_Bastia","TmnStdB",'Rain_Bastia','Temp_Petrig','Volume_C10','Hth2diff_S1','Hth2diff_S11','Hth2diff_S21',#'Rainfall_3','Rainfall_30', 'Volume_C10',
'TmnStdP', "TmnStdP_4",'Hydrometry','Debiet','Rainfall_3','Rainfall_7',"Rainfall_30", 'TmnStdP'], axis=1) #,'HydStd_8' "Hthdiff","Year", ,'Hth2diff'
X= Petri
y= Depth_to_P24
print(X.shape,y.shape )
(4168, 14) (4168, 1)
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.model_selection import TimeSeriesSplit
from sklearn.ensemble import RandomForestRegressor
#from sklearn.ensemble import ExtraTreesRegressor
from sklearn.feature_selection import SelectFromModel
from sklearn.metrics import accuracy_score
X_train,X_test, y_train,y_test= train_test_split(X ,y, test_size=0.105,random_state=1108,shuffle=False,)# ,n_splits=9.0max_train_size=365 random_state=1100,shuffle=Fals
X_train = X_train.values.reshape(-1,1) X_test = X_test.values.reshape(-1,1)
print(X_train.shape, y_train.shape, X_test.shape,y_test.shape)
(3730, 14) (3730, 1) (438, 14) (438, 1)
col_mask= X_train.isnull().any(axis=0); row_mask= X_train.isnull().any(axis=1); X_train.loc[row_mask,col_mask]
from Random Forest Regressor to Extra Trees Regressor¶
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import ExtraTreesRegressor
#from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
#GBr = GradientBoostingRegressor(random_state=100)
RFr = ExtraTreesRegressor(random_state=1100, n_jobs=3, n_estimators=12300,bootstrap=True,verbose=1,criterion="mse",max_features=None,min_samples_leaf=1)# log
type(X_train)
RFr.fit(X_train, y_train.values.ravel())
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 0.5s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 1.1s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 2.1s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 3.4s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 5.1s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 7.1s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 9.7s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 12.8s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 16.3s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 21.1s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 27.1s [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 33.8s [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 41.3s [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 50.3s [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 57.5s finished
ExtraTreesRegressor(bootstrap=True, max_features=None, n_estimators=12300,
n_jobs=3, random_state=1100, verbose=1)
RFr.feature_importances_
importances = RFr.feature_importances_
std = np.std([tree.feature_importances_ for tree in RFr.estimators_],
axis=0)
indices = np.argsort(importances)[::-1]
print("Feature ranking:")
for f in range(X.shape[1]):
print("%d. feature %d (%f)" % (f + 1, indices[f], importances[indices[f]]))
# Plot the feature importances of the forest
plt.figure()
plt.title("Feature importances")
plt.bar(range(X.shape[1]), importances[indices], color="crimson", yerr=std[indices], align="center")
plt.xticks(range(X.shape[1]), indices)
plt.xlim([-1, X.shape[1]])
plt.show()
Feature ranking: 1. feature 10 (0.320956) 2. feature 5 (0.194281) 3. feature 2 (0.152908) 4. feature 13 (0.097149) 5. feature 0 (0.046106) 6. feature 8 (0.045931) 7. feature 1 (0.025373) 8. feature 9 (0.023635) 9. feature 11 (0.021823) 10. feature 7 (0.020781) 11. feature 12 (0.016469) 12. feature 6 (0.013279) 13. feature 4 (0.012169) 14. feature 3 (0.009139)
Petricolumns= list(Petri.columns); Petricols=enumerate(Petricolumns); print (list(Petricols))
[(0, 'Hth2diff'), (1, 'Month'), (2, 'Rainfall_122'), (3, 'TmnStdP_4'), (4, 'TP_7'), (5, 'Vol_124'), (6, 'Vol_22'), (7, 'Vol_33'), (8, 'Vol_47'), (9, 'doy'), (10, 'Year'), (11, 'Week'), (12, 'Hyd_39'), (13, 'Hyd_128')]
Return the coefficient of determination 𝑅2 of the prediction.¶
RFr.score(X_test, y_test) # max_features=2, 0.7448
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 0.2s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 0.3s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 0.4s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 0.5s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 0.6s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 0.7s [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 0.9s [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 1.0s [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 1.2s [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 1.3s finished
-3.764810621914388
y_pred = RFr.predict(X_test)
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 0.2s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 0.3s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 0.4s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 0.5s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 0.6s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 0.7s [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 0.9s [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 1.0s [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 1.2s [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 1.3s finished
y_test = y_test.values.ravel() #values.reshape(-1,1)
print(y_test[0:20], type(y_test))
[174.295 174.30875 174.3225 174.33625 174.35 174.32 174.3 174.12 174.23 174.26 174.34 174.34 174.37 174.39 174.55 174.47 174.47 174.51 174.46 174.47 ] <class 'numpy.ndarray'>
RF regressor metrics¶
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
mape = np.mean(np.abs((y_test - y_pred) / np.abs(y_test)))
print('Mean Absolute Percentage Error (MAPE):', round(mape * 100, 2))
print('Accuracy:', round(100*(1 - mape), 2))
Mean Absolute Error: 1.1720349115530553 Mean Squared Error: 2.097612984802689 Root Mean Squared Error: 1.4483138419564625 Mean Absolute Percentage Error (MAPE): 0.67 Accuracy: 99.33
RF regressor predictions vs. observations¶
y_test[0:10]
dataset = pd.DataFrame()
dataset['y_test'] = y_test.tolist()#
dataset['y_pred'] = y_pred.tolist()
dataset
| y_test | y_pred | |
|---|---|---|
| 0 | 174.29500 | 174.290078 |
| 1 | 174.30875 | 174.298609 |
| 2 | 174.32250 | 174.346016 |
| 3 | 174.33625 | 174.414073 |
| 4 | 174.35000 | 174.484556 |
| ... | ... | ... |
| 433 | 175.33000 | 175.737674 |
| 434 | 175.24000 | 175.723419 |
| 435 | 175.19000 | 175.718603 |
| 436 | 175.21000 | 175.718640 |
| 437 | 175.29000 | 175.715753 |
438 rows × 2 columns
import seaborn as sns
sns.set_theme(style="darkgrid")
fig,ax= plt.subplots(1,1, figsize=(14,10),) #
sns.scatterplot(x=dataset.y_test, y=dataset.y_pred , data= dataset, s=4, color="forestgreen")#
#sns.scatterplot(x=y_train0, y=y_train0, color="orange", ax=ax);#data=y_train,
ax.minorticks_on()
ax.xaxis.grid(True, "minor", linewidth=.25)
ax.yaxis.grid(True, "minor", linewidth=.25);
!pip install pdpbox
from pdpbox.pdp import pdp_isolate, pdp_plot
feature= "Hyd_128" #"Temp_Bastia"
isolated= pdp_isolate( model=RFr ,dataset=Petri, model_features=Petri.columns, feature= feature) #
Multitarget regression for Petrignano¶
First I'll try the Extra Trees Regressor, after that Keras or MultiOutputRegressor.
#df10.dropna(inplace=True)
#dfp["Month"]= dfp.index.month
#dfp["Year"]= dfp.index.year
dfp.columns
Index(['Rain_Bastia', 'Depth_to_P24', 'Depth_to_P25', 'Temp_Bastia',
'Temp_Petrig', 'Volume_C10', 'Hydrometry', 'Debiet', 'Hth_to_P24',
'Hth_to_P25', 'CHt_P24', 'CHt_P25', 'head24', 'head25', 'Hth2diff',
'Hth2diff_S1', 'Hth2diff_S11', 'Hth2diff_S21', 'Month', 'Rainfall_3',
'Rainfall_7', 'Rainfall_122', 'Rainfall_30', 'TmnStdB', 'TmnStdP',
'TmnStdP_4', 'TP_7', 'Vol_124', 'Vol_22', 'Vol_33', 'Vol_47', 'doy',
'Year', 'Week', 'Hyd_39', 'Hyd_128'],
dtype='object')
# Just take data with decent data...
dfp0920= dfp["2009-01-01":"2020-06-24"]
dfp0920.dropna(inplace=True)
#
Hth_to_P25= dfp0920.loc[:,["head24","head25"]]#
Petri= dfp0920.drop(columns= ["Hth_to_P24",'Depth_to_P24',"Hth_to_P25","Depth_to_P25","CHt_P24","CHt_P25","head24","head25",'Hth2diff_S1','Hth2diff_S11','Hth2diff_S21',
'Temp_Bastia', 'Debiet','TmnStdB', 'TmnStdP_4','Temp_Petrig', 'Volume_C10'], axis=1) # ,'Hth2diff'
Petri.head() # ,"depthdiff","depth2diff","Hthdiff", "Hth2diff","T_Delta"
| Rain_Bastia | Hydrometry | Hth2diff | Month | Rainfall_3 | Rainfall_7 | Rainfall_122 | Rainfall_30 | TmnStdP | TP_7 | Vol_124 | Vol_22 | Vol_33 | Vol_47 | doy | Year | Week | Hyd_39 | Hyd_128 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Date | |||||||||||||||||||
| 2009-01-01 | 0.0 | 2.4 | 0.021735 | 1 | 0.0 | 0.0 | 70.8 | 6.3 | -2.557051 | 1.285714 | 4085576.064 | 626293.728 | 861848.640 | 1404765.504 | 1 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-02 | 0.0 | 2.5 | 0.021625 | 1 | 0.0 | 0.0 | 70.8 | 6.3 | -4.957051 | 1.285714 | 4080887.136 | 627008.256 | 861249.888 | 1402965.792 | 2 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-03 | 0.0 | 2.4 | 0.021607 | 1 | 0.0 | 0.0 | 70.8 | 6.3 | -3.557051 | 1.285714 | 4070869.920 | 621679.104 | 857704.032 | 1395009.216 | 3 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-04 | 0.0 | 2.4 | 0.021496 | 1 | 0.0 | 0.0 | 70.8 | 6.3 | -6.657051 | 1.285714 | 4063772.160 | 613121.184 | 857570.976 | 1392793.056 | 4 | 2009 | 1 | 94.9 | 312.5 |
| 2009-01-05 | 0.0 | 2.3 | 0.021474 | 1 | 0.0 | 0.0 | 70.8 | 6.3 | -9.557051 | 1.285714 | 4057692.192 | 610816.032 | 855811.872 | 1391827.968 | 5 | 2009 | 2 | 94.9 | 312.5 |
print(Petri.shape, Hth_to_P25.shape )
(4177, 19) (4177, 2)
Petri= Petri.drop(columns=['Rain_Bastia','Hydrometry',"Rainfall_7",'Rainfall_3',"Rainfall_30", 'TmnStdP'], axis=1) # ,"Year",'Temp_Petrig'"Temp_Bastia""TmnStdB",
#,"Rainfall_30" 'TmnStdP',"Debiet",'Volume_C10',"Temp_Petrig"
X= Petri
y= Hth_to_P25
print(X.shape,y.shape )
(4177, 13) (4177, 2)
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.model_selection import TimeSeriesSplit
from sklearn.ensemble import RandomForestRegressor
#from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_selection import SelectFromModel
from sklearn.metrics import accuracy_score
X_train,X_test, y_train,y_test= train_test_split(X ,y, test_size=0.105,random_state=1100,shuffle=False,)# ,n_splits=9.0max_train_size=365 random_state=1100,shuffle=False,
y.tail(10) # TimeSeriesSplit might not support 2 targetvals
| head24 | head25 | |
|---|---|---|
| Date | ||
| 2020-06-14 | 175.01 | 175.29 |
| 2020-06-16 | 174.67 | 175.23 |
| 2020-06-17 | 174.96 | 175.27 |
| 2020-06-18 | 175.01 | 175.28 |
| 2020-06-19 | 174.81 | 175.26 |
| 2020-06-20 | 174.59 | 175.20 |
| 2020-06-21 | 174.64 | 175.18 |
| 2020-06-22 | 174.66 | 175.17 |
| 2020-06-23 | 174.58 | 175.12 |
| 2020-06-24 | 174.33 | 175.03 |
print(X_train.shape, y_train.shape, X_test.shape,y_test.shape)
(3738, 13) (3738, 2) (439, 13) (439, 2)
Extra Trees Regressor¶
from sklearn.ensemble import GradientBoostingRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.model_selection import cross_val_score
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
#GBr = GradientBoostingRegressor(random_state=100)
RFr2 = ExtraTreesRegressor(random_state=1100, n_jobs=3, n_estimators=12300,bootstrap=False,verbose=1,criterion="mse",max_features="auto")# log
type(X_train)
RFr2.fit(X_train, y_train.values) #.ravel().ravel()
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.3s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 1.6s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 3.3s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 6.1s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 10.0s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 14.2s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 20.9s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 28.9s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 37.6s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 46.4s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 56.3s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 1.2min [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 1.4min [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 1.6min [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 1.9min [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 2.2min finished
ExtraTreesRegressor(n_estimators=12300, n_jobs=3, random_state=1100, verbose=1)
importances = RFr2.feature_importances_
std = np.std([tree.feature_importances_ for tree in RFr2.estimators_],
axis=0)
indices = np.argsort(importances)[::-1]
print("Feature ranking:")
for f in range(X.shape[1]):
print("%d. feature %d (%f)" % (f + 1, indices[f], importances[indices[f]]))
# Plot the feature importances of the forest
plt.figure()
plt.title("Feature importances")
plt.bar(range(X.shape[1]), importances[indices], color="crimson", yerr=std[indices], align="center")
plt.xticks(range(X.shape[1]), indices)
plt.xlim([-1, X.shape[1]])
plt.show()
Feature ranking: 1. feature 9 (0.322201) 2. feature 4 (0.201433) 3. feature 2 (0.152696) 4. feature 12 (0.098458) 5. feature 7 (0.046320) 6. feature 0 (0.039132) 7. feature 1 (0.026542) 8. feature 8 (0.024843) 9. feature 10 (0.022674) 10. feature 6 (0.021226) 11. feature 11 (0.017064) 12. feature 5 (0.013731) 13. feature 3 (0.013681)
Petricolumns= list(Petri.columns); Petricols=enumerate(Petricolumns); print (list(Petricols))
[(0, 'Hth2diff'), (1, 'Month'), (2, 'Rainfall_122'), (3, 'TP_7'), (4, 'Vol_124'), (5, 'Vol_22'), (6, 'Vol_33'), (7, 'Vol_47'), (8, 'doy'), (9, 'Year'), (10, 'Week'), (11, 'Hyd_39'), (12, 'Hyd_128')]
RFr2
Return the coefficient of determination 𝑅2 of the prediction.¶
RFr2.score(X_test, y_test) #
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 0.5s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 0.8s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 1.2s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 1.7s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 2.1s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 2.5s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 3.8s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 3.9s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 4.1s [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 4.2s [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 4.4s [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 4.5s [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 4.6s finished
-2.8923907082780835
y_pred = RFr2.predict(X_test)
[Parallel(n_jobs=3)]: Using backend ThreadingBackend with 3 concurrent workers. [Parallel(n_jobs=3)]: Done 44 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 194 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 444 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 794 tasks | elapsed: 0.0s [Parallel(n_jobs=3)]: Done 1244 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 1794 tasks | elapsed: 0.1s [Parallel(n_jobs=3)]: Done 2444 tasks | elapsed: 0.2s [Parallel(n_jobs=3)]: Done 3194 tasks | elapsed: 0.3s [Parallel(n_jobs=3)]: Done 4044 tasks | elapsed: 0.4s [Parallel(n_jobs=3)]: Done 4994 tasks | elapsed: 0.5s [Parallel(n_jobs=3)]: Done 6044 tasks | elapsed: 0.6s [Parallel(n_jobs=3)]: Done 7194 tasks | elapsed: 0.8s [Parallel(n_jobs=3)]: Done 8444 tasks | elapsed: 0.9s [Parallel(n_jobs=3)]: Done 9794 tasks | elapsed: 1.1s [Parallel(n_jobs=3)]: Done 11244 tasks | elapsed: 1.3s [Parallel(n_jobs=3)]: Done 12300 out of 12300 | elapsed: 1.4s finished
y_test
y_pred[0:10]
RF regressor metrics¶
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
mape = np.mean(np.abs((y_test - y_pred) / np.abs(y_test)))
print('Mean Absolute Percentage Error (MAPE):', round(mape * 100, 2))
print('Accuracy:', round(100*(1 - mape), 2))
Mean Absolute Error: 1.1243007123875501 Mean Squared Error: 1.998062809540579 Root Mean Squared Error: 1.4135284961897936 Mean Absolute Percentage Error (MAPE): head24 0.64 head25 0.65 dtype: float64 Accuracy: head24 99.36 head25 99.35 dtype: float64
RF regressor predictions vs. observations¶
dataset = pd.DataFrame() dataset['y_test'] = y_test.tolist()#.reshape(-1,1 ) dataset['y_pred'] = y_pred.tolist() dataset
y_test
Y_pred = pd.DataFrame({'head_24': y_pred[:, 0], 'head_25': y_pred[:, 1]} )
fig, ax = plt.subplots(figsize=(19,5))
sns.lineplot(x= y_test.index,y= y_test.head24, data=y_test, label="y_test.head24") #CHt_P25 Hth_to_P25
sns.lineplot(x= y_test.index ,y= Y_pred.head_24, data=Y_pred, label="Y_pred.head_24")
sns.lineplot(x= y_test.index,y= y_test.head25, data=y_test, label="y_test.head25")
sns.lineplot(x= y_test.index ,y= Y_pred.head_25 , data=Y_pred, label="Y_pred.head_25");
Keras¶
import math
from tensorflow import keras
from keras.models import Sequential
from keras.layers import Dense, Input
from keras.models import Model
from keras.layers import LSTM
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
df = df.drop('Date', axis=1)
scaler = MinMaxScaler(feature_range = (0, 1))
df_norm = scaler.fit_transform(df.values)
df = pd.DataFrame(df_norm, index=df.index, columns=df.columns)