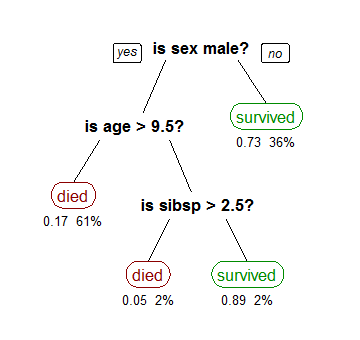In [1]:
import mglearn # credits to Muller and Guido (https://www.amazon.com/dp/1449369413/)
import matplotlib.pyplot as plt
%matplotlib inline
mglearn.plots.plot_tree_not_monotone()
Feature importances: [ 0. 1.]
Out[1]:
In [2]:
from sklearn.datasets import load_breast_cancer
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
cancer = load_breast_cancer()
X_train, X_test, y_train, y_test = train_test_split(cancer.data, cancer.target, stratify=cancer.target, random_state=42)
tree = DecisionTreeClassifier(random_state=0)
tree.fit(X_train, y_train)
print('Accuracy on the training subset: {:.3f}'.format(tree.score(X_train, y_train)))
print('Accuracy on the test subset: {:.3f}'.format(tree.score(X_test, y_test)))
Accuracy on the training subset: 1.000 Accuracy on the test subset: 0.937
In [3]:
tree = DecisionTreeClassifier(max_depth=4, random_state=0)
tree.fit(X_train, y_train)
print('Accuracy on the training subset: {:.3f}'.format(tree.score(X_train, y_train)))
print('Accuracy on the test subset: {:.3f}'.format(tree.score(X_test, y_test)))
Accuracy on the training subset: 0.988 Accuracy on the test subset: 0.951
In [4]:
import graphviz
from sklearn.tree import export_graphviz
export_graphviz(tree, out_file='cancertree.dot', class_names=['malignant', 'benign'], feature_names=cancer.feature_names,
impurity=False, filled=True)

In [ ]: