SYDE 556/750: Simulating Neurobiological Systems¶
Administration¶
- 2-slide presentations on project topics
- Mar 30th: Castricato, Guliani, Stoeckel, Tolooshams, Bradshaw, Reyes, Griffin, Zheng, Lee, Nguyen, Raghavan, Thompson
- Apr 3rd: Ganjidoost, Noukhovitch, Pacheco, Abarca, Lambert, Tsatskin, Barnard, Ansari, Kahn, Orr,
Memory¶
Readings: Serial Working Memory; Associative Memory
- We've seen how to represent symbol-like structures using vectors
- Typically high dimensional vectors
- How can we store those over short periods of time (working memory)
- Long periods of time?
- Recall Jackendoff's 4th challenge (same repn in long and working memory)
Attractor networks¶
- An attractor in dynamical systems theory is a system state (or states) towards which other states tend over time
- The standard analogy is to imagine the state space as a ‘hill-like’ topology which a ball travels through (tending downhill).

- In neural network research, attractor networks (networks with dynamical attractors) have long been thought relevant for various behaviours
- e.g., memory, integration, off-line updating of representations, repetitive pattern generation, noise reduction, etc.
The neural integrator is an example of an attractor network
The neural integrator can be thought of as a line attractor as in a)
- Actually an approximate one as in b)
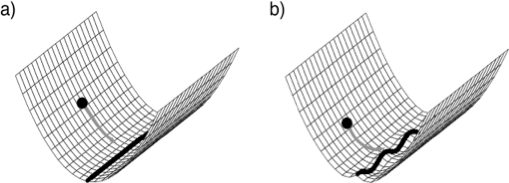
- Attractor networks were extensively examined in the ANN community (e.g. hopfield nets). Amit suggested that persistent activity could be associated with recurrent biological networks
- Persistent activity is found in motor, premotor, parietal, prefrontal, frontal, hippocampal, and inferotemporal cortex; and basal ganglia, midbrain, superior colliculus, and brainstem
- Focus to date is often on simple attractors:
- The NEF lets us easily generalize.
Generalizing dynamics¶
- A cyclic attractor (i.e. a set of attractive points that the system moves through)

Note: The hill and ball analogy doesn't work anymore.
This is an oscillator. Technically a nonlinear one, since it should be stable (the Simple Harmonic Oscillator is linear). Let's build a stable oscillator.
%pylab inline
import nengo
from nengo.utils.ensemble import response_curves
from nengo.dists import Uniform
model = nengo.Network('Oscillator')
freq = .25
scale = 1.1
N=300
with model:
stim = nengo.Node(lambda t: [.5,.5] if .1<t<.12 else [0,0])
inhib = nengo.Node(lambda t: [-1]*N if .8<t<.82 else [0]*N)
osc = nengo.Ensemble(N, dimensions=2, intercepts=Uniform(.3,1))
def feedback(x):
return scale*x[0]+freq*x[1], -freq*x[0]+scale*x[1]
nengo.Connection(osc, osc, function=feedback)
nengo.Connection(inhib, osc.neurons)
nengo.Connection(stim, osc)
osc_p = nengo.Probe(osc, synapse=.01)
WARNING: pylab import has clobbered these variables: ['seed'] `%matplotlib` prevents importing * from pylab and numpy
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/nonlinear_oscillator.py.cfg")
sim = nengo.Simulator(model)
sim.run(1)
x, A = response_curves(osc, sim)
figure(figsize=(4,2))
plot(x, A)
xlabel('x')
ylabel('firing rate (Hz)')
figure(figsize=(12,4))
subplot(1,2,1)
plot(sim.trange(), sim.data[osc_p]);
xlabel('Time (s)')
ylabel('State value')
subplot(1,2,2)
plot(sim.data[osc_p][:,0],sim.data[osc_p][:,1])
xlabel('$x_0$')
ylabel('$x_1$');
Building finished in 0:00:01. Simulating finished in 0:00:01.
- This also generalizes to chaotic attractors and many other fun dynamic networks (see Lecture 5).
- Can also generalize representation
Generalizing representation¶
- Generalizing a line attractor gives something much more interesting
- A plane attractor

- The attractor space is much more complicated, so you need a lot more neurons to do as good a job ($N^D$)
%pylab inline
#A 1D integrator with a few hundred neurons works great
import nengo
from nengo.utils.functions import piecewise
model = nengo.Network(label='1D Line Attractor', seed=5)
N = 300
tau = 0.01
with model:
stim = nengo.Node(piecewise({.3:[1], .5:[0] }))
neurons = nengo.Ensemble(N, dimensions=1)
nengo.Connection(stim, neurons, transform=tau, synapse=tau)
nengo.Connection(neurons, neurons, synapse=tau)
stim_p = nengo.Probe(stim)
neurons_p = nengo.Probe(neurons, synapse=.01)
sim = nengo.Simulator(model)
sim.run(4)
t=sim.trange()
plot(t, sim.data[stim_p], label = "stim")
plot(t, sim.data[neurons_p], label = "position")
legend(loc="best");
Populating the interactive namespace from numpy and matplotlib Building finished in 0:00:01.
WARNING: pylab import has clobbered these variables: ['piecewise'] `%matplotlib` prevents importing * from pylab and numpy
Simulating finished in 0:00:01.
#Need lots of neurons to get reasonable performance in higher D
import nengo
from nengo.utils.functions import piecewise
model = nengo.Network(label='2D Plane Attractor', seed=4)
N = 2000 #100
tau = 0.01
with model:
stim = nengo.Node(piecewise({.3:[1, -1], .5:[0, 0] }))
neurons = nengo.Ensemble(N, dimensions=2)
nengo.Connection(stim, neurons, transform=tau, synapse=tau)
nengo.Connection(neurons, neurons, synapse=tau)
stim_p = nengo.Probe(stim)
neurons_p = nengo.Probe(neurons, synapse=.01)
sim = nengo.Simulator(model)
sim.run(4)
t=sim.trange()
plot(t, sim.data[stim_p], label = "stim")
plot(t, sim.data[neurons_p], label = "position")
legend(loc="best");
Building finished in 0:00:01. Simulating finished in 0:00:01.
#Note that the representation saturates at the radius
with model:
stim.output = piecewise({.2:[1, -1], 1.2:[0, 0] })
sim = nengo.Simulator(model)
sim.run(4)
t=sim.trange()
plot(t, sim.data[stim_p], label = "stim")
plot(t, sim.data[neurons_p], label = "position");
Building finished in 0:00:01. Simulating finished in 0:00:02.
- So for higher dimensional memories, you may want the dimensions to be independent
- To make this easy, Nengo has 'ensemble arrays'
model = nengo.Network(label='Ensemble Array', seed=123)
N = 300
tau = 0.01
with model:
stim = nengo.Node(piecewise({.3:[1, -1], .5:[0, 0] }))
neurons = nengo.networks.EnsembleArray(N, n_ensembles=2)
nengo.Connection(stim, neurons.input, transform=tau, synapse=tau)
nengo.Connection(neurons.output, neurons.input, synapse=tau)
stim_p = nengo.Probe(stim)
neurons_p = nengo.Probe(neurons.output, synapse=.01)
sim = nengo.Simulator(model)
sim.run(4)
t=sim.trange()
plot(t, sim.data[stim_p], label = "stim")
plot(t, sim.data[neurons_p], label = "position");
Building finished in 0:00:01. Simulating finished in 0:00:02.
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/ensemble_array.py.cfg")
- And saturation effects are quite different
#Note that the representation saturates at the radius
with model:
stim.output = piecewise({.2:[1, -1], 1.2:[0, 0] })
sim = nengo.Simulator(model)
sim.run(4)
t=sim.trange()
plot(t, sim.data[stim_p], label = "stim")
plot(t, sim.data[neurons_p], label = "position");
Working memory¶
- We can build high dimensional working memories using plane attractors
- In general we want some saturation, and some independence between dimensions
- Gives both a 'soft normalization'
- And good temporal stability for fewer neurons
import nengo
from nengo import spa
def color_input(t):
if t < 0.15:
return 'BLUE'
elif 1.0 < t < 1.15:
return 'GREEN'
elif 1.7 < t < 1.85:
return 'RED'
else:
return '0'
model = spa.SPA(label="HighD Working Memory", seed=5)
dimensions = 32
with model:
model.color_in = spa.Buffer(dimensions=dimensions)
model.mem = spa.Memory(dimensions=dimensions, subdimensions=4,
synapse=0.1, neurons_per_dimension=50)
# Connect the buffers
cortical_actions = spa.Actions(
'mem = color_in'
)
model.cortical = spa.Cortical(cortical_actions)
model.inp = spa.Input(color_in=color_input)
model.config[nengo.Probe].synapse = nengo.Lowpass(0.03)
color_in = nengo.Probe(model.color_in.state.output)
mem = nengo.Probe(model.mem.state.output)
sim = nengo.Simulator(model)
sim.run(3.)
plt.figure(figsize=(10, 10))
vocab = model.get_default_vocab(dimensions)
plt.subplot(2, 1, 1)
plt.plot(sim.trange(), model.similarity(sim.data, color_in))
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("color")
plt.subplot(2, 1, 2)
plt.plot(sim.trange(), model.similarity(sim.data, mem))
plt.legend(fontsize='x-small')
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("memory")
plt.xlabel("time [s]");
//anaconda/lib/python2.7/site-packages/matplotlib/axes/_axes.py:519: UserWarning: No labelled objects found. Use label='...' kwarg on individual plots.
warnings.warn("No labelled objects found. "
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/simple_spa_wm.py.cfg")
# model.all_ensembles has all the neurons
model.all_ensembles[2].n_neurons
200
You can see interference effects here
If you recalled things from this memory, you'd probably have a 'recency effect'
- i.e. The things most recently put in memory would be best recalled
This is seen in human memory, but so is primacy
- How could we get primacy?
import nengo
from nengo import spa
def color_input(t):
if t < 0.15:
return 'BLUE'
elif 1.0 < t < 1.15:
return 'GREEN'
elif 1.7 < t < 1.85:
return 'RED'
else:
return '0'
model = spa.SPA(label="HighD Working Memory", seed=5)
dimensions = 32
with model:
model.color_in = spa.Buffer(dimensions=dimensions)
model.mem = spa.Memory(dimensions=dimensions, subdimensions=4,
synapse=0.1, neurons_per_dimension=50,
tau=-.2)
# Connect the buffers
cortical_actions = spa.Actions(
'mem = color_in'
)
model.cortical = spa.Cortical(cortical_actions)
model.inp = spa.Input(color_in=color_input)
model.config[nengo.Probe].synapse = nengo.Lowpass(0.03)
color_in = nengo.Probe(model.color_in.state.output)
mem = nengo.Probe(model.mem.state.output)
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/simple_spa_wm_primacy.py.cfg")
sim = nengo.Simulator(model)
sim.run(3.)
plt.figure(figsize=(10, 10))
vocab = model.get_default_vocab(dimensions)
plt.subplot(2, 1, 1)
plt.plot(sim.trange(), model.similarity(sim.data, color_in))
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("color")
plt.subplot(2, 1, 2)
plt.plot(sim.trange(), model.similarity(sim.data, mem))
plt.legend(fontsize='x-small')
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("memory")
plt.xlabel("time [s]");
- By making tau negative, we forced the recurrent connection to be positive
- So, whatever is in memory 'grows' over time
- This gives a 'primacy' effect
- i.e. Whatever you ran into first is what you remember
- You could also get this by changing connection weights
- Together, primacy and recency capture the most salient properties of human working memory
- You see these effects in both 'free recall' and 'serial recall'

- So far, the networks we've seen would work well for free recall presumably.
- How to include order information?
Serial Working Memory¶
An ordered list is a very simple kind of structure
To represent structures, we can use what we learned last time: vector binding
- Specifically, Semantic Pointers
In this case, we can do something like:
$Pos_0\circledast Item_0 + Pos_1\circledast Item_1 + ...$
- The $Item$ can just whatever vector we want to remember
- How should we generate $Pos$? What features does it need?
- It would be good if we could generate them on the fly, forever
- It would be good if $Pos_0$ "came before" $Pos_1$ in some sense
- There is a kind of vector that's ideal for this called a 'unitary vector'
- A unitary vector does not change length when convolved
- So, you can convolve forever
- The convolution of a unitary vector with itself is 'unlike' the original vector
- So, you can go fwd/backward with convolution/correlation
- A unitary vector does not change length when convolved
- This gives a way of 'counting' positions:
- $Pos_0 \circledast PlusOne = Pos_1$
- $Pos_1 \circledast PlusOne = Pos_2$
- $Pos_2 \circledast PlusOne = Pos_3$
- etc.
- And, $Pos_3 \circledast PlusOne' = Pos_2$
- We can put this representation together with primacy and recency, and we get something like this

- To do 'recall' from this representation, we want to remember what we've already recalled, so we can add a circuit like this

- This does a pretty good job of capturing lots of working memory data

- Order effects

- Error probability of items in the list (closer items are more likely to be incorrect, unless you change similarity between items)

- Item similarity effects

Arbitrary list lengths
This model can actually do lots more as well
- backwards recall, etc.
A slight variation does free recall
- you have to include non-bound item representations as well
You'll notice that it relies on a 'clean-up'
- This is a kind of long term memory
- We saw this show up before when talking about symbols
- How can we build such a thing in spiking neurons?
Long term memory¶
- In order for memories to be stable over the long term, they must be encoded in connection weights
- These kind of memories are often called 'associative memories'
- Auto-associative: recall the same thing you're shown
- e.g. for noise reduction
- pattern completion
- Hetero-associative: recall some arbitrary association
- e.g. for capturing domain relationships
- reasoning
- Auto-associative: recall the same thing you're shown
- Typical solutions in ANNs are
- Hopfield networks: A many-point attractor network
- Linear associators: Do SVD on the input/output to get a weight matrix
- Multi-layer Perceptron (MLP): Use backprop to train a network to compute the feedfwd mapping

Often worried about how to learn these
- We'll talk about learning in a later lecture
- Often the learning is so un-'biologically plausible' that it's best thought of as an optimization (like least squares in the NEF).
How to compute these kinds of mappings with the NEF/SPA?
- We want to 'recognize' specific vectors (in a vocabulary)
- We want only some neurons to fire when a given vector is present (sparse)
- We want to activate a given output vector if the input is recognized
Can do it in one layer with a feedforward approach
- Set the encoding vectors of some small set of neurons to be a vocabulary item
- That will pick out specific vectors
- Set the intercepts of neurons to be high in the positive direction
- So they will only fire if there is 'enough' of that vector in the input
- Set the linear transformation on the output of those neurons to be the desired output vector
- Since these are in the weights, the output will perfectly represent the desired output
- Set the encoding vectors of some small set of neurons to be a vocabulary item
We've seen several techniques that will make this easy
- Ensemble arrays: For collecting lots of small populations together
- SPA module: For defining vocabs, etc.
import nengo
from nengo import spa
seed=1
np.random.seed(seed)
model = spa.SPA("Associative Memory", seed=seed)
D = 32
vocab = spa.Vocabulary(D)
vocab.parse('BLUE+GREEN+RED')
noise_RED = vocab.parse("RED").v + .2*np.random.randn(D)
noise_RED = noise_RED/np.linalg.norm(noise_RED)
noise_GREEN = vocab.parse("GREEN").v + .2*np.random.randn(D)
noise_GREEN = noise_GREEN/np.linalg.norm(noise_GREEN)
def memory_input(t):
if t < 0.2:
return vocab.parse("BLUE").v
elif .2 < t < .5:
return noise_RED
elif .5 < t < .8:
return vocab.parse("RED").v
elif .8 < t < 1:
return noise_GREEN
else:
return vocab.parse("0").v
with model:
stim = nengo.Node(output=memory_input, label='input')
model.am = spa.AssociativeMemory(vocab)
nengo.Connection(stim, model.am.input)
in_p = nengo.Probe(stim)
out_p = nengo.Probe(model.am.output, synapse=0.03)
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/simple_cleanup.py.cfg")
sim = nengo.Simulator(model)
sim.run(1)
t = sim.trange()
figure(figsize=(10,10))
plt.subplot(2, 1, 1)
plt.plot(t, spa.similarity(sim.data[in_p], vocab))
plt.ylabel("Input")
plt.ylim(top=1.1)
plt.legend(vocab.keys, loc='best')
plt.subplot(2, 1, 2)
plt.plot(t, nengo.spa.similarity(sim.data[out_p], vocab))
plt.ylabel("Output")
plt.legend(vocab.keys, loc='best');
- This implementation has several nice features
- Extremely fast (1 synapse feedforward ~ 5ms)
- Fully spiking, integrates with SPA models
- Scales very well

- Scaling properties of a neural clean-up memory.
- A SP is formed by binding $k$ pairs of random SPs and adding
- Binding to random probes from that set.
- Figure shows the minimum number of dimensions required to recover a lexical item from the input 99% of the time.
- Averages over 200 simulations for each combination of $k$, $M$, and $D$ values.
- Vertical dashed line is approximate size of an adult lexicon.
- Horizontal dashed lines show the performance of a non-neural clean-up directly implementing the algorithm.
SPA Example¶
- This is the same example as for the symbols lecture, but with a working memory
- The WM is just a single attractor network
- The model is supposed to memorize the bound inputs and then answer questions about what is bound to what
%pylab inline
import nengo
from nengo import spa
def color_input(t):
if t < 0.25:
return 'RED'
elif t < 0.5:
return 'BLUE'
else:
return '0'
def shape_input(t):
if t < 0.25:
return 'CIRCLE'
elif t < 0.5:
return 'SQUARE'
else:
return '0'
def cue_input(t):
if t < 0.5:
return '0'
sequence = ['0', 'CIRCLE', 'RED', '0', 'SQUARE', 'BLUE']
idx = int(((t - 0.5) // (1. / len(sequence))) % len(sequence))
return sequence[idx]
seed=1
model = spa.SPA(label="Simple question answering", seed=seed)
dimensions = 32
vocab = model.get_default_vocab(dimensions)
vocab.parse('BLUE+RED+CIRCLE+SQUARE')
with model:
model.color_in = spa.Buffer(dimensions=dimensions)
model.shape_in = spa.Buffer(dimensions=dimensions)
model.conv = spa.Memory(dimensions=dimensions, subdimensions=4, synapse=0.4)
model.cue = spa.Buffer(dimensions=dimensions)
model.out = spa.Buffer(dimensions=dimensions)
model.am = spa.AssociativeMemory(vocab, threshold=0.1)
model.inp = spa.Input(color_in=color_input, shape_in=shape_input, cue=cue_input)
# Connect the buffers
cortical_actions = spa.Actions(
'conv = color_in * shape_in',
'out = conv * ~cue',
'am = out'
)
model.cortical = spa.Cortical(cortical_actions)
model.config[nengo.Probe].synapse = nengo.Lowpass(0.03)
color_in = nengo.Probe(model.color_in.state.output)
shape_in = nengo.Probe(model.shape_in.state.output)
cue = nengo.Probe(model.cue.state.output)
conv = nengo.Probe(model.conv.state.output)
out = nengo.Probe(model.out.state.output)
clean = nengo.Probe(model.am.output)
WARNING: pylab import has clobbered these variables: ['seed', 'piecewise'] `%matplotlib` prevents importing * from pylab and numpy
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/binding_with_memory.py.cfg")
sim = nengo.Simulator(model)
sim.run(3.)
plt.figure(figsize=(12, 10))
plt.subplot(6, 1, 1)
plt.plot(sim.trange(), model.similarity(sim.data, color_in))
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("color")
plt.subplot(6, 1, 2)
plt.plot(sim.trange(), model.similarity(sim.data, shape_in))
plt.legend(model.get_output_vocab('shape_in').keys, fontsize='x-small')
plt.ylabel("shape")
plt.subplot(6, 1, 3)
plt.plot(sim.trange(), model.similarity(sim.data, cue))
plt.legend(model.get_output_vocab('cue').keys, fontsize='x-small')
plt.ylabel("cue")
plt.subplot(6, 1, 4)
for pointer in ['RED * CIRCLE', 'BLUE * SQUARE']:
plt.plot(sim.trange(), vocab.parse(pointer).dot(sim.data[conv].T), label=pointer)
plt.legend(fontsize='x-small')
plt.ylabel("convolved")
plt.subplot(6, 1, 5)
plt.plot(sim.trange(), spa.similarity(sim.data[out], vocab))
plt.legend(model.get_output_vocab('out').keys, fontsize='x-small')
plt.ylabel("Output")
plt.xlabel("time [s]");
plt.subplot(6, 1, 6)
plt.plot(sim.trange(), spa.similarity(sim.data[clean], vocab))
plt.legend(model.get_output_vocab('am').keys, fontsize='x-small')
plt.ylabel("Cleaned Up Output")
plt.xlabel("time [s]");