Decision tree for classification in plain Python¶
A decision tree is a supervised machine learning model that can be used both for classification and regression. At its core, a decision tree uses a tree structure to predict an output value for a given input example. In the tree, each path from the root node to a leaf node represents a decision path that ends in a predicted value.
A simple example might look as follows:
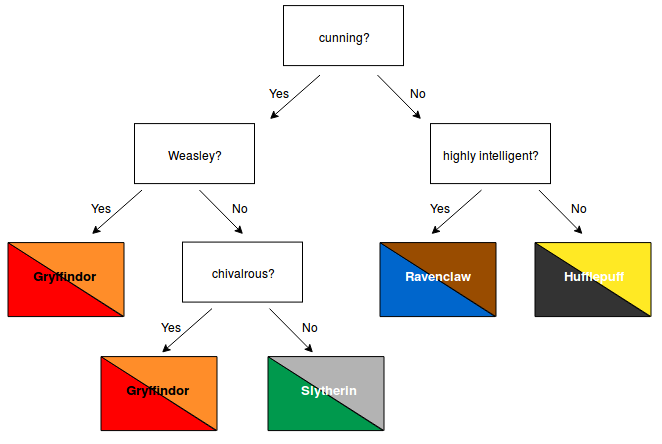
Decision trees have many advantages. For example, they are easy to understand and their decisions are easy to interpret. Also, they don't require a lot of data preparation. A more extensive list of their advantages and disadvantages can be found here.
CART training algorithm¶
In order to train a decision tree, various algorithms can be used. In this notebook we will focus on the CART algorithm (Classification and Regression Trees) for classification. The CART algorithm builds a binary tree in which every non-leaf node has exactly two children (corresponding to a yes/no answer).
Given a set of training examples and their labels, the algorithm repeatedly splits the training examples $D$ into two subsets $D_{left}, D_{right}$ using some feature $f$ and feature threshold $t_f$ such that samples with the same label are grouped together. At each node, the algorithm selects the split $\theta = (f, t_f)$ that produces the purest subsets, weighted by their size. Purity/impurity is measured using the Gini impurity.
So at each step, the algorithm selects the parameters $\theta$ that minimize the following cost function:
\begin{equation} J(D, \theta) = \frac{n_{left}}{n_{total}} G_{left} + \frac{n_{right}}{n_{total}} G_{right} \end{equation}- $D$: remaining training examples
- $n_{total}$ : number of remaining training examples
- $\theta = (f, t_f)$: feature and feature threshold
- $n_{left}/n_{right}$: number of samples in the left/right subset
- $G_{left}/G_{right}$: Gini impurity of the left/right subset
This step is repeated recursively until the maximum allowable depth is reached or the current number of samples $n_{total}$ drops below some minimum number. The original equations can be found here.
After building the tree, new examples can be classified by navigating through the tree, testing at each node the corresponding feature until a leaf node/prediction is reached.
Gini Impurity¶
Given $K$ different classification values $k \in \{1, ..., K\}$ the Gini impurity of node $m$ is computed as follows:
\begin{equation} G_m = 1 - \sum_{k=1}^{K} (p_{m,k})^2 \end{equation}where $p_{m, k}$ is the fraction of training examples with class $k$ among all training examples in node $m$.
The Gini impurity can be used to evaluate how good a potential split is. A split divides a given set of training examples into two groups. Gini measures how "mixed" the resulting groups are. A perfect separation (i.e. each group contains only samples of the same class) corresponds to a Gini impurity of 0. If the resulting groups contain equally many samples of each class, the Gini impurity will reach its highest value of 0.5
Caveats¶
Without regularization, decision trees are likely to overfit the training examples. This can be prevented using techniques like pruning or by providing a maximum allowed tree depth and/or a minimum number of samples required to split a node further.
from sklearn.datasets import load_iris
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
np.random.seed(123)
%matplotlib inline
Dataset¶
The iris dataset compromises 150 examples of 3 different species of iris flowers (Setosa, Versicolour, and Virginica). Each example is described by four attributes: sepal length (cm), sepal width (cm), petal length (cm), petal width (cm).
iris = load_iris()
X, y = iris.data, iris.target
# Split the data into a training and test set
X_train, X_test, y_train, y_test = train_test_split(X, y)
print(f'Shape X_train: {X_train.shape}')
print(f'Shape y_train: {y_train.shape}')
print(f'Shape X_test: {X_test.shape}')
print(f'Shape y_test: {y_test.shape}')
Shape X_train: (112, 4) Shape y_train: (112,) Shape X_test: (38, 4) Shape y_test: (38,)
Decision tree class¶
Parts of this code were inspired by this tutorial
class DecisionTree:
"""
Decision tree for classification
"""
def __init__(self):
self.root_dict = None
self.tree_dict = None
def split_dataset(self, X, y, feature_idx, threshold):
"""
Splits dataset X into two subsets, according to a given feature
and feature threshold.
Args:
X: 2D numpy array with data samples
y: 1D numpy array with labels
feature_idx: int, index of feature used for splitting the data
threshold: float, threshold used for splitting the data
Returns:
splits: dict containing the left and right subsets
and their labels
"""
left_idx = np.where(X[:, feature_idx] < threshold)
right_idx = np.where(X[:, feature_idx] >= threshold)
left_subset = X[left_idx]
y_left = y[left_idx]
right_subset = X[right_idx]
y_right = y[right_idx]
splits = {
'left': left_subset,
'y_left': y_left,
'right': right_subset,
'y_right': y_right,
}
return splits
def gini_impurity(self, y_left, y_right, n_left, n_right):
"""
Computes Gini impurity of a split.
Args:
y_left, y_right: target values of samples in left/right subset
n_left, n_right: number of samples in left/right subset
Returns:
gini_left: float, Gini impurity of left subset
gini_right: gloat, Gini impurity of right subset
"""
n_total = n_left + n_left
score_left, score_right = 0, 0
gini_left, gini_right = 0, 0
if n_left != 0:
for c in range(self.n_classes):
# For each class c, compute fraction of samples with class c
p_left = len(np.where(y_left == c)[0]) / n_left
score_left += p_left * p_left
gini_left = 1 - score_left
if n_right != 0:
for c in range(self.n_classes):
p_right = len(np.where(y_right == c)[0]) / n_right
score_right += p_right * p_right
gini_right = 1 - score_right
return gini_left, gini_right
def get_cost(self, splits):
"""
Computes cost of a split given the Gini impurity of
the left and right subset and the sizes of the subsets
Args:
splits: dict, containing params of current split
"""
y_left = splits['y_left']
y_right = splits['y_right']
n_left = len(y_left)
n_right = len(y_right)
n_total = n_left + n_right
gini_left, gini_right = self.gini_impurity(y_left, y_right, n_left, n_right)
cost = (n_left / n_total) * gini_left + (n_right / n_total) * gini_right
return cost
def find_best_split(self, X, y):
"""
Finds the best feature and feature index to split dataset X into
two groups. Checks every value of every attribute as a candidate
split.
Args:
X: 2D numpy array with data samples
y: 1D numpy array with labels
Returns:
best_split_params: dict, containing parameters of the best split
"""
n_samples, n_features = X.shape
best_feature_idx, best_threshold, best_cost, best_splits = np.inf, np.inf, np.inf, None
for feature_idx in range(n_features):
for i in range(n_samples):
current_sample = X[i]
threshold = current_sample[feature_idx]
splits = self.split_dataset(X, y, feature_idx, threshold)
cost = self.get_cost(splits)
if cost < best_cost:
best_feature_idx = feature_idx
best_threshold = threshold
best_cost = cost
best_splits = splits
best_split_params = {
'feature_idx': best_feature_idx,
'threshold': best_threshold,
'cost': best_cost,
'left': best_splits['left'],
'y_left': best_splits['y_left'],
'right': best_splits['right'],
'y_right': best_splits['y_right'],
}
return best_split_params
def build_tree(self, node_dict, depth, max_depth, min_samples):
"""
Builds the decision tree in a recursive fashion.
Args:
node_dict: dict, representing the current node
depth: int, depth of current node in the tree
max_depth: int, maximum allowed tree depth
min_samples: int, minimum number of samples needed to split a node further
Returns:
node_dict: dict, representing the full subtree originating from current node
"""
left_samples = node_dict['left']
right_samples = node_dict['right']
y_left_samples = node_dict['y_left']
y_right_samples = node_dict['y_right']
if len(y_left_samples) == 0 or len(y_right_samples) == 0:
node_dict["left_child"] = node_dict["right_child"] = self.create_terminal_node(np.append(y_left_samples, y_right_samples))
return None
if depth >= max_depth:
node_dict["left_child"] = self.create_terminal_node(y_left_samples)
node_dict["right_child"] = self.create_terminal_node(y_right_samples)
return None
if len(right_samples) < min_samples:
node_dict["right_child"] = self.create_terminal_node(y_right_samples)
else:
node_dict["right_child"] = self.find_best_split(right_samples, y_right_samples)
self.build_tree(node_dict["right_child"], depth+1, max_depth, min_samples)
if len(left_samples) < min_samples:
node_dict["left_child"] = self.create_terminal_node(y_left_samples)
else:
node_dict["left_child"] = self.find_best_split(left_samples, y_left_samples)
self.build_tree(node_dict["left_child"], depth+1, max_depth, min_samples)
return node_dict
def create_terminal_node(self, y):
"""
Creates a terminal node.
Given a set of labels the most common label is computed and
set as the classification value of the node.
Args:
y: 1D numpy array with labels
Returns:
classification: int, predicted class
"""
classification = max(set(y), key=list(y).count)
return classification
def train(self, X, y, max_depth, min_samples):
"""
Fits decision tree on a given dataset.
Args:
X: 2D numpy array with data samples
y: 1D numpy array with labels
max_depth: int, maximum allowed tree depth
min_samples: int, minimum number of samples needed to split a node further
"""
self.n_classes = len(set(y))
self.root_dict = self.find_best_split(X, y)
self.tree_dict = self.build_tree(self.root_dict, 1, max_depth, min_samples)
def predict(self, X, node):
"""
Predicts the class for a given input example X.
Args:
X: 1D numpy array, input example
node: dict, representing trained decision tree
Returns:
prediction: int, predicted class
"""
feature_idx = node['feature_idx']
threshold = node['threshold']
if X[feature_idx] < threshold:
if isinstance(node['left_child'], (int, np.integer)):
return node['left_child']
else:
prediction = self.predict(X, node['left_child'])
elif X[feature_idx] >= threshold:
if isinstance(node['right_child'], (int, np.integer)):
return node['right_child']
else:
prediction = self.predict(X, node['right_child'])
return prediction
Initializing and training the decision tree¶
tree = DecisionTree()
tree.train(X_train, y_train, max_depth=2, min_samples=1)
Printing the decision tree structure¶
def print_tree(node, depth=0):
if isinstance(node, (int, np.integer)):
print(f"{depth * ' '}predicted class: {iris.target_names[node]}")
else:
print(f"{depth * ' '}{iris.feature_names[node['feature_idx']]} < {node['threshold']}, "
f"cost of split: {round(node['cost'], 3)}")
print_tree(node["left_child"], depth+1)
print_tree(node["right_child"], depth+1)
print_tree(tree.tree_dict)
petal length (cm) < 3.0, cost of split: 0.346
sepal length (cm) < 5.4, cost of split: 0.0
predicted class: setosa
predicted class: setosa
petal width (cm) < 1.8, cost of split: 0.097
predicted class: versicolor
predicted class: virginica
Testing the decision tree¶
all_predictions = []
for i in range(X_test.shape[0]):
result = tree.predict(X_test[i], tree.tree_dict)
all_predictions.append(y_test[i] == result)
print(f"Accuracy on test set: {sum(all_predictions) / len(all_predictions)}")
Accuracy on test set: 0.9473684210526315