Bayesian Temporal Matrix Factorization¶
Published: October 8, 2019
Revised: October 8, 2020
Author: Xinyu Chen [GitHub homepage]
Download: This Jupyter notebook is at our GitHub repository. If you want to evaluate the code, please download the notebook from the transdim repository.
This notebook shows how to implement the Bayesian Temporal Matrix Factorization (BTMF), a fully Bayesian matrix factorization model, on some real-world data sets. To overcome the missing data problem in multivariate time series, BTMF takes into account both low-rank matrix structure and time series autoregression. For an in-depth discussion of BTMF, please see [1].
Abstract¶
Large-scale and multidimensional spatiotemporal data sets are becoming ubiquitous in many real-world applications such as monitoring traffic and air quality. Making predictions on these time series has become a critical challenge due to not only the large-scale and high-dimensional nature but also the considerable amount of missing data. In this work, we propose a Bayesian Temporal Matrix Factorization (BTMF) model for modeling multidimensional time series - and in particular spatiotemporal data - in the presence of missing values. By integrating low-rank matrix factorization and vector autoregressive (VAR) process into a single probabilistic graphical model, our model can effectively perform predictions without imputing those missing values. We develop efficient Gibbs sampling algorithms for model inference and test BTMF on several real-world spatiotemporal data sets (i.e., a typical kind of multivariate time series data) for both missing data imputation and short-term rolling prediction tasks. This post is mainly about BTMF and its Python implementation with an application of spatiotemporal data imputation.
Problem Description¶
We assume a spatiotemporal setting for multidimensional time series data throughout this work. In general, modern spatiotemporal data sets collected from sensor networks can be organized as matrix time series. For example, we can denote by matrix $Y\in\mathbb{R}^{N\times T}$ a multivariate time series collected from $N$ locations/sensors on $T$ time points, with each row $$\boldsymbol{y}_{i}=\left(y_{i,1},y_{i,2},...,y_{i,t-1},y_{i,t},y_{i,t+1},...,y_{i,T}\right)$$ corresponding to the time series collected at location $i$.
As mentioned, making accurate predictions on incomplete time series is very challenging, while missing data problem is almost inevitable in real-world applications. Figure 1 illustrates the prediction problem for incomplete time series data. Here we use $(i,t)\in\Omega$ to index the observed entries in matrix $Y$.
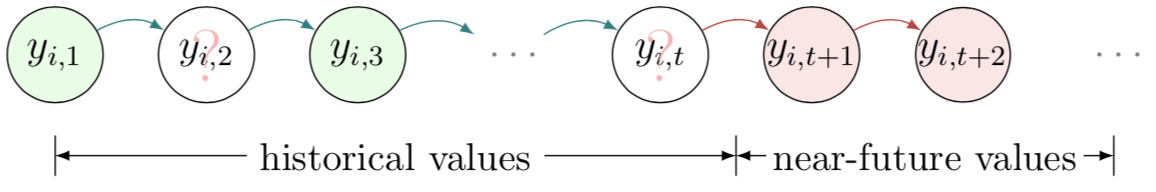
Figure 1: Illustration of multivariate time series and the prediction problem in the presence of missing values (green: observed data; white: missing data; red: prediction).
Model Description¶
Given a partially observed spatiotemporal matrix $Y\in\mathbb{R}^{N \times T}$, one can factorize it into a spatial factor matrix $W\in\mathbb{R}^{R \times N}$ and a temporal factor matrix $X\in\mathbb{R}^{R \times T}$ following general matrix factorization model: \begin{equation} Y\approx W^{\top}X, \label{btmf_equation1} \end{equation} and element-wise, we have \begin{equation} y_{it}\approx \boldsymbol{w}_{i}^\top\boldsymbol{x}_{t}, \quad \forall (i,t), \label{btmf_equation2} \end{equation} where vectors $\boldsymbol{w}_{i}$ and $\boldsymbol{x}_{t}$ refer to the $i$-th column of $W$ and the $t$-th column of $X$, respectively.
The standard matrix factorization model is a good approach to deal with the missing data problem; however, it cannot capture the dependencies among different columns in $X$, which are critical in modeling time series data. To better characterize the temporal dependencies and impose temporal smoothness, a novel AR regularizer is introduced on $X$ in TRMF (i.e., Temporal Regularizer Matrix Factorization proposed by Yu et al., 2016): \begin{equation} \label{equ:VAR} \begin{aligned} \boldsymbol{x}_{t+1}&=\sum\nolimits_{k=1}^{d}A_{k}\boldsymbol{x}_{t+1-h_k}+\boldsymbol{\epsilon}_t, \\ &=A^\top \boldsymbol{v}_{t+1}+\boldsymbol{\epsilon}_{t}, \\ \end{aligned} \end{equation} where $\mathcal{L}=\left\{h_1,\ldots,h_k,\ldots,h_d\right\}$ is a lag set ($d$ is the order of this AR model), each $A_k$ ($k\in\left\{1,...,d\right\}$) is a $R\times R$ coefficient matrix, and $\boldsymbol{\epsilon}_t$ is a zero mean Gaussian noise vector. For brevity, matrix $A\in \mathbb{R}^{(R d) \times R}$ and vector $\boldsymbol{v}_{t+1}\in \mathbb{R}^{(R d) \times 1}$ are defined as \begin{equation*} A=\left[A_{1}, \ldots, A_{d}\right]^{\top} ,\quad \boldsymbol{v}_{t+1}=\left[\begin{array}{c}{\boldsymbol{x}_{t+1-h_1}} \\ {\vdots} \\ {\boldsymbol{x}_{t+1-h_d}}\end{array}\right] . \end{equation*}

Figure 2: A graphical illustration of the rolling prediction scheme using BTMF (with VAR process) (green: observed data; white: missing data; red: prediction).
In Yu et al., 2016, to avoid overfitting and reduce the number of parameters, the coefficient matrix in TRMF is further assumed to be a diagonal $A_k=\text{diag}(\boldsymbol{\theta}_{k})$. Therefore, they have \begin{equation} \label{equ:AR} \boldsymbol{x}_{t+1}=\boldsymbol{\theta}_{1}\circledast\boldsymbol{x}_{t+1-h_1}+\cdots+\boldsymbol{\theta}_{d}\circledast\boldsymbol{x}_{t+1-h_d}+\boldsymbol{\epsilon}_t, \end{equation} where the symbol $\circledast$ denotes the element-wise Hadamard product. However, unlike this individual autoregressive (AR) process, a vector autoregressive (VAR) process is actually more powerful for capturing multivariate time series patterns.

Figure 3: A graphical illustration of the rolling prediction scheme using BTMF (with AR process) (green: observed data; white: missing data; red: prediction).
Bayesian Temporal Matrix Factorization Model¶
1 Model Specification¶
Following the general Bayesian probabilistic matrix factorization models (e.g., BPMF proposed by Salakhutdinov & Mnih, 2008), we assume that each observed entry in $Y$ follows a Gaussian distribution with precision $\tau$: \begin{equation} y_{i,t}\sim\mathcal{N}\left(\boldsymbol{w}_i^\top\boldsymbol{x}_t,\tau^{-1}\right),\quad \left(i,t\right)\in\Omega. \label{btmf_equation3} \end{equation}
On the spatial dimension, we use a simple Gaussian factor matrix without imposing any dependencies explicitly: \begin{equation} \boldsymbol{w}_i\sim\mathcal{N}\left(\boldsymbol{\mu}_{w},\Lambda_w^{-1}\right), \end{equation} and we place a conjugate Gaussian-Wishart prior on the mean vector and the precision matrix: \begin{equation} \boldsymbol{\mu}_w | \Lambda_w \sim\mathcal{N}\left(\boldsymbol{\mu}_0,(\beta_0\Lambda_w)^{-1}\right),\Lambda_w\sim\mathcal{W}\left(W_0,\nu_0\right), \end{equation} where $\boldsymbol{\mu}_0\in \mathbb{R}^{R}$ is a mean vector, $\mathcal{W}\left(W_0,\nu_0\right)$ is a Wishart distribution with a $R\times R$ scale matrix $W_0$ and $\nu_0$ degrees of freedom.
In modeling the temporal factor matrix $X$, we re-write the VAR process as: \begin{equation} \begin{aligned} \boldsymbol{x}_{t}&\sim\begin{cases} \mathcal{N}\left(\boldsymbol{0},I_R\right),&\text{if $t\in\left\{1,2,...,h_d\right\}$}, \\ \mathcal{N}\left(A^\top \boldsymbol{v}_{t},\Sigma\right),&\text{otherwise},\\ \end{cases}\\ \end{aligned} \label{btmf_equation5} \end{equation}
Since the mean vector is defined by VAR, we need to place the conjugate matrix normal inverse Wishart (MNIW) prior on the coefficient matrix $A$ and the covariance matrix $\Sigma$ as follows, \begin{equation} \begin{aligned} A\sim\mathcal{MN}_{(Rd)\times R}\left(M_0,\Psi_0,\Sigma\right),\quad \Sigma \sim\mathcal{IW}\left(S_0,\nu_0\right), \\ \end{aligned} \end{equation} where the probability density function for the $Rd$-by-$R$ random matrix $A$ has the form: \begin{equation} \begin{aligned} &p\left(A\mid M_0,\Psi_0,\Sigma\right) \\ =&\left(2\pi\right)^{-R^2d/2}\left|\Psi_0\right|^{-R/2}\left|\Sigma\right|^{-Rd/2} \\ &\times \exp\left(-\frac{1}{2}\text{tr}\left[\Sigma^{-1}\left(A-M_0\right)^{\top}\Psi_{0}^{-1}\left(A-M_0\right)\right]\right), \\ \end{aligned} \label{mnpdf} \end{equation} where $\Psi_0\in\mathbb{R}^{(Rd)\times (Rd)}$ and $\Sigma\in\mathbb{R}^{R\times R}$ are played as covariance matrices.
For the only remaining parameter $\tau$, we place a Gamma prior $\tau\sim\text{Gamma}\left(\alpha,\beta\right)$ where $\alpha$ and $\beta$ are the shape and rate parameters, respectively.
The above specifies the full generative process of BTMF, and we could also see the Bayesian graphical model shown in Figure 4. Several parameters are introduced to define the prior distributions for hyperparameters, including $\boldsymbol{\mu}_{0}$, $W_0$, $\nu_0$, $\beta_0$, $\alpha$, $\beta$, $M_0$, $\Psi_0$, and $S_0$. These parameters need to provided in advance when training the model. However, it should be noted that the specification of these parameters has little impact on the final results, as the training data will play a much more important role in defining the posteriors of the hyperparameters.

Figure 4: An overview graphical model of BTMF (time lag set: $\left\{1,2,...,d\right\}$). The shaded nodes ($y_{i,t}$) are the observed data in $\Omega$.
2 Model Inference¶
Given the complex structure of BTMF, it is intractable to write down the posterior distribution. Here we rely on the MCMC technique for Bayesian learning. In detail, we introduce a Gibbs sampling algorithm by deriving the full conditional distributions for all parameters and hyperparameters. Thanks to the use of conjugate priors in Figure 4, we can actually write down all the conditional distributions analytically. Below we summarize the Gibbs sampling procedure.
1) Sampling Factor Matrix $W$ and Its Hyperparameters¶
For programming convenience, we use $W\in\mathbb{R}^{N\times R}$ to replace $W\in\mathbb{R}^{R\times N}$.
import numpy as np
from numpy.linalg import inv as inv
from numpy.random import normal as normrnd
from scipy.linalg import khatri_rao as kr_prod
from scipy.stats import wishart
from scipy.stats import invwishart
from numpy.linalg import solve as solve
from numpy.linalg import cholesky as cholesky_lower
from scipy.linalg import cholesky as cholesky_upper
from scipy.linalg import solve_triangular as solve_ut
import matplotlib.pyplot as plt
%matplotlib inline
def mvnrnd_pre(mu, Lambda):
src = normrnd(size = (mu.shape[0],))
return solve_ut(cholesky_upper(Lambda, overwrite_a = True, check_finite = False),
src, lower = False, check_finite = False, overwrite_b = True) + mu
def cov_mat(mat, mat_bar):
mat = mat - mat_bar
return mat.T @ mat
def sample_factor_w(tau_sparse_mat, tau_ind, W, X, tau, beta0 = 1, vargin = 0):
"""Sampling N-by-R factor matrix W and its hyperparameters (mu_w, Lambda_w)."""
dim1, rank = W.shape
W_bar = np.mean(W, axis = 0)
temp = dim1 / (dim1 + beta0)
var_W_hyper = inv(np.eye(rank) + cov_mat(W, W_bar) + temp * beta0 * np.outer(W_bar, W_bar))
var_Lambda_hyper = wishart.rvs(df = dim1 + rank, scale = var_W_hyper)
var_mu_hyper = mvnrnd_pre(temp * W_bar, (dim1 + beta0) * var_Lambda_hyper)
if dim1 * rank ** 2 > 1e+8:
vargin = 1
if vargin == 0:
var1 = X.T
var2 = kr_prod(var1, var1)
var3 = (var2 @ tau_ind.T).reshape([rank, rank, dim1]) + var_Lambda_hyper[:, :, None]
var4 = var1 @ tau_sparse_mat.T + (var_Lambda_hyper @ var_mu_hyper)[:, None]
for i in range(dim1):
W[i, :] = mvnrnd_pre(solve(var3[:, :, i], var4[:, i]), var3[:, :, i])
elif vargin == 1:
for i in range(dim1):
pos0 = np.where(sparse_mat[i, :] != 0)
Xt = X[pos0[0], :]
var_mu = tau[i] * Xt.T @ sparse_mat[i, pos0[0]] + var_Lambda_hyper @ var_mu_hyper
var_Lambda = tau[i] * Xt.T @ Xt + var_Lambda_hyper
W[i, :] = mvnrnd_pre(solve(var_Lambda, var_mu), var_Lambda)
return W
2) Sampling VAR Coefficients $A$ and Its Hyperparameters¶
Foundations of VAR
Vector autoregression (VAR) is a multivariate extension of autoregression (AR). Formally, VAR for $R$-dimensional vectors $\boldsymbol{x}_{t}$ can be written as follows, \begin{equation} \begin{aligned} \boldsymbol{x}_{t}&=A_{1} \boldsymbol{x}_{t-h_1}+\cdots+A_{d} \boldsymbol{x}_{t-h_d}+\boldsymbol{\epsilon}_{t}, \\ &= A^\top \boldsymbol{v}_{t}+\boldsymbol{\epsilon}_{t},~t=h_d+1, \ldots, T, \\ \end{aligned} \end{equation} where \begin{equation} A=\left[A_{1}, \ldots, A_{d}\right]^{\top} \in \mathbb{R}^{(R d) \times R},\quad \boldsymbol{v}_{t}=\left[\begin{array}{c}{\boldsymbol{x}_{t-h_1}} \\ {\vdots} \\ {\boldsymbol{x}_{t-h_d}}\end{array}\right] \in \mathbb{R}^{(R d) \times 1}. \end{equation}
In the following, if we define \begin{equation} Z=\left[\begin{array}{c}{\boldsymbol{x}_{h_d+1}^{\top}} \\ {\vdots} \\ {\boldsymbol{x}_{T}^{\top}}\end{array}\right] \in \mathbb{R}^{(T-h_d) \times R},\quad Q=\left[\begin{array}{c}{\boldsymbol{v}_{h_d+1}^{\top}} \\ {\vdots} \\ {\boldsymbol{v}_{T}^{\top}}\end{array}\right] \in \mathbb{R}^{(T-h_d) \times(R d)}, \end{equation} then, we could write the above mentioned VAR as \begin{equation} \underbrace{Z}_{(T-h_d)\times R}\approx \underbrace{Q}_{(T-h_d)\times (Rd)}\times \underbrace{A}_{(Rd)\times R}. \end{equation}
To include temporal factors $\boldsymbol{x}_{t},t=1,...,h_d$, we also define $$Z_0=\left[\begin{array}{c}{\boldsymbol{x}_{1}^{\top}} \\ {\vdots} \\ {\boldsymbol{x}_{h_d}^{\top}}\end{array}\right] \in \mathbb{R}^{h_d \times R}.$$
Build a Bayesian VAR on temporal factors $\boldsymbol{x}_{t}$ \begin{equation} \begin{aligned} \boldsymbol{x}_{t}&\sim\begin{cases}\mathcal{N}\left(A^\top \boldsymbol{v}_{t},\Sigma\right),~\text{if $t\in\left\{h_d+1,...,T\right\}$},\\{\mathcal{N}\left(\boldsymbol{0},I_R\right),~\text{otherwise}}.\end{cases}\\ A&\sim\mathcal{MN}_{(Rd)\times R}\left(M_0,\Psi_0,\Sigma\right), \\ \Sigma &\sim\mathcal{IW}\left(S_0,\nu_0\right), \\ \end{aligned} \end{equation} where \begin{equation} \begin{aligned} &\mathcal{M N}_{(R d) \times R}\left(A | M_{0}, \Psi_{0}, \Sigma\right)\\ \propto|&\Sigma|^{-R d / 2} \exp \left(-\frac{1}{2} \operatorname{tr}\left[\Sigma^{-1}\left(A-M_{0}\right)^{\top} \Psi_{0}^{-1}\left(A-M_{0}\right)\right]\right), \\ \end{aligned} \end{equation} and \begin{equation} \mathcal{I} \mathcal{W}\left(\Sigma | S_{0}, \nu_{0}\right) \propto|\Sigma|^{-\left(\nu_{0}+R+1\right) / 2} \exp \left(-\frac{1}{2} \operatorname{tr}\left(\Sigma^{-1}S_{0}\right)\right). \end{equation}
Likelihood from temporal factors $\boldsymbol{x}_{t}$ \begin{equation} \begin{aligned} &\mathcal{L}\left(X\mid A,\Sigma\right) \\ \propto &\prod_{t=1}^{h_d}p\left(\boldsymbol{x}_{t}\mid \Sigma\right)\times \prod_{t=h_d+1}^{T}p\left(\boldsymbol{x}_{t}\mid A,\Sigma\right) \\ \propto &\left|\Sigma\right|^{-T/2}\exp\left\{-\frac{1}{2}\sum_{t=h_d+1}^{T}\left(\boldsymbol{x}_{t}-A^\top \boldsymbol{v}_{t}\right)^\top\Sigma^{-1}\left(\boldsymbol{x}_{t}-A^\top \boldsymbol{v}_{t}\right)\right\} \\ \propto &\left|\Sigma\right|^{-T/2}\exp\left\{-\frac{1}{2}\text{tr}\left[\Sigma^{-1}\left(Z_0^\top Z_0+\left(Z-QA\right)^\top \left(Z-QA\right)\right)\right]\right\} \end{aligned} \end{equation}
Posterior distribution
Consider \begin{equation} \begin{aligned} &\left(A-M_{0}\right)^{\top} \Psi_{0}^{-1}\left(A-M_{0}\right)+S_0+Z_0^\top Z_0+\left(Z-QA\right)^\top \left(Z-QA\right) \\ =&A^\top\left(\Psi_0^{-1}+Q^\top Q\right)A-A^\top\left(\Psi_0^{-1}M_0+Q^\top Z\right) \\ &-\left(\Psi_0^{-1}M_0+Q^\top Z\right)^\top A \\ &+\left(\Psi_0^{-1}M_0+Q^\top Z\right)^\top\left(\Psi_0^{-1}+Q^\top Q\right)\left(\Psi_0^{-1}M_0+Q^\top Z\right) \\ &-\left(\Psi_0^{-1}M_0+Q^\top Z\right)^\top\left(\Psi_0^{-1}+Q^\top Q\right)\left(\Psi_0^{-1}M_0+Q^\top Z\right) \\ &+M_0^\top\Psi_0^{-1}M_0+S_0+Z_0^\top Z_0+Z^\top Z \\ =&\left(A-M^{*}\right)^\top\left(\Psi^{*}\right)^{-1}\left(A-M^{*}\right)+S^{*}, \\ \end{aligned} \end{equation} which is in the form of $\mathcal{MN}\left(\cdot\right)$ and $\mathcal{IW}\left(\cdot\right)$.
The $Rd$-by-$R$ matrix $A$ has a matrix normal distribution, and $R$-by-$R$ covariance matrix $\Sigma$ has an inverse Wishart distribution, that is, \begin{equation} A \sim \mathcal{M N}_{(R d) \times R}\left(M^{*}, \Psi^{*}, \Sigma\right), \quad \Sigma \sim \mathcal{I} \mathcal{W}\left(S^{*}, \nu^{*}\right), \end{equation} with \begin{equation} \begin{cases} {\Psi^{*}=\left(\Psi_{0}^{-1}+Q^{\top} Q\right)^{-1}}, \\ {M^{*}=\Psi^{*}\left(\Psi_{0}^{-1} M_{0}+Q^{\top} Z\right)}, \\ {S^{*}=S_{0}+Z^\top Z+M_0^\top\Psi_0^{-1}M_0-\left(M^{*}\right)^\top\left(\Psi^{*}\right)^{-1}M^{*}}, \\ {\nu^{*}=\nu_{0}+T-h_d}. \end{cases} \end{equation}
def mnrnd(M, U, V):
"""
Generate matrix normal distributed random matrix.
M is a m-by-n matrix, U is a m-by-m matrix, and V is a n-by-n matrix.
"""
dim1, dim2 = M.shape
X0 = np.random.randn(dim1, dim2)
P = cholesky_lower(U)
Q = cholesky_lower(V)
return M + P @ X0 @ Q.T
def sample_var_coefficient(X, time_lags):
dim, rank = X.shape
d = time_lags.shape[0]
tmax = np.max(time_lags)
Z_mat = X[tmax : dim, :]
Q_mat = np.zeros((dim - tmax, rank * d))
for k in range(d):
Q_mat[:, k * rank : (k + 1) * rank] = X[tmax - time_lags[k] : dim - time_lags[k], :]
var_Psi0 = np.eye(rank * d) + Q_mat.T @ Q_mat
var_Psi = inv(var_Psi0)
var_M = var_Psi @ Q_mat.T @ Z_mat
var_S = np.eye(rank) + Z_mat.T @ Z_mat - var_M.T @ var_Psi0 @ var_M
Sigma = invwishart.rvs(df = rank + dim - tmax, scale = var_S)
return mnrnd(var_M, var_Psi, Sigma), Sigma
3) Sampling Factor Matrix $X$¶
Posterior distribution \begin{equation} \begin{aligned} y_{it}&\sim\mathcal{N}\left(\boldsymbol{w}_{i}^\top\boldsymbol{x}_{t},\tau^{-1}\right),~\left(i,t\right)\in\Omega, \\ \boldsymbol{x}_{t}&\sim\begin{cases}\mathcal{N}\left(\sum_{k=1}^{d}A_{k} \boldsymbol{x}_{t-h_k},\Sigma\right),~\text{if $t\in\left\{h_d+1,...,T\right\}$},\\{\mathcal{N}\left(\boldsymbol{0},I\right),~\text{otherwise}}.\end{cases}\\ \end{aligned} \end{equation}
If $t\in\left\{1,...,h_d\right\}$, parameters of the posterior distribution $\mathcal{N}\left(\boldsymbol{x}_{t}\mid \boldsymbol{\mu}_{t}^{*},\Sigma_{t}^{*}\right)$ are \begin{equation} \begin{aligned} \Sigma_{t}^{*}&=\left(\sum_{k=1, h_{d}<t+h_{k} \leq T}^{d} {A}_{k}^{\top} \Sigma^{-1} A_{k}+\tau\sum_{i:(i,t)\in\Omega}\boldsymbol{w}_{i}\boldsymbol{w}_{i}^\top+I\right)^{-1}, \\ \boldsymbol{\mu}_{t}^{*}&=\Sigma_{t}^{*}\left(\sum_{k=1, h_{d}<t+h_{k} \leq T}^{d} A_{k}^{\top} \Sigma^{-1} \boldsymbol{\psi}_{t+h_{k}}+\tau\sum_{i:(i,t)\in\Omega}\boldsymbol{w}_{i}y_{it}\right). \\ \end{aligned} \end{equation}
If $t\in\left\{h_d+1,...,T\right\}$, then parameters of the posterior distribution $\mathcal{N}\left(\boldsymbol{x}_{t}\mid \boldsymbol{\mu}_{t}^{*},\Sigma_{t}^{*}\right)$ are \begin{equation} \begin{aligned} \Sigma_{t}^{*}&=\left(\sum_{k=1, h_{d}<t+h_{k} \leq T}^{d} {A}_{k}^{\top} \Sigma^{-1} A_{k}+\tau\sum_{i:(i,t)\in\Omega}\boldsymbol{w}_{i}\boldsymbol{w}_{i}^\top+\Sigma^{-1}\right)^{-1}, \\ \boldsymbol{\mu}_{t}^{*}&=\Sigma_{t}^{*}\left(\sum_{k=1, h_{d}<t+h_{k} \leq T}^{d} A_{k}^{\top} \Sigma^{-1} \boldsymbol{\psi}_{t+h_{k}}+\tau\sum_{i:(i,t)\in\Omega}\boldsymbol{w}_{i}y_{it}+\Sigma^{-1}\sum_{k=1}^{d}A_{k}\boldsymbol{x}_{t-h_k}\right), \\ \end{aligned} \end{equation} where $$\boldsymbol{\psi}_{t+h_k}=\boldsymbol{x}_{t+h_k}-\sum_{l=1,l\neq k}^{d}A_{l}\boldsymbol{x}_{t+h_k-h_l}.$$
def sample_factor_x(tau_sparse_mat, tau_ind, time_lags, W, X, A, Lambda_x):
"""Sampling T-by-R factor matrix X."""
dim2, rank = X.shape
tmax = np.max(time_lags)
tmin = np.min(time_lags)
d = time_lags.shape[0]
A0 = np.dstack([A] * d)
for k in range(d):
A0[k * rank : (k + 1) * rank, :, k] = 0
mat0 = Lambda_x @ A.T
mat1 = np.einsum('kij, jt -> kit', A.reshape([d, rank, rank]), Lambda_x)
mat2 = np.einsum('kit, kjt -> ij', mat1, A.reshape([d, rank, rank]))
var1 = W.T
var2 = kr_prod(var1, var1)
var3 = (var2 @ tau_ind).reshape([rank, rank, dim2]) + Lambda_x[:, :, None]
var4 = var1 @ tau_sparse_mat
for t in range(dim2):
Mt = np.zeros((rank, rank))
Nt = np.zeros(rank)
Qt = mat0 @ X[t - time_lags, :].reshape(rank * d)
index = list(range(0, d))
if t >= dim2 - tmax and t < dim2 - tmin:
index = list(np.where(t + time_lags < dim2))[0]
elif t < tmax:
Qt = np.zeros(rank)
index = list(np.where(t + time_lags >= tmax))[0]
if t < dim2 - tmin:
Mt = mat2.copy()
temp = np.zeros((rank * d, len(index)))
n = 0
for k in index:
temp[:, n] = X[t + time_lags[k] - time_lags, :].reshape(rank * d)
n += 1
temp0 = X[t + time_lags[index], :].T - np.einsum('ijk, ik -> jk', A0[:, :, index], temp)
Nt = np.einsum('kij, jk -> i', mat1[index, :, :], temp0)
var3[:, :, t] = var3[:, :, t] + Mt
if t < tmax:
var3[:, :, t] = var3[:, :, t] - Lambda_x + np.eye(rank)
X[t, :] = mvnrnd_pre(solve(var3[:, :, t], var4[:, t] + Nt + Qt), var3[:, :, t])
return X
4) Sampling Precision $\tau$¶
def sample_precision_tau(sparse_mat, mat_hat, ind):
var_alpha = 1e-6 + 0.5 * np.sum(ind, axis = 1)
var_beta = 1e-6 + 0.5 * np.sum(((sparse_mat - mat_hat) ** 2) * ind, axis = 1)
return np.random.gamma(var_alpha, 1 / var_beta)
def sample_precision_scalar_tau(sparse_mat, mat_hat, ind):
var_alpha = 1e-6 + 0.5 * np.sum(ind)
var_beta = 1e-6 + 0.5 * np.sum(((sparse_mat - mat_hat) ** 2) * ind)
return np.random.gamma(var_alpha, 1 / var_beta)
def compute_mape(var, var_hat):
return np.sum(np.abs(var - var_hat) / var) / var.shape[0]
def compute_rmse(var, var_hat):
return np.sqrt(np.sum((var - var_hat) ** 2) / var.shape[0])
5) BTMF Implementation¶
def BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "factor"):
"""Bayesian Temporal Matrix Factorization, BTMF."""
dim1, dim2 = sparse_mat.shape
d = time_lags.shape[0]
W = init["W"]
X = init["X"]
if np.isnan(sparse_mat).any() == False:
ind = sparse_mat != 0
pos_obs = np.where(ind)
pos_test = np.where((dense_mat != 0) & (sparse_mat == 0))
elif np.isnan(sparse_mat).any() == True:
pos_test = np.where((dense_mat != 0) & (np.isnan(sparse_mat)))
ind = ~np.isnan(sparse_mat)
pos_obs = np.where(ind)
sparse_mat[np.isnan(sparse_mat)] = 0
dense_test = dense_mat[pos_test]
del dense_mat
tau = np.ones(dim1)
W_plus = np.zeros((dim1, rank))
X_plus = np.zeros((dim2, rank))
A_plus = np.zeros((rank * d, rank))
temp_hat = np.zeros(len(pos_test[0]))
show_iter = 200
mat_hat_plus = np.zeros((dim1, dim2))
for it in range(burn_iter + gibbs_iter):
tau_ind = tau[:, None] * ind
tau_sparse_mat = tau[:, None] * sparse_mat
W = sample_factor_w(tau_sparse_mat, tau_ind, W, X, tau)
A, Sigma = sample_var_coefficient(X, time_lags)
X = sample_factor_x(tau_sparse_mat, tau_ind, time_lags, W, X, A, inv(Sigma))
mat_hat = W @ X.T
if option == "factor":
tau = sample_precision_tau(sparse_mat, mat_hat, ind)
elif option == "pca":
tau = sample_precision_scalar_tau(sparse_mat, mat_hat, ind)
tau = tau * np.ones(dim1)
temp_hat += mat_hat[pos_test]
if (it + 1) % show_iter == 0 and it < burn_iter:
temp_hat = temp_hat / show_iter
print('Iter: {}'.format(it + 1))
print('MAPE: {:.6}'.format(compute_mape(dense_test, temp_hat)))
print('RMSE: {:.6}'.format(compute_rmse(dense_test, temp_hat)))
temp_hat = np.zeros(len(pos_test[0]))
print()
if it + 1 > burn_iter:
W_plus += W
X_plus += X
A_plus += A
mat_hat_plus += mat_hat
mat_hat = mat_hat_plus / gibbs_iter
W = W_plus / gibbs_iter
X = X_plus / gibbs_iter
A = A_plus / gibbs_iter
print('Imputation MAPE: {:.6}'.format(compute_mape(dense_test, mat_hat[:, : dim2][pos_test])))
print('Imputation RMSE: {:.6}'.format(compute_rmse(dense_test, mat_hat[:, : dim2][pos_test])))
print()
mat_hat[mat_hat < 0] = 0
return mat_hat, W, X, A
Evaluation on Guangzhou Speed Data¶
Scenario setting:
- Tensor size: $214\times 61\times 144$ (road segment, day, time of day)
- Non-random missing (NM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Non-random missing (NM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1])[:, :, np.newaxis] + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 10
- Time lags: {1, 2, 144}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
time_lags = np.array([1, 2, 144])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.104269 RMSE: 4.36081 Iter: 400 MAPE: 0.101626 RMSE: 4.30256 Iter: 600 MAPE: 0.101615 RMSE: 4.3037 Iter: 800 MAPE: 0.101683 RMSE: 4.30592 Iter: 1000 MAPE: 0.101673 RMSE: 4.30681 Imputation MAPE: 0.101755 Imputation RMSE: 4.30853 Running time: 933 seconds
Scenario setting:
- Tensor size: $214\times 61\times 144$ (road segment, day, time of day)
- Random missing (RM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 80
- Time lags: {1, 2, 144}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 80
time_lags = np.array([1, 2, 144])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0778319 RMSE: 3.33635 Iter: 400 MAPE: 0.0768004 RMSE: 3.31004 Iter: 600 MAPE: 0.0764412 RMSE: 3.30118 Iter: 800 MAPE: 0.076182 RMSE: 3.29244 Iter: 1000 MAPE: 0.0761008 RMSE: 3.29149 Imputation MAPE: 0.0760784 Imputation RMSE: 3.29119 Running time: 6351 seconds
Scenario setting:
- Tensor size: $214\times 61\times 144$ (road segment, day, time of day)
- Random missing (RM)
- 60% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.6 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 80
- Time lags: {1, 2, 144}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 80
time_lags = np.array([1, 2, 144])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0842265 RMSE: 3.59357 Iter: 400 MAPE: 0.0831631 RMSE: 3.55573 Iter: 600 MAPE: 0.0826056 RMSE: 3.53509 Iter: 800 MAPE: 0.0822812 RMSE: 3.52181 Iter: 1000 MAPE: 0.0820294 RMSE: 3.51294 Imputation MAPE: 0.0819592 Imputation RMSE: 3.51001 Running time: 6355 seconds
Evaluation on Birmingham Parking Data¶
Scenario setting:
- Tensor size: $30\times 77\times 18$ (parking slot, day, time of day)
- Non-random missing (NM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Non-random missing (NM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1])[:, :, np.newaxis] + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 20
- Time lags: {1, 2, 18}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 18])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.160954 RMSE: 99.1409 Iter: 400 MAPE: 0.130435 RMSE: 80.5966 Iter: 600 MAPE: 0.133369 RMSE: 80.2392 Iter: 800 MAPE: 0.127632 RMSE: 77.7049 Iter: 1000 MAPE: 0.133299 RMSE: 74.0348 Imputation MAPE: 0.13379 Imputation RMSE: 69.5338 Running time: 204 seconds
Scenario setting:
- Tensor size: $30\times 77\times 18$ (parking slot, day, time of day)
- Random missing (RM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 20
- Time lags: {1, 2, 18}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 18])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0437143 RMSE: 21.7522 Iter: 400 MAPE: 0.0356367 RMSE: 17.9024 Iter: 600 MAPE: 0.0345634 RMSE: 17.0658 Iter: 800 MAPE: 0.0340878 RMSE: 16.2093 Iter: 1000 MAPE: 0.0347393 RMSE: 16.63 Imputation MAPE: 0.0343784 Imputation RMSE: 18.1942 Running time: 204 seconds
Scenario setting:
- Tensor size: $30\times 77\times 18$ (parking slot, day, time of day)
- Random missing (RM)
- 60% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.6 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 20
- Time lags: {1, 2, 18}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 18])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0832529 RMSE: 64.7848 Iter: 400 MAPE: 0.0700006 RMSE: 33.6699 Iter: 600 MAPE: 0.0663554 RMSE: 27.0075 Iter: 800 MAPE: 0.0629423 RMSE: 26.593 Iter: 1000 MAPE: 0.0619536 RMSE: 27.0718 Imputation MAPE: 0.0616117 Imputation RMSE: 25.9018 Running time: 204 seconds
Evaluation on Hangzhou Flow Data¶
Scenario setting:
- Tensor size: $80\times 25\times 108$ (metro station, day, time of day)
- Non-random missing (NM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Non-random missing (NM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1])[:, :, np.newaxis] + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 108}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 108])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.280562 RMSE: 103.909 Iter: 400 MAPE: 0.27785 RMSE: 84.921 Iter: 600 MAPE: 0.267921 RMSE: 68.0001 Iter: 800 MAPE: 0.258384 RMSE: 58.105 Iter: 1000 MAPE: 0.259447 RMSE: 48.4835 Imputation MAPE: 0.259309 Imputation RMSE: 46.0055 Running time: 626 seconds
Scenario setting:
- Tensor size: $80\times 25\times 108$ (metro station, day, time of day)
- Random missing (RM)
- 40% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.4 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 108}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 108])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.230024 RMSE: 41.7666 Iter: 400 MAPE: 0.237999 RMSE: 34.9787 Iter: 600 MAPE: 0.238276 RMSE: 31.4654 Iter: 800 MAPE: 0.237635 RMSE: 30.5488 Iter: 1000 MAPE: 0.238353 RMSE: 30.1015 Imputation MAPE: 0.239989 Imputation RMSE: 30.0307 Running time: 597 seconds
Scenario setting:
- Tensor size: $80\times 25\times 108$ (metro station, day, time of day)
- Random missing (RM)
- 60% missing rate
import time
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
missing_rate = 0.6 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 108}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 108])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.241568 RMSE: 47.9988 Iter: 400 MAPE: 0.253166 RMSE: 40.6155 Iter: 600 MAPE: 0.260333 RMSE: 37.0129 Iter: 800 MAPE: 0.26306 RMSE: 36.3198 Iter: 1000 MAPE: 0.255018 RMSE: 35.4753 Imputation MAPE: 0.251851 Imputation RMSE: 35.2499 Running time: 598 seconds
Evaluation on Seattle Speed Data¶
Scenario setting:
- Tensor size: $323\times 28\times 288$ (road segment, day, time of day)
- Non-random missing (NM)
- 40% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Seattle-data-set/tensor.npz')['arr_0']
dim = dense_tensor.shape
missing_rate = 0.4 # Non-random missing (NM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1])[:, :, np.newaxis] + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 10
- Time lags: {1, 2, 288}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 10
time_lags = np.array([1, 2, 288])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0963432 RMSE: 5.48416 Iter: 400 MAPE: 0.0926811 RMSE: 5.34828 Iter: 600 MAPE: 0.0926641 RMSE: 5.3471 Iter: 800 MAPE: 0.0926791 RMSE: 5.34896 Iter: 1000 MAPE: 0.0926937 RMSE: 5.349 Imputation MAPE: 0.0927048 Imputation RMSE: 5.35061 Running time: 1015 seconds
Scenario setting:
- Tensor size: $323\times 28\times 288$ (road segment, day, time of day)
- Random missing (RM)
- 40% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Seattle-data-set/tensor.npz')['arr_0']
dim = dense_tensor.shape
missing_rate = 0.4 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 50
- Time lags: {1, 2, 288}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
time_lags = np.array([1, 2, 288])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.060615 RMSE: 3.78165 Iter: 400 MAPE: 0.0598819 RMSE: 3.76034 Iter: 600 MAPE: 0.0596581 RMSE: 3.75081 Iter: 800 MAPE: 0.059572 RMSE: 3.74568 Iter: 1000 MAPE: 0.05955 RMSE: 3.7446 Imputation MAPE: 0.059578 Imputation RMSE: 3.74506 Running time: 2911 seconds
Scenario setting:
- Tensor size: $323\times 28\times 288$ (road segment, day, time of day)
- Random missing (RM)
- 60% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Seattle-data-set/tensor.npz')['arr_0']
dim = dense_tensor.shape
missing_rate = 0.6 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dense_mat = dense_tensor.reshape([dim[0], dim[1] * dim[2]])
sparse_mat = sparse_tensor.reshape([dim[0], dim[1] * dim[2]])
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 50
- Time lags: {1, 2, 288}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 50
time_lags = np.array([1, 2, 288])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0634663 RMSE: 3.9178 Iter: 400 MAPE: 0.0619926 RMSE: 3.8534 Iter: 600 MAPE: 0.061864 RMSE: 3.84633 Iter: 800 MAPE: 0.0617191 RMSE: 3.84037 Iter: 1000 MAPE: 0.0616136 RMSE: 3.83723 Imputation MAPE: 0.0615285 Imputation RMSE: 3.83463 Running time: 2920 seconds
Evaluation on London Movement Speed Data¶
London movement speed data set is is a city-wide hourly traffic speeddataset collected in London.
- Collected from 200,000+ road segments.
- 720 time points in April 2019.
- 73% missing values in the original data.
| Observation rate | $>90\%$ | $>80\%$ | $>70\%$ | $>60\%$ | $>50\%$ |
|---|---|---|---|---|---|
| Number of roads | 17,666 | 27,148 | 35,912 | 44,352 | 52,727 |
If want to test on the full dataset, you could consider the following setting for masking observations as missing values.
import numpy as np
np.random.seed(1000)
mask_rate = 0.20
dense_mat = np.load('../datasets/London-data-set/hourly_speed_mat.npy')
pos_obs = np.where(dense_mat != 0)
num = len(pos_obs[0])
sample_ind = np.random.choice(num, size = int(mask_rate * num), replace = False)
sparse_mat = dense_mat.copy()
sparse_mat[pos_obs[0][sample_ind], pos_obs[1][sample_ind]] = 0
Notably, you could also consider to evaluate the model on a subset of the data with the following setting.
import numpy as np
np.random.seed(1000)
missing_rate = 0.4
dense_mat = np.load('../datasets/London-data-set/hourly_speed_mat.npy')
binary_mat = dense_mat.copy()
binary_mat[binary_mat != 0] = 1
pos = np.where(np.sum(binary_mat, axis = 1) > 0.7 * binary_mat.shape[1])
dense_mat = dense_mat[pos[0], :]
## Non-random missing (NM)
binary_mat = np.zeros(dense_mat.shape)
random_mat = np.random.rand(dense_mat.shape[0], 30)
for i1 in range(dense_mat.shape[0]):
for i2 in range(30):
binary_mat[i1, i2 * 24 : (i2 + 1) * 24] = np.round(random_mat[i1, i2] + 0.5 - missing_rate)
sparse_mat = dense_mat * binary_mat
Model setting:
- Low rank: 20
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 24])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.101369 RMSE: 2.52938 Iter: 400 MAPE: 0.0943416 RMSE: 2.32419 Iter: 600 MAPE: 0.0942804 RMSE: 2.32234 Iter: 800 MAPE: 0.0942548 RMSE: 2.32151 Iter: 1000 MAPE: 0.0942522 RMSE: 2.32131 Imputation MAPE: 0.0942448 Imputation RMSE: 2.32114 Running time: 3434 seconds
import numpy as np
np.random.seed(1000)
missing_rate = 0.4
dense_mat = np.load('../datasets/London-data-set/hourly_speed_mat.npy')
binary_mat = dense_mat.copy()
binary_mat[binary_mat != 0] = 1
pos = np.where(np.sum(binary_mat, axis = 1) > 0.7 * binary_mat.shape[1])
dense_mat = dense_mat[pos[0], :]
## Random missing (RM)
random_mat = np.random.rand(dense_mat.shape[0], dense_mat.shape[1])
binary_mat = np.round(random_mat + 0.5 - missing_rate)
sparse_mat = dense_mat * binary_mat
Model setting:
- Low rank: 20
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 24])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.101091 RMSE: 2.53104 Iter: 400 MAPE: 0.0915506 RMSE: 2.26009 Iter: 600 MAPE: 0.0915279 RMSE: 2.25953 Iter: 800 MAPE: 0.0915008 RMSE: 2.25885 Iter: 1000 MAPE: 0.0914937 RMSE: 2.25851 Imputation MAPE: 0.0914913 Imputation RMSE: 2.25831 Running time: 3707 seconds
import numpy as np
np.random.seed(1000)
missing_rate = 0.6
dense_mat = np.load('../datasets/London-data-set/hourly_speed_mat.npy')
binary_mat = dense_mat.copy()
binary_mat[binary_mat != 0] = 1
pos = np.where(np.sum(binary_mat, axis = 1) > 0.7 * binary_mat.shape[1])
dense_mat = dense_mat[pos[0], :]
## Random missing (RM)
random_mat = np.random.rand(dense_mat.shape[0], dense_mat.shape[1])
binary_mat = np.round(random_mat + 0.5 - missing_rate)
sparse_mat = dense_mat * binary_mat
Model setting:
- Low rank: 20
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 20
time_lags = np.array([1, 2, 24])
init = {"W": 0.01 * np.random.randn(dim1, rank), "X": 0.01 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter)
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.118065 RMSE: 2.97448 Iter: 400 MAPE: 0.0933272 RMSE: 2.30121 Iter: 600 MAPE: 0.0933082 RMSE: 2.30072 Iter: 800 MAPE: 0.0933009 RMSE: 2.30048 Iter: 1000 MAPE: 0.093254 RMSE: 2.29921 Imputation MAPE: 0.0931998 Imputation RMSE: 2.29791 Running time: 3668 seconds
Eavluation on NYC Taxi Flow Data¶
Scenario setting:
- Tensor size: $30\times 30\times 1461$ (origin, destination, time)
- Random missing (RM)
- 40% missing rate
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/NYC-data-set/tensor.mat')['tensor'].astype(np.float32)
dim = dense_tensor.shape
missing_rate = 0.4 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 24])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.438729 RMSE: 4.51639 Iter: 400 MAPE: 0.441871 RMSE: 4.58073 Iter: 600 MAPE: 0.442271 RMSE: 4.58424 Iter: 800 MAPE: 0.442407 RMSE: 4.58285 Iter: 1000 MAPE: 0.442621 RMSE: 4.58954 Imputation MAPE: 0.442656 Imputation RMSE: 4.5902 Running time: 576 seconds
Scenario setting:
- Tensor size: $30\times 30\times 1461$ (origin, destination, time)
- Random missing (RM)
- 60% missing rate
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/NYC-data-set/tensor.mat')['tensor'].astype(np.float32)
dim = dense_tensor.shape
missing_rate = 0.6 # Random missing (RM)
sparse_tensor = dense_tensor * np.round(np.random.rand(dim[0], dim[1], dim[2]) + 0.5 - missing_rate)
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 24])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.449161 RMSE: 4.69922 Iter: 400 MAPE: 0.454175 RMSE: 4.77746 Iter: 600 MAPE: 0.455749 RMSE: 4.81528 Iter: 800 MAPE: 0.455906 RMSE: 4.81052 Iter: 1000 MAPE: 0.456352 RMSE: 4.81612 Imputation MAPE: 0.456358 Imputation RMSE: 4.81239 Running time: 498 seconds
Scenario setting:
- Tensor size: $30\times 30\times 1461$ (origin, destination, time)
- Non-random missing (NM)
- 40% missing rate
import scipy.io
import numpy as np
np.random.seed(1000)
dense_tensor = scipy.io.loadmat('../datasets/NYC-data-set/tensor.mat')['tensor']
dim = dense_tensor.shape
nm_tensor = np.random.rand(dim[0], dim[1], dim[2])
missing_rate = 0.4 # Non-random missing (NM)
binary_tensor = np.zeros(dense_tensor.shape)
for i1 in range(dim[0]):
for i2 in range(dim[1]):
for i3 in range(61):
binary_tensor[i1, i2, i3 * 24 : (i3 + 1) * 24] = np.round(nm_tensor[i1, i2, i3] + 0.5 - missing_rate)
sparse_tensor = dense_tensor * binary_tensor
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
del dense_tensor, sparse_tensor
Model setting:
- Low rank: 30
- Time lags: {1, 2, 24}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 24])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.444464 RMSE: 4.65518 Iter: 400 MAPE: 0.447948 RMSE: 4.75414 Iter: 600 MAPE: 0.448308 RMSE: 4.75636 Iter: 800 MAPE: 0.448359 RMSE: 4.7692 Iter: 1000 MAPE: 0.448241 RMSE: 4.76313 Imputation MAPE: 0.448418 Imputation RMSE: 4.77218 Running time: 517 seconds
Evaluation on Pacific Surface Temperature Data¶
Scenario setting:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Random missing (RM)
- 40% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], dense_tensor.shape[2])
missing_rate = 0.4
## Random missing (RM)
binary_tensor = np.round(random_tensor + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
Model setting:
- Low rank: 30
- Time lags: {1, 2, 12}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 12])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0137389 RMSE: 0.457494 Iter: 400 MAPE: 0.0137986 RMSE: 0.45871 Iter: 600 MAPE: 0.0137956 RMSE: 0.458598 Iter: 800 MAPE: 0.0137929 RMSE: 0.45853 Iter: 1000 MAPE: 0.0137936 RMSE: 0.458537 Imputation MAPE: 0.0137925 Imputation RMSE: 0.458508 Running time: 344 seconds
Scenario setting:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Random missing (RM)
- 60% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], dense_tensor.shape[2])
missing_rate = 0.6
## Random missing (RM)
binary_tensor = np.round(random_tensor + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
Model setting:
- Low rank: 30
- Time lags: {1, 2, 12}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 12])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.014609 RMSE: 0.489699 Iter: 400 MAPE: 0.0145158 RMSE: 0.48626 Iter: 600 MAPE: 0.0145027 RMSE: 0.485859 Iter: 800 MAPE: 0.0145018 RMSE: 0.485809 Iter: 1000 MAPE: 0.0144967 RMSE: 0.485647 Imputation MAPE: 0.0144962 Imputation RMSE: 0.485584 Running time: 358 seconds
Scenario setting:
- Tensor size: $30\times 84\times 396$ (location x, location y, month)
- Non-random missing (NM)
- 40% missing rate
import numpy as np
np.random.seed(1000)
dense_tensor = np.load('../datasets/Temperature-data-set/tensor.npy').astype(np.float32)
pos = np.where(dense_tensor[:, 0, :] > 50)
dense_tensor[pos[0], :, pos[1]] = 0
random_tensor = np.random.rand(dense_tensor.shape[0], dense_tensor.shape[1], int(dense_tensor.shape[2] / 3))
missing_rate = 0.4
## Non-random missing (NM)
binary_tensor = np.zeros(dense_tensor.shape)
for i1 in range(dense_tensor.shape[0]):
for i2 in range(dense_tensor.shape[1]):
for i3 in range(int(dense_tensor.shape[2] / 3)):
binary_tensor[i1, i2, i3 * 3 : (i3 + 1) * 3] = np.round(random_tensor[i1, i2, i3] + 0.5 - missing_rate)
sparse_tensor = dense_tensor.copy()
sparse_tensor[binary_tensor == 0] = np.nan
sparse_tensor[sparse_tensor == 0] = np.nan
dim1, dim2, dim3 = dense_tensor.shape
dense_mat = np.zeros((dim1 * dim2, dim3))
sparse_mat = np.zeros((dim1 * dim2, dim3))
for i in range(dim1):
dense_mat[i * dim2 : (i + 1) * dim2, :] = dense_tensor[i, :, :]
sparse_mat[i * dim2 : (i + 1) * dim2, :] = sparse_tensor[i, :, :]
Model setting:
- Low rank: 30
- Time lags: {1, 2, 12}
- The number of burn-in iterations: 1000
- The number of Gibbs iterations: 200
import time
start = time.time()
dim1, dim2 = sparse_mat.shape
rank = 30
time_lags = np.array([1, 2, 12])
init = {"W": 0.1 * np.random.randn(dim1, rank), "X": 0.1 * np.random.randn(dim2, rank)}
burn_iter = 1000
gibbs_iter = 200
mat_hat, W, X, A = BTMF(dense_mat, sparse_mat, init, rank, time_lags, burn_iter, gibbs_iter, option = "pca")
end = time.time()
print('Running time: %d seconds'%(end - start))
Iter: 200 MAPE: 0.0144348 RMSE: 0.478619 Iter: 400 MAPE: 0.0144376 RMSE: 0.478621 Iter: 600 MAPE: 0.0144356 RMSE: 0.478508 Iter: 800 MAPE: 0.0144289 RMSE: 0.478346 Iter: 1000 MAPE: 0.0144225 RMSE: 0.478176 Imputation MAPE: 0.0144058 Imputation RMSE: 0.477715 Running time: 354 seconds