Yanouk's samples - Hypo and Hypermethylated loci (3x)¶
Instructions for Visualization¶
- Install IGV (free software) http://www.broadinstitute.org/software/igv/home
In [4]:
from IPython.display import HTML
HTML('<iframe src=http://www.broadinstitute.org/software/igv/home width=100% height=650></iframe>')
Out[4]:
- Open IGV
- Go to File > Load from URL...
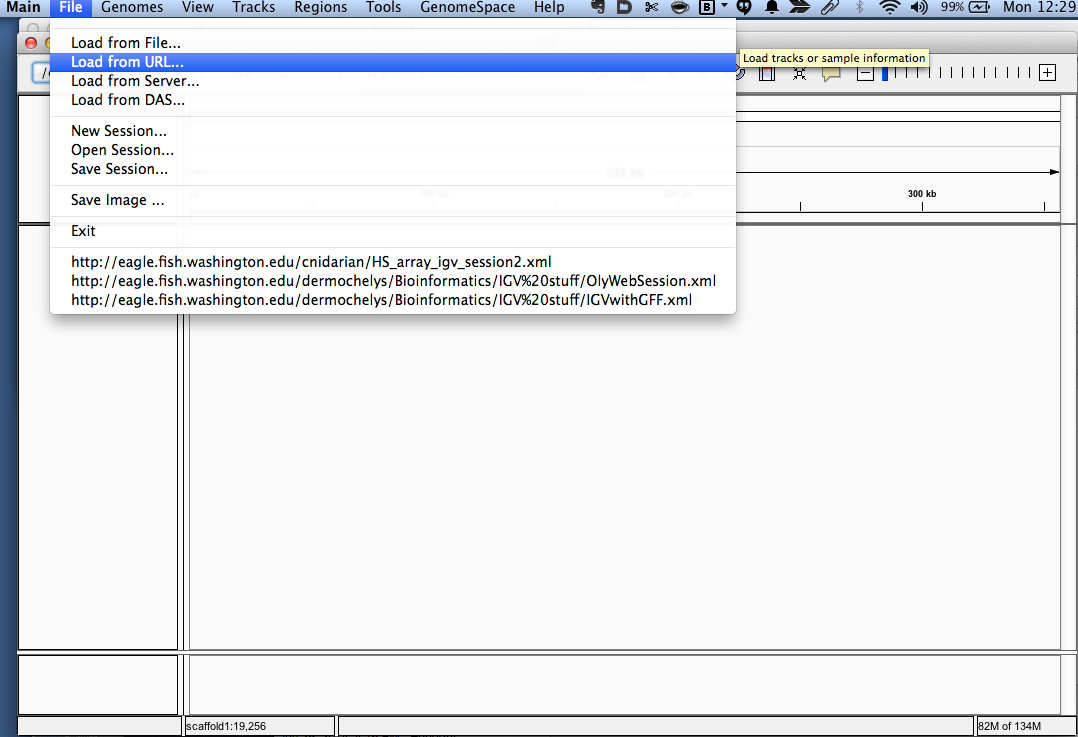
- Enter
http://eagle.fish.washington.edu/cnidarian/igv_session_YElarvae.xml
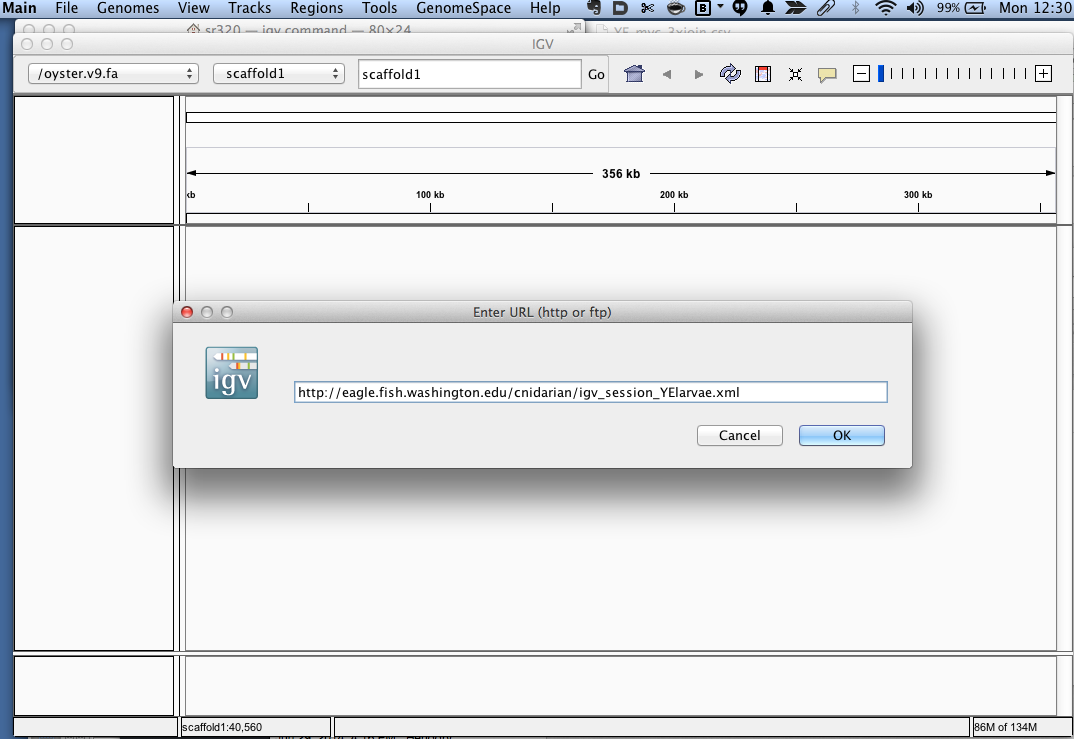
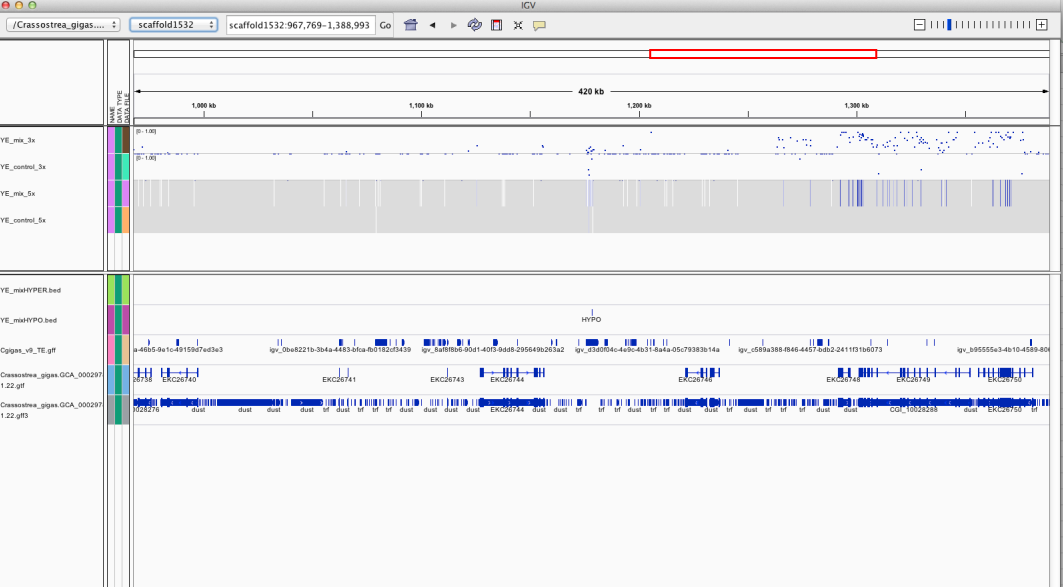
Voila- you should get a window similar to the what is at the top of the page.
Tracks include methylation data at 3x and 5x coverage and a track representing loci that are hypermethylated in mix library and hypomethylated in mix sample.
IGV xml to directly load all tracks above
http://eagle.fish.washington.edu/cnidarian/igv_session_YElarvae.xml
In [12]:
!cat /Volumes/web/cnidarian/igv_session_YElarvae.xml
<?xml version="1.0" encoding="UTF-8" standalone="no"?>
<Session genome="http://eagle.fish.washington.edu/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa" locus="scaffold334:356874-366720" version="5">
<Resources>
<Resource path="http://eagle.fish.washington.edu/cnidarian/filt_YE_mix_22smCG5x.igv"/>
<Resource path="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.gtf"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/YE_mixHYPER.bed"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/filt_YE_control_22smCG5x.igv"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/zearly_th_genomecov.bedgraph"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/filt_YE_control_22smCG3x.igv"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/filt_YE_mix_22smCG3x.igv"/>
<Resource path="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.gff3"/>
<Resource path="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE.gff"/>
<Resource path="http://eagle.fish.washington.edu/cnidarian/YE_mixHYPO.bed"/>
</Resources>
<Panel height="233" name="DataPanel" width="1133">
<Track altColor="0,0,178" autoScale="true" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="http://eagle.fish.washington.edu/cnidarian/filt_YE_mix_22smCG3x.igv_0.000" name="YE_mix_3x" normalize="false" renderer="BAR_CHART" sortable="true" visible="true" windowFunction="mean">
<DataRange baseline="0.0" drawBaseline="true" flipAxis="false" maximum="1.0" minimum="0.0" type="LINEAR"/>
</Track>
<Track altColor="0,0,178" autoScale="true" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="http://eagle.fish.washington.edu/cnidarian/filt_YE_control_22smCG3x.igv_0.000" name="YE_control_3x" normalize="false" renderer="BAR_CHART" sortable="true" visible="true" windowFunction="mean">
<DataRange baseline="0.0" drawBaseline="true" flipAxis="false" maximum="0.667" minimum="0.0" type="LINEAR"/>
</Track>
<Track altColor="0,0,178" autoScale="true" color="0,0,178" colorScale="ContinuousColorScale;0.0;1.0;255,255,255;0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="http://eagle.fish.washington.edu/cnidarian/filt_YE_mix_22smCG5x.igv_0.000" name="YE_mix_5x" normalize="false" renderer="BAR_CHART" sortable="true" visible="true" windowFunction="mean">
<DataRange baseline="0.0" drawBaseline="true" flipAxis="false" maximum="0.8" minimum="0.0" type="LINEAR"/>
</Track>
<Track altColor="0,0,178" autoScale="true" color="0,0,178" colorScale="ContinuousColorScale;0.0;1.0;255,255,255;0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="http://eagle.fish.washington.edu/cnidarian/filt_YE_control_22smCG5x.igv_0.000" name="YE_control_5x" normalize="false" renderer="BAR_CHART" sortable="true" visible="true" windowFunction="mean">
<DataRange baseline="0.0" drawBaseline="true" flipAxis="false" maximum="1.0" minimum="0.0" type="LINEAR"/>
</Track>
<Track altColor="0,0,178" autoScale="true" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="http://eagle.fish.washington.edu/cnidarian/zearly_th_genomecov.bedgraph" name="zearly_th_genomecov.bedgraph" normalize="false" renderer="BAR_CHART" sortable="true" visible="true" windowFunction="mean">
<DataRange baseline="0.0" drawBaseline="true" flipAxis="false" maximum="1725.0" minimum="0.0" type="LINEAR"/>
</Track>
</Panel>
<Panel height="615" name="FeaturePanel" width="1133">
<Track altColor="0,0,178" autoScale="false" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" id="Reference sequence" name="Reference sequence" sortable="false" visible="true"/>
<Track altColor="0,0,178" autoScale="false" clazz="org.broad.igv.track.FeatureTrack" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" height="45" id="http://eagle.fish.washington.edu/cnidarian/YE_mixHYPER.bed" name="YE_mixHYPER.bed" renderer="BASIC_FEATURE" sortable="false" visible="true" windowFunction="count"/>
<Track altColor="0,0,178" autoScale="false" clazz="org.broad.igv.track.FeatureTrack" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" height="45" id="http://eagle.fish.washington.edu/cnidarian/YE_mixHYPO.bed" name="YE_mixHYPO.bed" renderer="BASIC_FEATURE" sortable="false" visible="true" windowFunction="count"/>
<Track altColor="0,0,178" autoScale="false" clazz="org.broad.igv.track.FeatureTrack" color="0,0,178" displayMode="SQUISHED" featureVisibilityWindow="-1" fontSize="10" height="45" id="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_TE.gff" name="Cgigas_v9_TE.gff" renderer="GENE_TRACK" sortable="false" visible="true" windowFunction="count"/>
<Track altColor="0,0,178" autoScale="false" clazz="org.broad.igv.track.FeatureTrack" color="0,0,178" displayMode="COLLAPSED" featureVisibilityWindow="-1" fontSize="10" height="45" id="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.gtf" name="Crassostrea_gigas.GCA_000297895.1.22.gtf" renderer="GENE_TRACK" sortable="false" visible="true" windowFunction="count"/>
<Track altColor="0,0,178" autoScale="false" clazz="org.broad.igv.track.FeatureTrack" color="0,0,178" displayMode="EXPANDED" featureVisibilityWindow="-1" fontSize="10" height="45" id="http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.gff3" name="Crassostrea_gigas.GCA_000297895.1.22.gff3" renderer="GENE_TRACK" sortable="false" visible="true" windowFunction="count"/>
</Panel>
<PanelLayout dividerFractions="0.39965986394557823"/>
</Session>
Adding some larvae RNA-seq Tracks¶
In [8]:
bgdir="/iplant/home/sr320/Cgigas_v9/Zhang/bgraph"
!ils {bgdir}
/iplant/home/sr320/Cgigas_v9/Zhang/bgraph: C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_E-2014-03-08-07-36-07.971 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_early-2014-07-08-09-17-15.654 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_Fgo-2014-03-07-10-07-12.617 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_G3-2014-03-06-16-26-41.026 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_late-2014-07-08-09-15-42.258 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_Man1-2014-07-02-08-36-42.069 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_Mgo1.3-2014-03-07-14-06-07.430 C- /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/RNAseq2bedgraph_Mgo-2014-03-07-14-03-33.677
In [9]:
folder="RNAseq2bedgraph_late-2014-07-08-09-15-42.258"
id="late"
In [10]:
cd /Volumes/web/cnidarian/
/Volumes/web/cnidarian
In [ ]:
!icd /iplant/home/sr320/Cgigas_v9/Zhang/bgraph/{folder}
!imv accepted_hits.bam z{id}_tophat.bam
!imv bedtools_genomecov_output z{id}_th_genomecov.bedgraph
!iget -f z{id}_th_genomecov.bedgraph
!iget -f z{id}_tophat.bam
!igvtools toTDF z{id}_th_genomecov.bedgraph z{id}_th_genomecov.bedgraph.tdf /Volumes/web/cnidarian/cgigas_v9_genome03.chrom.sizes
!iput z{id}_th_genomecov.bedgraph.tdf
In [ ]: