Linearly Inseparable Datasets¶
The Non-Convex Regions¶
Non-Convex Regions¶
- What are non-convex regions?
Linearly Inseparable Regions¶
- What are linearly inseparable regions?
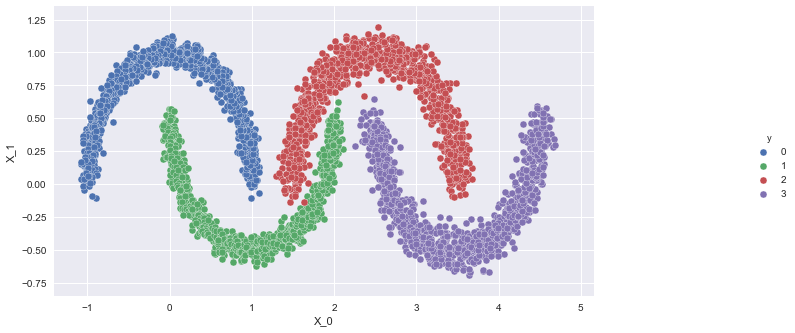
Here is an image of linearly inseparable, non-convex regions that we would like to identify by clustering.
Let us look at just the data points:
## Exercise:
Let us now create this dataset of 4 almonds using half-moons.
- Create a dataframe, moon_df with the structure given the X, y.
| X_0 | X_1 | y |
| X[:,0] | X[:,1] | y |
In [2]:
from matplotlib import pyplot as plt
from sklearn.datasets import make_moons
import numpy as np
import seaborn as sns
import pandas as pd
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
N_Samples = 1000
D = 2
K = 4
X, y = make_moons(n_samples = 2*N_Samples, noise=0.05, shuffle = False)
x_vec, y_vec = make_moons(n_samples = 2*N_Samples, noise=0.08, shuffle = False)
x_vec[:,0] += 2.5
y_vec += 2
X = np.concatenate((X, x_vec), axis=0)
y = np.concatenate((y, y_vec), axis=0)
use the pd.DataFrame() command and sns.pairplot(x_vars, y_vars, hue, data)
In [ ]:
moon_df = pd.DataFrame({'X_0':X[:,0], 'X_1':X[:,1], 'y':y})
g = sns.pairplot(x_vars="X_0", y_vars="X_1", hue="y", data = moon_df)
g.fig.set_size_inches(14, 6)
sns.despine()
In [ ]:
ref_tmp_var = False
try:
ref_assert_var = False
moon_df_ = pd.DataFrame({'X_0':X[:,0], 'X_1':X[:,1], 'y':y})
import numpy as np
if np.all(moon_df['X_0'] == moon_df_['X_0']) and np.all(moon_df['X_1'] == moon_df_['X_1']) and np.all(moon_df['y'] == moon_df_['y']):
ref_assert_var = True
out = g
else:
ref_assert_var = False
except Exception:
print('Please follow the instructions given and use the same variables provided in the instructions.')
else:
if ref_assert_var:
ref_tmp_var = True
else:
print('Please follow the instructions given and use the same variables provided in the instructions.')
assert ref_tmp_var
Linearly Inseparable Datasets¶
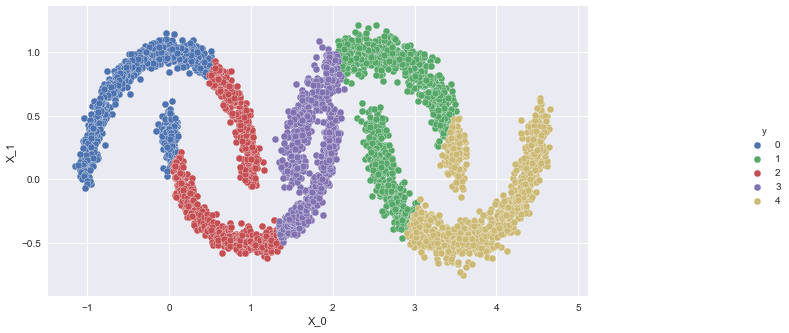
Applying k-means to the dataset:We see that the k-means failed. Let us now try GMMs
## Exercise:
- Apply GMMs on the dataset and visualize the graph, g.
- Assign the cluster assignments of each data point to gmm_cluster column in the blob_df dataframe.
In [4]:
import matplotlib as mpl
from matplotlib import pyplot as plt
from sklearn.datasets import make_moons
import numpy as np
import seaborn as sns
import pandas as pd
from sklearn import datasets
from sklearn import mixture
from sklearn.mixture import GaussianMixture
#Generate the half moon data-set (4-halfmoons)
N_Samples = 1000
X, y = make_moons(n_samples = 2*N_Samples, noise=0.05, shuffle = False)
x_vec, y_vec = make_moons(n_samples = 2*N_Samples, noise=0.08, shuffle = False)
x_vec[:,0] += 2.5
y_vec += 2
X = np.concatenate((X, x_vec), axis=0)
y = np.concatenate((y, y_vec), axis=0)
#visualizing using seaborn library
moon_df = pd.DataFrame({'X_0':X[:,0],'X_1':X[:,1], 'y':y})
Use the code from GMMs.
In [ ]:
mixture_model = GaussianMixture(n_components = 4, covariance_type = 'spherical')
mixture_model.fit(X)
moon_df['gmm_clus'] = mixture_model.predict(X)
# Plot the clusters
g = sns.pairplot(x_vars="X_0", y_vars="X_1", hue="gmm_clus", data = moon_df)
g.fig.set_size_inches(14, 6)
sns.despine()
In [ ]:
ref_tmp_var = False
try:
ref_assert_var = False
mixture_model_ = GaussianMixture(n_components = 4, covariance_type = 'spherical')
mixture_model_.fit(X)
y_hat = mixture_model_.predict(X)
import numpy as np
if len(y_hat) == len(moon_df['gmm_clus']):
ref_assert_var = True
out = g
else:
ref_assert_var = False
except Exception:
print('Please follow the instructions given and use the same variables provided in the instructions.')
else:
if ref_assert_var:
ref_tmp_var = True
else:
print('Please follow the instructions given and use the same variables provided in the instructions.')
assert ref_tmp_var