DNA Methylation during early development in oysters¶
This notebook includes procedures for analyzing data desribed in --- In theory anyone should be able to reproduce our work in IPython, if they have the following software installed.
- BSMAP
- R module for IPython
- SQLShare python client
The general premise is you simply provide minimal information including a working directory and location of software. The notebook is divided into X sections
0
At the beginning of each section specfic instructions on user actions will be described. Please post any comments and questions in issues .
In [414]:
!rsync -avz /Volumes/Bay3_scratch/BiGo_manny /Volumes/web/whale
building file list ... done BiGo_manny/ BiGo_manny/BiGo_lar_fastq.tgz ^C
In [13]:
#set locaiton of sqlshare python
pt="/Users/sr320/sqlshare-pythonclient/tools/"
Downloading Files¶
In [3]:
#The first thing you will need to do is set a working directory. Simply include the path to your working directory within quotes.
lwd="/Volumes/Bay3_scratch/BiGo_manny"
In [4]:
cd {lwd}
/Volumes/Bay3_scratch/BiGo_manny
In [ ]:
#This command downloads a archived file including six BS-seq libraries.
!wget http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_HTSdata/BiGo_larvae_cc/BiGo_lar_fastq.tgz
In [3]:
ls
BiGo_lar_fastq.tgz
In [424]:
!ls mcf*
mcf_M1_R1.fastq mcf_M3_R2.fastq mcf_T1D5_R1.fastq mcf_T3D3_R2.fastq mcf_M1_R2.fastq mcf_T1D3_R1.fastq mcf_T1D5_R2.fastq mcf_T3D5_R1.fastq mcf_M3_R1.fastq mcf_T1D3_R2.fastq mcf_T3D3_R1.fastq mcf_T3D5_R2.fastq
In [425]:
!tar -zcvf BiGo_lar_fastq_mcf.tgz mcf*
a mcf_M1_R1.fastq a mcf_M1_R2.fastq a mcf_M3_R1.fastq a mcf_M3_R2.fastq a mcf_T1D3_R1.fastq a mcf_T1D3_R2.fastq a mcf_T1D5_R1.fastq a mcf_T1D5_R2.fastq a mcf_T3D3_R1.fastq a mcf_T3D3_R2.fastq a mcf_T3D5_R1.fastq a mcf_T3D5_R2.fastq
In [6]:
#oyster genome
!wget http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa
--2014-07-08 10:20:04-- http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_ensembl_tracks/Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa Resolving eagle.fish.washington.edu... 128.95.149.81 Connecting to eagle.fish.washington.edu|128.95.149.81|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 567608307 (541M) [text/plain] Saving to: `Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa' 100%[======================================>] 567,608,307 10.7M/s in 51s 2014-07-08 10:20:56 (10.5 MB/s) - `Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa' saved [567608307/567608307]
Running BSMAP¶
In [5]:
#setting some variables for files but also
#location of BSMAP (if not in PATH)
bsmaploc="/Volumes/Bay3/Software/BSMAP/bsmap-2.74/"
Another option is running BSMAP on iPlant
In [6]:
lib="M1"
In [7]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 2
BSMAP v2.74 Start at: Wed Jul 9 15:11:17 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 18 secs passed total_kmers: 43046721 Create seed table. 70 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(2 threads) Input read file #1: mcf_M1_R1.fastq (format: FASTQ) Input read file #2: mcf_M1_R2.fastq (format: FASTQ) Output file: bsmap_out_M1.sam (format: SAM) Thread #0: 50000 read pairs finished. 100 secs passed Thread #1: 100000 read pairs finished. 100 secs passed Thread #0: 150000 read pairs finished. 128 secs passed Thread #1: 200000 read pairs finished. 128 secs passed Thread #0: 250000 read pairs finished. 157 secs passed Thread #1: 300000 read pairs finished. 158 secs passed Thread #0: 350000 read pairs finished. 183 secs passed Thread #1: 400000 read pairs finished. 183 secs passed Thread #0: 450000 read pairs finished. 207 secs passed Thread #1: 500000 read pairs finished. 208 secs passed Thread #0: 550000 read pairs finished. 233 secs passed Thread #1: 600000 read pairs finished. 234 secs passed Thread #0: 650000 read pairs finished. 264 secs passed Thread #1: 700000 read pairs finished. 265 secs passed Thread #0: 750000 read pairs finished. 291 secs passed Thread #1: 800000 read pairs finished. 292 secs passed Thread #0: 850000 read pairs finished. 317 secs passed Thread #1: 900000 read pairs finished. 318 secs passed Thread #0: 950000 read pairs finished. 343 secs passed Thread #1: 1000000 read pairs finished. 344 secs passed Thread #0: 1050000 read pairs finished. 367 secs passed Thread #1: 1100000 read pairs finished. 368 secs passed Thread #0: 1150000 read pairs finished. 392 secs passed Thread #1: 1200000 read pairs finished. 394 secs passed Thread #0: 1250000 read pairs finished. 419 secs passed Thread #1: 1300000 read pairs finished. 420 secs passed Thread #0: 1350000 read pairs finished. 451 secs passed Thread #1: 1400000 read pairs finished. 454 secs passed Thread #0: 1450000 read pairs finished. 481 secs passed Thread #1: 1500000 read pairs finished. 483 secs passed Thread #0: 1550000 read pairs finished. 503 secs passed Thread #1: 1600000 read pairs finished. 505 secs passed Thread #0: 1650000 read pairs finished. 526 secs passed Thread #1: 1700000 read pairs finished. 527 secs passed Thread #0: 1750000 read pairs finished. 549 secs passed Thread #1: 1800000 read pairs finished. 550 secs passed Thread #0: 1850000 read pairs finished. 571 secs passed Thread #1: 1900000 read pairs finished. 572 secs passed Thread #0: 1950000 read pairs finished. 593 secs passed Thread #1: 2000000 read pairs finished. 595 secs passed Thread #0: 2050000 read pairs finished. 617 secs passed Thread #1: 2100000 read pairs finished. 618 secs passed Thread #0: 2150000 read pairs finished. 639 secs passed Thread #1: 2200000 read pairs finished. 640 secs passed Thread #0: 2250000 read pairs finished. 662 secs passed Thread #1: 2300000 read pairs finished. 663 secs passed Thread #0: 2350000 read pairs finished. 687 secs passed Thread #1: 2400000 read pairs finished. 689 secs passed Thread #0: 2450000 read pairs finished. 709 secs passed Thread #1: 2500000 read pairs finished. 711 secs passed Thread #0: 2550000 read pairs finished. 731 secs passed Thread #1: 2600000 read pairs finished. 733 secs passed Thread #0: 2650000 read pairs finished. 754 secs passed Thread #1: 2700000 read pairs finished. 755 secs passed Thread #0: 2750000 read pairs finished. 777 secs passed Thread #1: 2800000 read pairs finished. 778 secs passed Thread #0: 2850000 read pairs finished. 799 secs passed Thread #1: 2900000 read pairs finished. 800 secs passed Thread #0: 2950000 read pairs finished. 824 secs passed Thread #1: 3000000 read pairs finished. 827 secs passed Thread #0: 3050000 read pairs finished. 856 secs passed Thread #1: 3100000 read pairs finished. 857 secs passed Thread #0: 3150000 read pairs finished. 878 secs passed Thread #1: 3200000 read pairs finished. 879 secs passed Thread #0: 3250000 read pairs finished. 899 secs passed Thread #1: 3300000 read pairs finished. 901 secs passed Thread #0: 3350000 read pairs finished. 922 secs passed Thread #1: 3400000 read pairs finished. 923 secs passed Thread #0: 3450000 read pairs finished. 943 secs passed Thread #1: 3500000 read pairs finished. 944 secs passed Thread #0: 3550000 read pairs finished. 966 secs passed Thread #1: 3600000 read pairs finished. 968 secs passed Thread #0: 3650000 read pairs finished. 990 secs passed Thread #1: 3700000 read pairs finished. 991 secs passed Thread #0: 3750000 read pairs finished. 1013 secs passed Thread #1: 3800000 read pairs finished. 1015 secs passed Thread #0: 3850000 read pairs finished. 1040 secs passed Thread #1: 3900000 read pairs finished. 1042 secs passed Thread #0: 3950000 read pairs finished. 1067 secs passed Thread #1: 4000000 read pairs finished. 1068 secs passed Thread #0: 4050000 read pairs finished. 1090 secs passed Thread #1: 4100000 read pairs finished. 1091 secs passed Thread #0: 4150000 read pairs finished. 1111 secs passed Thread #1: 4200000 read pairs finished. 1112 secs passed Thread #0: 4250000 read pairs finished. 1133 secs passed Thread #1: 4300000 read pairs finished. 1133 secs passed Thread #0: 4350000 read pairs finished. 1158 secs passed Thread #1: 4400000 read pairs finished. 1159 secs passed Thread #0: 4450000 read pairs finished. 1181 secs passed Thread #1: 4500000 read pairs finished. 1182 secs passed Thread #0: 4550000 read pairs finished. 1204 secs passed Thread #1: 4600000 read pairs finished. 1205 secs passed Thread #0: 4650000 read pairs finished. 1226 secs passed Thread #1: 4700000 read pairs finished. 1227 secs passed Thread #0: 4750000 read pairs finished. 1247 secs passed Thread #1: 4800000 read pairs finished. 1248 secs passed Thread #0: 4850000 read pairs finished. 1270 secs passed Thread #1: 4900000 read pairs finished. 1271 secs passed Thread #0: 4950000 read pairs finished. 1292 secs passed Thread #1: 5000000 read pairs finished. 1293 secs passed Thread #0: 5050000 read pairs finished. 1313 secs passed Thread #1: 5100000 read pairs finished. 1314 secs passed Thread #0: 5150000 read pairs finished. 1332 secs passed Thread #1: 5200000 read pairs finished. 1334 secs passed Thread #0: 5250000 read pairs finished. 1352 secs passed Thread #1: 5300000 read pairs finished. 1355 secs passed Thread #0: 5350000 read pairs finished. 1373 secs passed Thread #1: 5400000 read pairs finished. 1375 secs passed Thread #0: 5450000 read pairs finished. 1393 secs passed Thread #1: 5500000 read pairs finished. 1395 secs passed Thread #0: 5550000 read pairs finished. 1415 secs passed Thread #1: 5600000 read pairs finished. 1419 secs passed Thread #0: 5650000 read pairs finished. 1437 secs passed Thread #1: 5700000 read pairs finished. 1439 secs passed Thread #0: 5750000 read pairs finished. 1456 secs passed Thread #1: 5800000 read pairs finished. 1459 secs passed Thread #0: 5850000 read pairs finished. 1476 secs passed Thread #1: 5900000 read pairs finished. 1478 secs passed Thread #0: 5950000 read pairs finished. 1500 secs passed Thread #1: 6000000 read pairs finished. 1501 secs passed Thread #0: 6050000 read pairs finished. 1525 secs passed Thread #1: 6100000 read pairs finished. 1527 secs passed Thread #0: 6150000 read pairs finished. 1546 secs passed Thread #1: 6200000 read pairs finished. 1548 secs passed Thread #0: 6250000 read pairs finished. 1568 secs passed Thread #1: 6300000 read pairs finished. 1570 secs passed Thread #0: 6350000 read pairs finished. 1619 secs passed Thread #1: 6400000 read pairs finished. 1620 secs passed Thread #0: 6450000 read pairs finished. 1648 secs passed Thread #1: 6500000 read pairs finished. 1649 secs passed Thread #0: 6550000 read pairs finished. 1675 secs passed Thread #1: 6600000 read pairs finished. 1676 secs passed Thread #0: 6650000 read pairs finished. 1702 secs passed Thread #1: 6700000 read pairs finished. 1704 secs passed Thread #0: 6750000 read pairs finished. 1733 secs passed Thread #1: 6800000 read pairs finished. 1735 secs passed Thread #0: 6850000 read pairs finished. 1756 secs passed Thread #1: 6900000 read pairs finished. 1759 secs passed Thread #0: 6950000 read pairs finished. 1786 secs passed Thread #1: 7000000 read pairs finished. 1788 secs passed Thread #0: 7050000 read pairs finished. 1815 secs passed Thread #1: 7100000 read pairs finished. 1817 secs passed Thread #0: 7150000 read pairs finished. 1839 secs passed Thread #1: 7200000 read pairs finished. 1840 secs passed Thread #0: 7250000 read pairs finished. 1864 secs passed Thread #1: 7300000 read pairs finished. 1865 secs passed Thread #0: 7350000 read pairs finished. 1890 secs passed Thread #1: 7400000 read pairs finished. 1891 secs passed Thread #0: 7450000 read pairs finished. 1913 secs passed Thread #1: 7500000 read pairs finished. 1913 secs passed Thread #0: 7550000 read pairs finished. 1936 secs passed Thread #1: 7600000 read pairs finished. 1937 secs passed Thread #0: 7650000 read pairs finished. 1957 secs passed Thread #1: 7700000 read pairs finished. 1958 secs passed Thread #0: 7750000 read pairs finished. 1983 secs passed Thread #1: 7800000 read pairs finished. 1985 secs passed Thread #0: 7850000 read pairs finished. 2005 secs passed Thread #1: 7900000 read pairs finished. 2006 secs passed Thread #0: 7950000 read pairs finished. 2025 secs passed Thread #1: 8000000 read pairs finished. 2026 secs passed Thread #0: 8023643 read pairs finished. 2035 secs passed Total number of aligned reads: pairs: 3982552 (50%) single a: 1489441 (19%) single b: 1430645 (18%) Done. Finished at Wed Jul 9 15:45:12 2014 Total time consumed: 2035 secs
In [8]:
lib="T1D3"
In [9]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 4
BSMAP v2.74 Start at: Wed Jul 9 15:45:14 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 15 secs passed total_kmers: 43046721 Create seed table. 61 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: mcf_T1D3_R1.fastq (format: FASTQ) Input read file #2: mcf_T1D3_R2.fastq (format: FASTQ) Output file: bsmap_out_T1D3.sam (format: SAM) Thread #2: 50000 read pairs finished. 97 secs passed Thread #0: 100000 read pairs finished. 98 secs passed Thread #1: 200000 read pairs finished. 99 secs passed Thread #3: 150000 read pairs finished. 99 secs passed Thread #2: 250000 read pairs finished. 129 secs passed Thread #0: 300000 read pairs finished. 130 secs passed Thread #1: 350000 read pairs finished. 131 secs passed Thread #3: 400000 read pairs finished. 132 secs passed Thread #2: 450000 read pairs finished. 155 secs passed Thread #0: 500000 read pairs finished. 157 secs passed Thread #1: 550000 read pairs finished. 157 secs passed Thread #3: 600000 read pairs finished. 158 secs passed Thread #2: 650000 read pairs finished. 178 secs passed Thread #0: 700000 read pairs finished. 181 secs passed Thread #1: 750000 read pairs finished. 181 secs passed Thread #3: 800000 read pairs finished. 182 secs passed Thread #2: 850000 read pairs finished. 201 secs passed Thread #0: 900000 read pairs finished. 205 secs passed Thread #1: 950000 read pairs finished. 205 secs passed Thread #3: 1000000 read pairs finished. 207 secs passed Thread #2: 1050000 read pairs finished. 227 secs passed Thread #0: 1100000 read pairs finished. 231 secs passed Thread #1: 1150000 read pairs finished. 232 secs passed Thread #3: 1200000 read pairs finished. 233 secs passed Thread #2: 1250000 read pairs finished. 252 secs passed Thread #0: 1300000 read pairs finished. 257 secs passed Thread #1: 1350000 read pairs finished. 257 secs passed Thread #3: 1400000 read pairs finished. 258 secs passed Thread #2: 1450000 read pairs finished. 276 secs passed Thread #0: 1500000 read pairs finished. 280 secs passed Thread #1: 1550000 read pairs finished. 280 secs passed Thread #3: 1600000 read pairs finished. 281 secs passed Thread #2: 1650000 read pairs finished. 299 secs passed Thread #0: 1700000 read pairs finished. 303 secs passed Thread #1: 1750000 read pairs finished. 303 secs passed Thread #3: 1800000 read pairs finished. 304 secs passed Thread #2: 1850000 read pairs finished. 322 secs passed Thread #0: 1900000 read pairs finished. 325 secs passed Thread #1: 1950000 read pairs finished. 326 secs passed Thread #3: 2000000 read pairs finished. 327 secs passed Thread #2: 2050000 read pairs finished. 344 secs passed Thread #0: 2100000 read pairs finished. 348 secs passed Thread #1: 2150000 read pairs finished. 348 secs passed Thread #3: 2200000 read pairs finished. 349 secs passed Thread #2: 2250000 read pairs finished. 366 secs passed Thread #0: 2300000 read pairs finished. 370 secs passed Thread #1: 2350000 read pairs finished. 370 secs passed Thread #3: 2400000 read pairs finished. 371 secs passed Thread #2: 2450000 read pairs finished. 388 secs passed Thread #0: 2500000 read pairs finished. 392 secs passed Thread #1: 2550000 read pairs finished. 393 secs passed Thread #3: 2600000 read pairs finished. 394 secs passed Thread #2: 2650000 read pairs finished. 410 secs passed Thread #0: 2700000 read pairs finished. 414 secs passed Thread #1: 2750000 read pairs finished. 415 secs passed Thread #3: 2800000 read pairs finished. 415 secs passed Thread #2: 2850000 read pairs finished. 432 secs passed Thread #0: 2900000 read pairs finished. 436 secs passed Thread #1: 2950000 read pairs finished. 437 secs passed Thread #3: 3000000 read pairs finished. 438 secs passed Thread #2: 3050000 read pairs finished. 456 secs passed Thread #0: 3100000 read pairs finished. 460 secs passed Thread #1: 3150000 read pairs finished. 460 secs passed Thread #3: 3200000 read pairs finished. 461 secs passed Thread #2: 3250000 read pairs finished. 478 secs passed Thread #0: 3300000 read pairs finished. 482 secs passed Thread #1: 3350000 read pairs finished. 483 secs passed Thread #3: 3400000 read pairs finished. 484 secs passed Thread #2: 3450000 read pairs finished. 500 secs passed Thread #0: 3500000 read pairs finished. 505 secs passed Thread #1: 3550000 read pairs finished. 506 secs passed Thread #3: 3600000 read pairs finished. 507 secs passed Thread #2: 3650000 read pairs finished. 523 secs passed Thread #0: 3700000 read pairs finished. 527 secs passed Thread #1: 3750000 read pairs finished. 528 secs passed Thread #3: 3800000 read pairs finished. 529 secs passed Thread #2: 3850000 read pairs finished. 545 secs passed Thread #0: 3900000 read pairs finished. 549 secs passed Thread #1: 3950000 read pairs finished. 550 secs passed Thread #3: 4000000 read pairs finished. 552 secs passed Thread #2: 4050000 read pairs finished. 569 secs passed Thread #0: 4100000 read pairs finished. 573 secs passed Thread #1: 4150000 read pairs finished. 573 secs passed Thread #3: 4200000 read pairs finished. 575 secs passed Thread #2: 4250000 read pairs finished. 591 secs passed Thread #0: 4300000 read pairs finished. 595 secs passed Thread #1: 4350000 read pairs finished. 595 secs passed Thread #3: 4400000 read pairs finished. 597 secs passed Thread #2: 4450000 read pairs finished. 614 secs passed Thread #0: 4500000 read pairs finished. 618 secs passed Thread #1: 4550000 read pairs finished. 619 secs passed Thread #3: 4600000 read pairs finished. 621 secs passed Thread #2: 4650000 read pairs finished. 638 secs passed Thread #1: 4750000 read pairs finished. 641 secs passed Thread #0: 4700000 read pairs finished. 641 secs passed Thread #3: 4800000 read pairs finished. 643 secs passed Thread #2: 4850000 read pairs finished. 660 secs passed Thread #1: 4900000 read pairs finished. 664 secs passed Thread #0: 4950000 read pairs finished. 664 secs passed Thread #3: 5000000 read pairs finished. 665 secs passed Thread #2: 5016400 read pairs finished. 667 secs passed Total number of aligned reads: pairs: 2717832 (54%) single a: 1117376 (22%) single b: 1178713 (23%) Done. Finished at Wed Jul 9 15:56:21 2014 Total time consumed: 667 secs
In [10]:
lib="T1D5"
In [11]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 4
BSMAP v2.74 Start at: Wed Jul 9 15:56:22 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 13 secs passed total_kmers: 43046721 Create seed table. 52 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: mcf_T1D5_R1.fastq (format: FASTQ) Input read file #2: mcf_T1D5_R2.fastq (format: FASTQ) Output file: bsmap_out_T1D5.sam (format: SAM) Thread #1: 50000 read pairs finished. 82 secs passed Thread #2: 100000 read pairs finished. 83 secs passed Thread #0: 150000 read pairs finished. 84 secs passed Thread #3: 200000 read pairs finished. 85 secs passed Thread #1: 250000 read pairs finished. 109 secs passed Thread #2: 300000 read pairs finished. 110 secs passed Thread #0: 350000 read pairs finished. 111 secs passed Thread #3: 400000 read pairs finished. 112 secs passed Thread #1: 450000 read pairs finished. 136 secs passed Thread #2: 500000 read pairs finished. 136 secs passed Thread #0: 550000 read pairs finished. 137 secs passed Thread #3: 600000 read pairs finished. 138 secs passed Thread #1: 650000 read pairs finished. 164 secs passed Thread #2: 700000 read pairs finished. 165 secs passed Thread #0: 750000 read pairs finished. 166 secs passed Thread #3: 800000 read pairs finished. 167 secs passed Thread #1: 850000 read pairs finished. 193 secs passed Thread #2: 900000 read pairs finished. 193 secs passed Thread #0: 950000 read pairs finished. 194 secs passed Thread #3: 1000000 read pairs finished. 195 secs passed Thread #1: 1050000 read pairs finished. 220 secs passed Thread #2: 1100000 read pairs finished. 221 secs passed Thread #0: 1150000 read pairs finished. 221 secs passed Thread #3: 1200000 read pairs finished. 222 secs passed Thread #1: 1250000 read pairs finished. 247 secs passed Thread #2: 1300000 read pairs finished. 248 secs passed Thread #0: 1350000 read pairs finished. 249 secs passed Thread #3: 1400000 read pairs finished. 249 secs passed Thread #1: 1450000 read pairs finished. 274 secs passed Thread #2: 1500000 read pairs finished. 275 secs passed Thread #0: 1550000 read pairs finished. 275 secs passed Thread #3: 1600000 read pairs finished. 276 secs passed Thread #1: 1650000 read pairs finished. 301 secs passed Thread #2: 1700000 read pairs finished. 302 secs passed Thread #0: 1750000 read pairs finished. 302 secs passed Thread #3: 1800000 read pairs finished. 303 secs passed Thread #1: 1850000 read pairs finished. 329 secs passed Thread #2: 1900000 read pairs finished. 330 secs passed Thread #0: 1950000 read pairs finished. 331 secs passed Thread #3: 2000000 read pairs finished. 332 secs passed Thread #1: 2050000 read pairs finished. 357 secs passed Thread #2: 2100000 read pairs finished. 358 secs passed Thread #0: 2150000 read pairs finished. 359 secs passed Thread #3: 2200000 read pairs finished. 360 secs passed Thread #1: 2250000 read pairs finished. 384 secs passed Thread #2: 2300000 read pairs finished. 385 secs passed Thread #0: 2350000 read pairs finished. 386 secs passed Thread #3: 2400000 read pairs finished. 387 secs passed Thread #1: 2450000 read pairs finished. 412 secs passed Thread #2: 2500000 read pairs finished. 413 secs passed Thread #0: 2550000 read pairs finished. 414 secs passed Thread #3: 2600000 read pairs finished. 415 secs passed Thread #1: 2650000 read pairs finished. 439 secs passed Thread #2: 2700000 read pairs finished. 440 secs passed Thread #0: 2750000 read pairs finished. 441 secs passed Thread #3: 2800000 read pairs finished. 442 secs passed Thread #1: 2850000 read pairs finished. 466 secs passed Thread #2: 2900000 read pairs finished. 467 secs passed Thread #0: 2950000 read pairs finished. 469 secs passed Thread #3: 3000000 read pairs finished. 470 secs passed Thread #1: 3050000 read pairs finished. 494 secs passed Thread #2: 3100000 read pairs finished. 494 secs passed Thread #0: 3150000 read pairs finished. 496 secs passed Thread #3: 3200000 read pairs finished. 497 secs passed Thread #1: 3250000 read pairs finished. 522 secs passed Thread #2: 3300000 read pairs finished. 522 secs passed Thread #0: 3350000 read pairs finished. 523 secs passed Thread #3: 3400000 read pairs finished. 524 secs passed Thread #1: 3450000 read pairs finished. 550 secs passed Thread #2: 3500000 read pairs finished. 550 secs passed Thread #0: 3550000 read pairs finished. 552 secs passed Thread #3: 3600000 read pairs finished. 552 secs passed Thread #1: 3650000 read pairs finished. 577 secs passed Thread #2: 3700000 read pairs finished. 578 secs passed Thread #0: 3750000 read pairs finished. 579 secs passed Thread #3: 3800000 read pairs finished. 580 secs passed Thread #1: 3850000 read pairs finished. 603 secs passed Thread #2: 3900000 read pairs finished. 605 secs passed Thread #0: 3950000 read pairs finished. 606 secs passed Thread #3: 4000000 read pairs finished. 609 secs passed Thread #1: 4050000 read pairs finished. 632 secs passed Thread #2: 4100000 read pairs finished. 633 secs passed Thread #0: 4150000 read pairs finished. 634 secs passed Thread #3: 4200000 read pairs finished. 637 secs passed Thread #1: 4250000 read pairs finished. 659 secs passed Thread #2: 4300000 read pairs finished. 660 secs passed Thread #0: 4350000 read pairs finished. 661 secs passed Thread #3: 4400000 read pairs finished. 665 secs passed Thread #1: 4450000 read pairs finished. 687 secs passed Thread #2: 4500000 read pairs finished. 688 secs passed Thread #0: 4550000 read pairs finished. 688 secs passed Thread #3: 4600000 read pairs finished. 691 secs passed Thread #1: 4650000 read pairs finished. 714 secs passed Thread #2: 4700000 read pairs finished. 715 secs passed Thread #0: 4750000 read pairs finished. 715 secs passed Thread #3: 4800000 read pairs finished. 717 secs passed Thread #1: 4850000 read pairs finished. 742 secs passed Thread #2: 4900000 read pairs finished. 743 secs passed Thread #0: 4950000 read pairs finished. 743 secs passed Thread #3: 5000000 read pairs finished. 744 secs passed Thread #1: 5050000 read pairs finished. 769 secs passed Thread #2: 5100000 read pairs finished. 770 secs passed Thread #0: 5150000 read pairs finished. 770 secs passed Thread #3: 5200000 read pairs finished. 771 secs passed Thread #1: 5250000 read pairs finished. 796 secs passed Thread #2: 5300000 read pairs finished. 798 secs passed Thread #3: 5400000 read pairs finished. 799 secs passed Thread #0: 5350000 read pairs finished. 800 secs passed Thread #1: 5450000 read pairs finished. 823 secs passed Thread #2: 5500000 read pairs finished. 824 secs passed Thread #3: 5550000 read pairs finished. 826 secs passed Thread #0: 5600000 read pairs finished. 827 secs passed Thread #1: 5650000 read pairs finished. 849 secs passed Thread #2: 5700000 read pairs finished. 851 secs passed Thread #3: 5750000 read pairs finished. 854 secs passed Thread #0: 5800000 read pairs finished. 855 secs passed Thread #1: 5850000 read pairs finished. 877 secs passed Thread #2: 5900000 read pairs finished. 878 secs passed Thread #3: 5950000 read pairs finished. 881 secs passed Thread #0: 6000000 read pairs finished. 882 secs passed Thread #0: 6179274 read pairs finished. 898 secs passed Thread #1: 6050000 read pairs finished. 903 secs passed Thread #2: 6100000 read pairs finished. 903 secs passed Thread #3: 6150000 read pairs finished. 905 secs passed Total number of aligned reads: pairs: 2725017 (44%) single a: 1704213 (28%) single b: 1415455 (23%) Done. Finished at Wed Jul 9 16:11:28 2014 Total time consumed: 906 secs
In [12]:
lib="M3"
In [13]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 4
BSMAP v2.74 Start at: Wed Jul 9 16:11:29 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 12 secs passed total_kmers: 43046721 Create seed table. 52 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: mcf_M3_R1.fastq (format: FASTQ) Input read file #2: mcf_M3_R2.fastq (format: FASTQ) Output file: bsmap_out_M3.sam (format: SAM) Thread #0: 50000 read pairs finished. 75 secs passed Thread #2: 100000 read pairs finished. 76 secs passed Thread #1: 150000 read pairs finished. 77 secs passed Thread #3: 200000 read pairs finished. 77 secs passed Thread #0: 250000 read pairs finished. 100 secs passed Thread #2: 300000 read pairs finished. 101 secs passed Thread #1: 350000 read pairs finished. 102 secs passed Thread #3: 400000 read pairs finished. 102 secs passed Thread #0: 450000 read pairs finished. 125 secs passed Thread #2: 500000 read pairs finished. 126 secs passed Thread #1: 550000 read pairs finished. 127 secs passed Thread #3: 600000 read pairs finished. 127 secs passed Thread #0: 650000 read pairs finished. 149 secs passed Thread #2: 700000 read pairs finished. 151 secs passed Thread #1: 750000 read pairs finished. 152 secs passed Thread #3: 800000 read pairs finished. 152 secs passed Thread #0: 850000 read pairs finished. 174 secs passed Thread #2: 900000 read pairs finished. 175 secs passed Thread #1: 950000 read pairs finished. 177 secs passed Thread #3: 1000000 read pairs finished. 178 secs passed Thread #0: 1050000 read pairs finished. 199 secs passed Thread #2: 1100000 read pairs finished. 200 secs passed Thread #1: 1150000 read pairs finished. 202 secs passed Thread #3: 1200000 read pairs finished. 203 secs passed Thread #0: 1250000 read pairs finished. 225 secs passed Thread #2: 1300000 read pairs finished. 226 secs passed Thread #1: 1350000 read pairs finished. 229 secs passed Thread #3: 1400000 read pairs finished. 229 secs passed Thread #0: 1450000 read pairs finished. 251 secs passed Thread #2: 1500000 read pairs finished. 252 secs passed Thread #1: 1550000 read pairs finished. 254 secs passed Thread #3: 1600000 read pairs finished. 255 secs passed Thread #0: 1650000 read pairs finished. 276 secs passed Thread #2: 1700000 read pairs finished. 277 secs passed Thread #1: 1750000 read pairs finished. 279 secs passed Thread #3: 1800000 read pairs finished. 280 secs passed Thread #0: 1850000 read pairs finished. 301 secs passed Thread #2: 1900000 read pairs finished. 301 secs passed Thread #1: 1950000 read pairs finished. 303 secs passed Thread #3: 2000000 read pairs finished. 304 secs passed Thread #0: 2050000 read pairs finished. 325 secs passed Thread #2: 2100000 read pairs finished. 326 secs passed Thread #1: 2150000 read pairs finished. 328 secs passed Thread #3: 2200000 read pairs finished. 329 secs passed Thread #0: 2250000 read pairs finished. 350 secs passed Thread #2: 2300000 read pairs finished. 351 secs passed Thread #1: 2350000 read pairs finished. 353 secs passed Thread #3: 2400000 read pairs finished. 354 secs passed Thread #0: 2450000 read pairs finished. 374 secs passed Thread #2: 2500000 read pairs finished. 377 secs passed Thread #1: 2550000 read pairs finished. 380 secs passed Thread #3: 2600000 read pairs finished. 380 secs passed Thread #0: 2650000 read pairs finished. 400 secs passed Thread #2: 2700000 read pairs finished. 403 secs passed Thread #1: 2750000 read pairs finished. 405 secs passed Thread #3: 2800000 read pairs finished. 406 secs passed Thread #0: 2850000 read pairs finished. 426 secs passed Thread #2: 2900000 read pairs finished. 428 secs passed Thread #1: 2950000 read pairs finished. 430 secs passed Thread #3: 3000000 read pairs finished. 431 secs passed Thread #0: 3050000 read pairs finished. 450 secs passed Thread #2: 3100000 read pairs finished. 451 secs passed Thread #1: 3150000 read pairs finished. 454 secs passed Thread #3: 3200000 read pairs finished. 455 secs passed Thread #0: 3250000 read pairs finished. 474 secs passed Thread #2: 3300000 read pairs finished. 475 secs passed Thread #1: 3350000 read pairs finished. 478 secs passed Thread #3: 3400000 read pairs finished. 481 secs passed Thread #0: 3450000 read pairs finished. 501 secs passed Thread #2: 3500000 read pairs finished. 502 secs passed Thread #1: 3550000 read pairs finished. 503 secs passed Thread #3: 3600000 read pairs finished. 505 secs passed Thread #0: 3650000 read pairs finished. 527 secs passed Thread #2: 3700000 read pairs finished. 527 secs passed Thread #1: 3750000 read pairs finished. 527 secs passed Thread #3: 3800000 read pairs finished. 530 secs passed Thread #0: 3850000 read pairs finished. 551 secs passed Thread #2: 3900000 read pairs finished. 552 secs passed Thread #1: 3950000 read pairs finished. 552 secs passed Thread #3: 4000000 read pairs finished. 554 secs passed Thread #0: 4050000 read pairs finished. 576 secs passed Thread #2: 4100000 read pairs finished. 577 secs passed Thread #1: 4150000 read pairs finished. 578 secs passed Thread #3: 4200000 read pairs finished. 578 secs passed Thread #0: 4250000 read pairs finished. 601 secs passed Thread #2: 4300000 read pairs finished. 601 secs passed Thread #1: 4350000 read pairs finished. 603 secs passed Thread #3: 4400000 read pairs finished. 604 secs passed Thread #0: 4450000 read pairs finished. 625 secs passed Thread #2: 4500000 read pairs finished. 626 secs passed Thread #1: 4550000 read pairs finished. 627 secs passed Thread #3: 4600000 read pairs finished. 628 secs passed Thread #0: 4650000 read pairs finished. 650 secs passed Thread #2: 4700000 read pairs finished. 650 secs passed Thread #1: 4750000 read pairs finished. 651 secs passed Thread #3: 4800000 read pairs finished. 652 secs passed Thread #0: 4850000 read pairs finished. 674 secs passed Thread #2: 4900000 read pairs finished. 675 secs passed Thread #1: 4950000 read pairs finished. 675 secs passed Thread #3: 5000000 read pairs finished. 676 secs passed Thread #0: 5050000 read pairs finished. 697 secs passed Thread #2: 5100000 read pairs finished. 699 secs passed Thread #1: 5150000 read pairs finished. 700 secs passed Thread #3: 5200000 read pairs finished. 702 secs passed Thread #0: 5250000 read pairs finished. 722 secs passed Thread #2: 5300000 read pairs finished. 723 secs passed Thread #1: 5350000 read pairs finished. 725 secs passed Thread #3: 5400000 read pairs finished. 726 secs passed Thread #0: 5450000 read pairs finished. 747 secs passed Thread #2: 5500000 read pairs finished. 748 secs passed Thread #1: 5550000 read pairs finished. 749 secs passed Thread #3: 5600000 read pairs finished. 750 secs passed Thread #0: 5650000 read pairs finished. 771 secs passed Thread #2: 5700000 read pairs finished. 772 secs passed Thread #1: 5750000 read pairs finished. 773 secs passed Thread #3: 5800000 read pairs finished. 774 secs passed Thread #0: 5850000 read pairs finished. 795 secs passed Thread #2: 5900000 read pairs finished. 797 secs passed Thread #1: 5950000 read pairs finished. 797 secs passed Thread #3: 6000000 read pairs finished. 798 secs passed Thread #0: 6050000 read pairs finished. 820 secs passed Thread #2: 6100000 read pairs finished. 823 secs passed Thread #1: 6150000 read pairs finished. 824 secs passed Thread #3: 6200000 read pairs finished. 824 secs passed Thread #0: 6250000 read pairs finished. 845 secs passed Thread #2: 6300000 read pairs finished. 848 secs passed Thread #1: 6350000 read pairs finished. 849 secs passed Thread #3: 6400000 read pairs finished. 850 secs passed Thread #0: 6450000 read pairs finished. 870 secs passed Thread #2: 6500000 read pairs finished. 872 secs passed Thread #1: 6550000 read pairs finished. 874 secs passed Thread #3: 6600000 read pairs finished. 875 secs passed Thread #0: 6650000 read pairs finished. 894 secs passed Thread #2: 6700000 read pairs finished. 896 secs passed Thread #1: 6750000 read pairs finished. 899 secs passed Thread #3: 6800000 read pairs finished. 900 secs passed Thread #0: 6850000 read pairs finished. 918 secs passed Thread #2: 6900000 read pairs finished. 920 secs passed Thread #1: 6950000 read pairs finished. 924 secs passed Thread #3: 7000000 read pairs finished. 925 secs passed Thread #0: 7050000 read pairs finished. 944 secs passed Thread #2: 7100000 read pairs finished. 944 secs passed Thread #1: 7150000 read pairs finished. 948 secs passed Thread #3: 7200000 read pairs finished. 949 secs passed Thread #0: 7250000 read pairs finished. 968 secs passed Thread #2: 7300000 read pairs finished. 969 secs passed Thread #1: 7350000 read pairs finished. 973 secs passed Thread #3: 7400000 read pairs finished. 974 secs passed Thread #0: 7450000 read pairs finished. 992 secs passed Thread #2: 7500000 read pairs finished. 993 secs passed Thread #1: 7550000 read pairs finished. 997 secs passed Thread #3: 7600000 read pairs finished. 998 secs passed Thread #0: 7650000 read pairs finished. 1016 secs passed Thread #2: 7700000 read pairs finished. 1017 secs passed Thread #1: 7750000 read pairs finished. 1021 secs passed Thread #3: 7800000 read pairs finished. 1022 secs passed Thread #2: 7867124 read pairs finished. 1025 secs passed Thread #0: 7850000 read pairs finished. 1035 secs passed Total number of aligned reads: pairs: 4545285 (58%) single a: 1331429 (17%) single b: 1233410 (16%) Done. Finished at Wed Jul 9 16:28:44 2014 Total time consumed: 1035 secs
In [14]:
lib="T3D3"
In [15]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 4
BSMAP v2.74 Start at: Wed Jul 9 16:28:45 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 13 secs passed total_kmers: 43046721 Create seed table. 51 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: mcf_T3D3_R1.fastq (format: FASTQ) Input read file #2: mcf_T3D3_R2.fastq (format: FASTQ) Output file: bsmap_out_T3D3.sam (format: SAM) Thread #2: 50000 read pairs finished. 75 secs passed Thread #1: 100000 read pairs finished. 76 secs passed Thread #0: 150000 read pairs finished. 77 secs passed Thread #3: 200000 read pairs finished. 77 secs passed Thread #2: 250000 read pairs finished. 100 secs passed Thread #1: 300000 read pairs finished. 102 secs passed Thread #0: 350000 read pairs finished. 102 secs passed Thread #3: 400000 read pairs finished. 103 secs passed Thread #2: 450000 read pairs finished. 125 secs passed Thread #1: 500000 read pairs finished. 126 secs passed Thread #0: 550000 read pairs finished. 127 secs passed Thread #3: 600000 read pairs finished. 128 secs passed Thread #2: 650000 read pairs finished. 150 secs passed Thread #1: 700000 read pairs finished. 151 secs passed Thread #0: 750000 read pairs finished. 153 secs passed Thread #3: 800000 read pairs finished. 154 secs passed Thread #2: 850000 read pairs finished. 175 secs passed Thread #1: 900000 read pairs finished. 176 secs passed Thread #0: 950000 read pairs finished. 178 secs passed Thread #3: 1000000 read pairs finished. 179 secs passed Thread #2: 1050000 read pairs finished. 200 secs passed Thread #1: 1100000 read pairs finished. 201 secs passed Thread #0: 1150000 read pairs finished. 203 secs passed Thread #3: 1200000 read pairs finished. 204 secs passed Thread #2: 1250000 read pairs finished. 226 secs passed Thread #1: 1300000 read pairs finished. 227 secs passed Thread #0: 1350000 read pairs finished. 228 secs passed Thread #3: 1400000 read pairs finished. 229 secs passed Thread #2: 1450000 read pairs finished. 252 secs passed Thread #1: 1500000 read pairs finished. 254 secs passed Thread #0: 1550000 read pairs finished. 255 secs passed Thread #3: 1600000 read pairs finished. 256 secs passed Thread #2: 1650000 read pairs finished. 278 secs passed Thread #1: 1700000 read pairs finished. 279 secs passed Thread #0: 1750000 read pairs finished. 280 secs passed Thread #3: 1800000 read pairs finished. 281 secs passed Thread #2: 1850000 read pairs finished. 304 secs passed Thread #1: 1900000 read pairs finished. 305 secs passed Thread #0: 1950000 read pairs finished. 305 secs passed Thread #3: 2000000 read pairs finished. 306 secs passed Thread #2: 2050000 read pairs finished. 330 secs passed Thread #1: 2100000 read pairs finished. 331 secs passed Thread #0: 2150000 read pairs finished. 331 secs passed Thread #3: 2200000 read pairs finished. 332 secs passed Thread #2: 2250000 read pairs finished. 354 secs passed Thread #1: 2300000 read pairs finished. 356 secs passed Thread #0: 2350000 read pairs finished. 357 secs passed Thread #3: 2400000 read pairs finished. 358 secs passed Thread #2: 2450000 read pairs finished. 383 secs passed Thread #0: 2550000 read pairs finished. 384 secs passed Thread #1: 2500000 read pairs finished. 384 secs passed Thread #3: 2600000 read pairs finished. 385 secs passed Thread #2: 2650000 read pairs finished. 408 secs passed Thread #0: 2700000 read pairs finished. 410 secs passed Thread #1: 2750000 read pairs finished. 411 secs passed Thread #3: 2800000 read pairs finished. 412 secs passed Thread #2: 2850000 read pairs finished. 434 secs passed Thread #0: 2900000 read pairs finished. 436 secs passed Thread #1: 2950000 read pairs finished. 437 secs passed Thread #3: 3000000 read pairs finished. 438 secs passed Thread #2: 3050000 read pairs finished. 460 secs passed Thread #0: 3100000 read pairs finished. 461 secs passed Thread #1: 3150000 read pairs finished. 463 secs passed Thread #3: 3200000 read pairs finished. 464 secs passed Thread #2: 3250000 read pairs finished. 485 secs passed Thread #0: 3300000 read pairs finished. 488 secs passed Thread #1: 3350000 read pairs finished. 489 secs passed Thread #3: 3400000 read pairs finished. 490 secs passed Thread #2: 3450000 read pairs finished. 511 secs passed Thread #0: 3500000 read pairs finished. 513 secs passed Thread #1: 3550000 read pairs finished. 515 secs passed Thread #3: 3600000 read pairs finished. 515 secs passed Thread #2: 3650000 read pairs finished. 536 secs passed Thread #0: 3700000 read pairs finished. 539 secs passed Thread #1: 3750000 read pairs finished. 540 secs passed Thread #3: 3800000 read pairs finished. 541 secs passed Thread #2: 3850000 read pairs finished. 562 secs passed Thread #0: 3900000 read pairs finished. 563 secs passed Thread #1: 3950000 read pairs finished. 565 secs passed Thread #3: 4000000 read pairs finished. 566 secs passed Thread #2: 4050000 read pairs finished. 587 secs passed Thread #0: 4100000 read pairs finished. 588 secs passed Thread #1: 4150000 read pairs finished. 590 secs passed Thread #3: 4200000 read pairs finished. 591 secs passed Thread #2: 4250000 read pairs finished. 613 secs passed Thread #0: 4300000 read pairs finished. 614 secs passed Thread #1: 4350000 read pairs finished. 616 secs passed Thread #3: 4400000 read pairs finished. 617 secs passed Thread #2: 4450000 read pairs finished. 638 secs passed Thread #0: 4500000 read pairs finished. 640 secs passed Thread #1: 4550000 read pairs finished. 642 secs passed Thread #3: 4600000 read pairs finished. 643 secs passed Thread #2: 4650000 read pairs finished. 663 secs passed Thread #0: 4700000 read pairs finished. 665 secs passed Thread #1: 4750000 read pairs finished. 667 secs passed Thread #3: 4800000 read pairs finished. 668 secs passed Thread #2: 4850000 read pairs finished. 689 secs passed Thread #0: 4900000 read pairs finished. 690 secs passed Thread #1: 4950000 read pairs finished. 692 secs passed Thread #3: 5000000 read pairs finished. 693 secs passed Thread #2: 5050000 read pairs finished. 713 secs passed Thread #0: 5100000 read pairs finished. 715 secs passed Thread #1: 5150000 read pairs finished. 717 secs passed Thread #3: 5200000 read pairs finished. 718 secs passed Thread #2: 5250000 read pairs finished. 742 secs passed Thread #0: 5300000 read pairs finished. 743 secs passed Thread #1: 5350000 read pairs finished. 744 secs passed Thread #3: 5400000 read pairs finished. 745 secs passed Thread #2: 5450000 read pairs finished. 768 secs passed Thread #0: 5500000 read pairs finished. 769 secs passed Thread #1: 5550000 read pairs finished. 770 secs passed Thread #3: 5600000 read pairs finished. 771 secs passed Thread #2: 5650000 read pairs finished. 794 secs passed Thread #0: 5700000 read pairs finished. 795 secs passed Thread #1: 5750000 read pairs finished. 796 secs passed Thread #3: 5800000 read pairs finished. 796 secs passed Thread #2: 5850000 read pairs finished. 819 secs passed Thread #0: 5900000 read pairs finished. 820 secs passed Thread #1: 5950000 read pairs finished. 821 secs passed Thread #3: 6000000 read pairs finished. 822 secs passed Thread #2: 6050000 read pairs finished. 844 secs passed Thread #0: 6100000 read pairs finished. 845 secs passed Thread #1: 6150000 read pairs finished. 848 secs passed Thread #3: 6200000 read pairs finished. 848 secs passed Thread #2: 6250000 read pairs finished. 871 secs passed Thread #0: 6300000 read pairs finished. 871 secs passed Thread #1: 6350000 read pairs finished. 873 secs passed Thread #3: 6400000 read pairs finished. 874 secs passed Thread #2: 6450000 read pairs finished. 896 secs passed Thread #0: 6500000 read pairs finished. 897 secs passed Thread #1: 6550000 read pairs finished. 899 secs passed Thread #3: 6600000 read pairs finished. 899 secs passed Thread #2: 6650000 read pairs finished. 922 secs passed Thread #0: 6700000 read pairs finished. 922 secs passed Thread #1: 6750000 read pairs finished. 923 secs passed Thread #3: 6800000 read pairs finished. 924 secs passed Thread #2: 6850000 read pairs finished. 948 secs passed Thread #0: 6900000 read pairs finished. 949 secs passed Thread #1: 6950000 read pairs finished. 949 secs passed Thread #3: 7000000 read pairs finished. 951 secs passed Thread #1: 7109789 read pairs finished. 954 secs passed Thread #2: 7050000 read pairs finished. 968 secs passed Thread #0: 7100000 read pairs finished. 969 secs passed Total number of aligned reads: pairs: 4047152 (57%) single a: 1332491 (19%) single b: 1205102 (17%) Done. Finished at Wed Jul 9 16:44:54 2014 Total time consumed: 969 secs
In [16]:
lib="T3D5"
In [17]:
! {bsmaploc}bsmap \
-a mcf_{lib}_R1.fastq \
-b mcf_{lib}_R2.fastq \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-o bsmap_out_{lib}.sam \
-p 4
BSMAP v2.74 Start at: Wed Jul 9 16:44:56 2014 Input reference file: Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 12 secs passed total_kmers: 43046721 Create seed table. 50 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: mcf_T3D5_R1.fastq (format: FASTQ) Input read file #2: mcf_T3D5_R2.fastq (format: FASTQ) Output file: bsmap_out_T3D5.sam (format: SAM) Thread #3: 50000 read pairs finished. 74 secs passed Thread #2: 100000 read pairs finished. 75 secs passed Thread #0: 150000 read pairs finished. 76 secs passed Thread #1: 200000 read pairs finished. 76 secs passed Thread #3: 250000 read pairs finished. 98 secs passed Thread #2: 300000 read pairs finished. 100 secs passed Thread #0: 350000 read pairs finished. 101 secs passed Thread #1: 400000 read pairs finished. 102 secs passed Thread #3: 450000 read pairs finished. 123 secs passed Thread #2: 500000 read pairs finished. 124 secs passed Thread #0: 550000 read pairs finished. 126 secs passed Thread #1: 600000 read pairs finished. 127 secs passed Thread #3: 650000 read pairs finished. 148 secs passed Thread #2: 700000 read pairs finished. 149 secs passed Thread #0: 750000 read pairs finished. 151 secs passed Thread #1: 800000 read pairs finished. 152 secs passed Thread #3: 850000 read pairs finished. 173 secs passed Thread #2: 900000 read pairs finished. 174 secs passed Thread #0: 950000 read pairs finished. 177 secs passed Thread #1: 1000000 read pairs finished. 178 secs passed Thread #3: 1050000 read pairs finished. 198 secs passed Thread #2: 1100000 read pairs finished. 199 secs passed Thread #0: 1150000 read pairs finished. 201 secs passed Thread #1: 1200000 read pairs finished. 202 secs passed Thread #3: 1250000 read pairs finished. 223 secs passed Thread #2: 1300000 read pairs finished. 224 secs passed Thread #0: 1350000 read pairs finished. 226 secs passed Thread #1: 1400000 read pairs finished. 227 secs passed Thread #3: 1450000 read pairs finished. 248 secs passed Thread #2: 1500000 read pairs finished. 249 secs passed Thread #0: 1550000 read pairs finished. 251 secs passed Thread #1: 1600000 read pairs finished. 252 secs passed Thread #3: 1650000 read pairs finished. 274 secs passed Thread #2: 1700000 read pairs finished. 275 secs passed Thread #0: 1750000 read pairs finished. 276 secs passed Thread #1: 1800000 read pairs finished. 277 secs passed Thread #3: 1850000 read pairs finished. 299 secs passed Thread #2: 1900000 read pairs finished. 300 secs passed Thread #0: 1950000 read pairs finished. 301 secs passed Thread #1: 2000000 read pairs finished. 302 secs passed Thread #3: 2050000 read pairs finished. 324 secs passed Thread #2: 2100000 read pairs finished. 325 secs passed Thread #0: 2150000 read pairs finished. 326 secs passed Thread #1: 2200000 read pairs finished. 327 secs passed Thread #3: 2250000 read pairs finished. 350 secs passed Thread #2: 2300000 read pairs finished. 351 secs passed Thread #0: 2350000 read pairs finished. 351 secs passed Thread #1: 2400000 read pairs finished. 352 secs passed Thread #3: 2450000 read pairs finished. 375 secs passed Thread #2: 2500000 read pairs finished. 376 secs passed Thread #0: 2550000 read pairs finished. 377 secs passed Thread #1: 2600000 read pairs finished. 378 secs passed Thread #3: 2650000 read pairs finished. 400 secs passed Thread #2: 2700000 read pairs finished. 400 secs passed Thread #0: 2750000 read pairs finished. 402 secs passed Thread #1: 2800000 read pairs finished. 403 secs passed Thread #3: 2850000 read pairs finished. 426 secs passed Thread #2: 2900000 read pairs finished. 427 secs passed Thread #0: 2950000 read pairs finished. 428 secs passed Thread #1: 3000000 read pairs finished. 429 secs passed Thread #3: 3050000 read pairs finished. 451 secs passed Thread #2: 3100000 read pairs finished. 452 secs passed Thread #0: 3150000 read pairs finished. 454 secs passed Thread #1: 3200000 read pairs finished. 455 secs passed Thread #3: 3250000 read pairs finished. 476 secs passed Thread #2: 3300000 read pairs finished. 477 secs passed Thread #0: 3350000 read pairs finished. 479 secs passed Thread #1: 3400000 read pairs finished. 480 secs passed Thread #3: 3450000 read pairs finished. 507 secs passed Thread #2: 3500000 read pairs finished. 508 secs passed Thread #0: 3550000 read pairs finished. 512 secs passed Thread #1: 3600000 read pairs finished. 515 secs passed Thread #2: 3700000 read pairs finished. 534 secs passed Thread #3: 3650000 read pairs finished. 535 secs passed Thread #0: 3750000 read pairs finished. 538 secs passed Thread #1: 3800000 read pairs finished. 541 secs passed Thread #2: 3850000 read pairs finished. 560 secs passed Thread #3: 3900000 read pairs finished. 560 secs passed Thread #0: 3950000 read pairs finished. 563 secs passed Thread #1: 4000000 read pairs finished. 566 secs passed Thread #2: 4050000 read pairs finished. 585 secs passed Thread #3: 4100000 read pairs finished. 585 secs passed Thread #0: 4150000 read pairs finished. 587 secs passed Thread #1: 4200000 read pairs finished. 591 secs passed Thread #2: 4250000 read pairs finished. 611 secs passed Thread #3: 4300000 read pairs finished. 611 secs passed Thread #0: 4350000 read pairs finished. 613 secs passed Thread #1: 4400000 read pairs finished. 616 secs passed Thread #2: 4450000 read pairs finished. 636 secs passed Thread #3: 4500000 read pairs finished. 637 secs passed Thread #0: 4550000 read pairs finished. 638 secs passed Thread #1: 4600000 read pairs finished. 640 secs passed Thread #2: 4650000 read pairs finished. 662 secs passed Thread #3: 4700000 read pairs finished. 663 secs passed Thread #0: 4750000 read pairs finished. 663 secs passed Thread #1: 4800000 read pairs finished. 665 secs passed Thread #2: 4850000 read pairs finished. 686 secs passed Thread #3: 4900000 read pairs finished. 688 secs passed Thread #0: 4950000 read pairs finished. 690 secs passed Thread #1: 5000000 read pairs finished. 691 secs passed Thread #2: 5050000 read pairs finished. 712 secs passed Thread #3: 5100000 read pairs finished. 714 secs passed Thread #0: 5150000 read pairs finished. 715 secs passed Thread #1: 5200000 read pairs finished. 717 secs passed Thread #2: 5250000 read pairs finished. 739 secs passed Thread #3: 5300000 read pairs finished. 739 secs passed Thread #0: 5350000 read pairs finished. 741 secs passed Thread #1: 5400000 read pairs finished. 742 secs passed Thread #2: 5450000 read pairs finished. 764 secs passed Thread #3: 5500000 read pairs finished. 764 secs passed Thread #0: 5550000 read pairs finished. 765 secs passed Thread #1: 5600000 read pairs finished. 766 secs passed Thread #2: 5650000 read pairs finished. 788 secs passed Thread #3: 5700000 read pairs finished. 791 secs passed Thread #1: 5800000 read pairs finished. 792 secs passed Thread #0: 5750000 read pairs finished. 793 secs passed Thread #2: 5850000 read pairs finished. 813 secs passed Thread #3: 5900000 read pairs finished. 816 secs passed Thread #1: 5950000 read pairs finished. 818 secs passed Thread #0: 6000000 read pairs finished. 819 secs passed Thread #2: 6050000 read pairs finished. 838 secs passed Thread #3: 6100000 read pairs finished. 841 secs passed Thread #1: 6150000 read pairs finished. 843 secs passed Thread #0: 6200000 read pairs finished. 844 secs passed Thread #2: 6250000 read pairs finished. 863 secs passed Thread #3: 6300000 read pairs finished. 865 secs passed Thread #1: 6350000 read pairs finished. 867 secs passed Thread #0: 6400000 read pairs finished. 869 secs passed Thread #2: 6450000 read pairs finished. 889 secs passed Thread #3: 6500000 read pairs finished. 891 secs passed Thread #1: 6550000 read pairs finished. 893 secs passed Thread #0: 6600000 read pairs finished. 894 secs passed Thread #2: 6650000 read pairs finished. 914 secs passed Thread #3: 6700000 read pairs finished. 916 secs passed Thread #1: 6750000 read pairs finished. 918 secs passed Thread #0: 6800000 read pairs finished. 919 secs passed Thread #2: 6850000 read pairs finished. 939 secs passed Thread #3: 6900000 read pairs finished. 940 secs passed Thread #1: 6950000 read pairs finished. 942 secs passed Thread #0: 7000000 read pairs finished. 944 secs passed Thread #1: 7125800 read pairs finished. 953 secs passed Thread #2: 7050000 read pairs finished. 959 secs passed Thread #3: 7100000 read pairs finished. 960 secs passed Total number of aligned reads: pairs: 4092568 (57%) single a: 1250715 (18%) single b: 1133306 (16%) Done. Finished at Wed Jul 9 17:00:56 2014 Total time consumed: 960 secs
methratio¶
In [18]:
lib="M1"
In [19]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 17:00:58 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 17:01:50 2014: reading bsmap_out_M1.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 17:09:01 2014: read 10000000 lines @ Wed Jul 9 17:09:38 2014: combining CpG methylation from both strands ... @ Wed Jul 9 17:10:11 2014: writing methratio_out_M1.txt ... @ Wed Jul 9 17:22:53 2014: done. total 8716465 valid mappings, 48671759 covered cytosines, average coverage: 1.78 fold.
In [20]:
lib="T1D3"
In [21]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 17:22:55 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 17:23:46 2014: reading bsmap_out_T1D3.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 17:27:44 2014: combining CpG methylation from both strands ... @ Wed Jul 9 17:28:17 2014: writing methratio_out_T1D3.txt ... @ Wed Jul 9 17:35:59 2014: done. total 5759215 valid mappings, 26507310 covered cytosines, average coverage: 1.32 fold.
In [22]:
lib="T1D5"
In [23]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 17:36:01 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 17:36:51 2014: reading bsmap_out_T1D5.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 17:43:03 2014: combining CpG methylation from both strands ... @ Wed Jul 9 17:43:37 2014: writing methratio_out_T1D5.txt ... @ Wed Jul 9 17:56:54 2014: done. total 6974208 valid mappings, 45446465 covered cytosines, average coverage: 1.54 fold.
In [24]:
lib="M3"
In [25]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 17:56:56 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 17:57:47 2014: reading bsmap_out_M3.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 18:05:09 2014: read 10000000 lines @ Wed Jul 9 18:06:22 2014: combining CpG methylation from both strands ... @ Wed Jul 9 18:06:55 2014: writing methratio_out_M3.txt ... @ Wed Jul 9 18:20:40 2014: done. total 9773223 valid mappings, 53389886 covered cytosines, average coverage: 1.78 fold.
In [26]:
lib="T3D3"
In [27]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 18:20:42 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 18:21:33 2014: reading bsmap_out_T3D3.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 18:28:52 2014: read 10000000 lines @ Wed Jul 9 18:29:19 2014: combining CpG methylation from both strands ... @ Wed Jul 9 18:29:51 2014: writing methratio_out_T3D3.txt ... @ Wed Jul 9 18:43:22 2014: done. total 8847902 valid mappings, 52255860 covered cytosines, average coverage: 1.65 fold.
In [28]:
lib="T3D5"
In [29]:
!python {bsmaploc}methratio.py \
-d Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa \
-u -z -g \
-o methratio_out_{lib}.txt \
-s {bsmaploc}samtools \
bsmap_out_{lib}.sam \
@ Wed Jul 9 18:43:23 2014: reading reference Crassostrea_gigas.GCA_000297895.1.22.dna_sm.genome.fa ... @ Wed Jul 9 18:44:14 2014: reading bsmap_out_T3D5.sam ... [samopen] SAM header is present: 7658 sequences. @ Wed Jul 9 18:52:58 2014: read 10000000 lines @ Wed Jul 9 18:53:22 2014: combining CpG methylation from both strands ... @ Wed Jul 9 18:53:55 2014: writing methratio_out_T3D5.txt ... @ Wed Jul 9 19:07:18 2014: done. total 8808414 valid mappings, 51732152 covered cytosines, average coverage: 1.69 fold.
Converting methratio files for methylkit
In [39]:
!cat ./scripts/mr3x.awk
#!/usr/bin/awk -f
!awk {if ($8 >= 3) print $1,$2,$3,$4,$5,$6,$7,$8,$9,$10,$11,$12}
In [40]:
!cat ./scripts/mr_gg.awk.sh
#!/usr/bin/awk -f
BEGIN{ print "chr.Base\tchr\tbase\tstrand\tcoverage\tfreqC\tfreqT" }
{
if ($3 == "+") {
strand="F"
} else {
strand="R"
}
FC=($7/$8)*100
FT=(1-($7/$8))*100
chrbase=$1"."$2
printf "%s\t%s\t%s\t%s\t%d\t%.2f\t%.2f\n",
chrbase, $1, $2, strand, $8, FC, FT
}
In [30]:
for i in ("M1","T1D3","T1D5", "M3", "T3D3", "T3D5"):
!echo {i}
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out_{i}.txt> methratio_out_{i}CG.txt
!awk -f ./scripts/mr3x.awk methratio_out_{i}CG.txt > mr3x.{i}.txt
!awk -f ./scripts/mr_gg.awk.sh mr3x.{i}.txt > mkfmt_{i}.txt
M1 T1D3 T1D5 M3 T3D3 T3D5
Methylkit¶
In [31]:
%pylab inline
Populating the interactive namespace from numpy and matplotlib
In [32]:
%load_ext rpy2.ipython
In [33]:
%R library(methylKit)
Out[33]:
<StrVector - Python:0x1004440e0 / R:0x1119abd48> [str, str, str, ..., str, str, str]
In [34]:
%R library(data.table)
data.table 1.9.2 For help type: help("data.table")
In [35]:
%R library(GenomicRanges)
Loading required package: BiocGenerics
Loading required package: parallel
Attaching package: ‘BiocGenerics’
The following objects are masked from ‘package:parallel’:
clusterApply, clusterApplyLB, clusterCall, clusterEvalQ,
clusterExport, clusterMap, parApply, parCapply, parLapply,
parLapplyLB, parRapply, parSapply, parSapplyLB
The following object is masked from ‘package:stats’:
xtabs
The following objects are masked from ‘package:base’:
anyDuplicated, append, as.data.frame, as.vector, cbind, colnames,
duplicated, eval, evalq, Filter, Find, get, intersect, is.unsorted,
lapply, Map, mapply, match, mget, order, paste, pmax, pmax.int,
pmin, pmin.int, Position, rank, rbind, Reduce, rep.int, rownames,
sapply, setdiff, sort, table, tapply, union, unique, unlist
Loading required package: IRanges
Loading required package: XVector
Attaching package: ‘GenomicRanges’
The following object is masked from ‘package:data.table’:
last
In [29]:
ls mkfmt*
mkfmt_CgM1.txt mkfmt_CgM3.txt mkfmt_CgT1D3.txt mkfmt_CgT1D5.txt mkfmt_CgT3D3.txt mkfmt_CgT3D5.txt mkfmt_CgM1txt mkfmt_CgM3txt mkfmt_CgT1D3txt mkfmt_CgT1D5txt mkfmt_CgT3D3txt mkfmt_CgT3D5txt
In [36]:
%%R file.list <- list
('mkfmt_M1.txt',
'mkfmt_T1D3.txt',
'mkfmt_T1D5.txt',
'mkfmt_M3.txt',
'mkfmt_T3D3.txt',
'mkfmt_T3D5.txt'
)
In [37]:
%R myobj=read(file.list,sample.id=list("1_sperm","1_72hpf","1_120hpf","2_sperm","2_72hpf","2_120hpf"),assembly="v9",treatment=c(0,0,0,1,1,1))
Out[37]:
<ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...] <ListVector - Python:0x10ae60680 / R:0x10f3bc5c0> [ListV..., ListV..., ListV..., ListV..., ListV..., ListV...]
In [38]:
%%R
meth<-unite(myobj)
head(meth)
nrow(meth)
getCorrelation(meth,plot=T)
hc<- clusterSamples(meth, dist="correlation", method="ward", plot=T)
PCA<-PCASamples(meth)
1_sperm 1_72hpf 1_120hpf 2_sperm 2_72hpf 2_120hpf 1_sperm 1.0000000 0.8185855 0.8381244 0.8028188 0.8048533 0.8064576 1_72hpf 0.8185855 1.0000000 0.8162468 0.8077710 0.8160557 0.8195972 1_120hpf 0.8381244 0.8162468 1.0000000 0.8260072 0.8346218 0.8373328 2_sperm 0.8028188 0.8077710 0.8260072 1.0000000 0.8813808 0.8815682 2_72hpf 0.8048533 0.8160557 0.8346218 0.8813808 1.0000000 0.8799352 2_120hpf 0.8064576 0.8195972 0.8373328 0.8815682 0.8799352 1.0000000 KernSmooth 2.23 loaded Copyright M. P. Wand 1997-2009
ID target loci¶
Summary: methration files filtered for CG and 3x coverage will be sligthly modified and uploaded to SQLshare for querying.
In [40]:
#example file generated above that will be used
!head mr3x.M1.txt
C12768 103 + TTCGT 0.167 6.00 1 6 6 6 0.030 0.564 C12768 119 + TTCGT 0.250 4.00 1 4 4 4 0.046 0.699 C12768 145 + TTCGT 0.000 3.00 0 3 3 3 0.000 0.562 C12806 56 + AACGA 0.000 4.00 0 4 4 4 0.000 0.490 C12806 76 + ATCGC 0.200 5.00 1 5 5 5 0.036 0.624 C12806 78 + CGCGT 0.200 5.00 1 5 5 5 0.036 0.624 C12806 105 + ATCGG 0.250 4.00 1 4 5 5 0.046 0.699 C12806 142 + ATCGG 0.375 8.00 3 8 8 8 0.137 0.694 C12924 19 + AACGA 0.000 4.00 0 4 4 4 0.000 0.490 C12924 30 + ATCGT 0.000 4.00 0 5 4 5 0.000 0.490
In [200]:
pwd
Out[200]:
u'/Volumes/Bay3 scratch/BiGo_manny'
In [232]:
for i in ("M1","T1D3","T1D5", "M3", "T3D3", "T3D5"):
!echo {i}
!awk -f ./scripts/psqls.awk mr3x.{i}.txt > short.{i}.txt
!echo "loci {i}ratio" >> {i}.head
!cat {i}.head short.{i}.txt > sql_{i}.txt
!sed '/NA/d' sql_{i}.txt > sqlr_{i}.txt
M1 T1D3 T1D5 M3 T3D3 T3D5
In [60]:
#fyi
!cat ./scripts/psqls.awk
#!/usr/bin/awk -f
!awk {print ($1"."$2),$5}
In [344]:
!head sqlr_M1.txt
loci M1ratio C12768.103 0.167 C12768.119 0.250 C12768.145 0.000 C12806.56 0.000 C12806.76 0.200 C12806.78 0.200 C12806.105 0.250 C12806.142 0.375 C12924.19 0.000
In [447]:
!wc -l sqlr_M1.txt
1212738 sqlr_M1.txt
SQLShare¶
In [43]:
#set locaiton of sqlshare python
pt="/Users/sr320/sqlshare-pythonclient/tools/"
In [258]:
!python {pt}multiupload.py sqlr_*
uploading sT3D3 uploading sT3D3 into [] uploading sqlr_T3D3.txt uploading sqlr_T3D3.txt into ['sqlr_T3D3.txt'] processing chunk line 0 to 1157463 (4.96696996689 s elapsed) pushing sqlr_T3D3.txt... parsing 567A0318... finished sqlr_T3D3.txt Successfully uploaded sqlr_T3D3.txt
In [426]:
!python {pt}fetchdata.py -s "SELECT *, ([M1ratio]+[T1D3ratio]+[T1D5ratio])/3 as mean1 FROM [sr320@washington.edu].[sqlr_M1.txt]m1 inner join [sr320@washington.edu].[sqlr_T1D5.txt]t1d5 on m1.loci=t1d5.loci inner join [sr320@washington.edu].[sqlr_T1D3.txt]t1d3 on m1.loci=t1d3.loci where [M1ratio]-(([M1ratio]+[T1D3ratio]+[T1D5ratio])/3) < abs(.2) and [T1D3ratio]-(([M1ratio]+[T1D3ratio]+[T1D5ratio])/3) < abs(.2) and [T1D5ratio]-(([M1ratio]+[T1D3ratio]+[T1D5ratio])/3) < abs(.2)" -f tsv -o cglarv_lineage_1.txt
In [427]:
!python {pt}fetchdata.py -s "SELECT *, ([M1ratio]+[T1D3ratio]+[T1D5ratio])/3 as mean1 FROM [sr320@washington.edu].[sqlr_M1.txt]m1 inner join [sr320@washington.edu].[sqlr_T1D5.txt]t1d5 on m1.loci=t1d5.loci inner join [sr320@washington.edu].[sqlr_T1D3.txt]t1d3 on m1.loci=t1d3.loci" -f tsv -o tocountlarv_lineage_1.txt
In [428]:
!head tocountlarv_lineage_1.txt
In [429]:
!wc -l tocountlarv_lineage_1.txt
69313 tocountlarv_lineage_1.txt
In [430]:
!head cglarv_lineage_1.txt
In [431]:
!python {pt}fetchdata.py -s "SELECT *, ([M3ratio]+[T3D3ratio]+[T3D5ratio])/3 as mean3 FROM [sr320@washington.edu].[sqlr_M3.txt]M3 inner join [sr320@washington.edu].[sqlr_T3D5.txt]T3D5 on M3.loci=T3D5.loci inner join [sr320@washington.edu].[sqlr_T3D3.txt]T3D3 on M3.loci=T3D3.loci where [M3ratio]-(([M3ratio]+[T3D3ratio]+[T3D5ratio])/3) < abs(.2) and [T3D3ratio]-(([M3ratio]+[T3D3ratio]+[T3D5ratio])/3) < abs(.2) and [T3D5ratio]-(([M3ratio]+[T3D3ratio]+[T3D5ratio])/3) < abs(.2)" -f tsv -o cglarv_lineage_3.txt
In [432]:
!head cglarv_lineage_3.txt
In [433]:
for i in ("1","3"):
!echo {i}
!awk -f ./scripts/joinr.awk cglarv_lineage_{i}.txt > cglarv_lineage_{i}trm.txt
!sed 's/loci/loci{i}/g' <cglarv_lineage_{i}trm.txt> sq_cglarv_lineage_{i}.txt
1 3
In [434]:
!head sq_cglarv_lineage_1.txt
In [435]:
!head sq_cglarv_lineage_3.txt
In [436]:
!python {pt}multiupload.py sq_cglarv_*
uploading sq_cglarv_lineage_1.txt uploading sq_cglarv_lineage_1.txt into ['sq_cglarv_lineage_1.txt'] processing chunk line 0 to 63043 (0.0331981182098 s elapsed) pushing sq_cglarv_lineage_1.txt... parsing 19EC8DBE... finished sq_cglarv_lineage_1.txt Successfully uploaded sq_cglarv_lineage_1.txt uploading sq_cglarv_lineage_3.txt uploading sq_cglarv_lineage_3.txt into ['sq_cglarv_lineage_3.txt'] processing chunk line 0 to 245244 (0.118955850601 s elapsed) pushing sq_cglarv_lineage_3.txt... parsing B1F7682A... finished sq_cglarv_lineage_3.txt Successfully uploaded sq_cglarv_lineage_3.txt
In [437]:
!python {pt}fetchdata.py -s "SELECT *, abs(mean1 - mean3) as meandiff FROM [sr320@washington.edu].[sq_cglarv_lineage_1.txt]c1 inner join [sr320@washington.edu].[sq_cglarv_lineage_3.txt]c3 on c1.loci1=c3.loci3 where abs(mean1 - mean3) >= 0.3" -f tsv -o cglarv_lineage_differ.txt
In [443]:
#QC check commmon loci
!python {pt}fetchdata.py -s "SELECT *, abs(mean1 - mean3) as meandiff FROM [sr320@washington.edu].[sq_cglarv_lineage_1.txt]c1 inner join [sr320@washington.edu].[sq_cglarv_lineage_3.txt]c3 on c1.loci1=c3.loci3" -f tsv -o commonloci.txt
In [445]:
!head commonloci.txt
In [446]:
!wc -l commonloci.txt
35475 commonloci.txt
In [438]:
!head cglarv_lineage_differ.txt
In [439]:
!awk '{print $1}' cglarv_lineage_differ.txt > cglarv_lin_a
!tail -n +2 cglarv_lin_a > cglarv_lin_b
!tr '.' "\t" <cglarv_lin_b> cglarv_lin_c
!awk '{print $1, $2, ($2+1), "DML_lin" }' cglarv_lin_c > cglarv_lin_d
!tr ' ' "\t" <cglarv_lin_d> cglarv_lineage_differ.bed
In [440]:
!head cglarv_lineage_differ.bed
C22094 1004 1005 DML_lin C25394 2981 2982 DML_lin C25670 2403 2404 DML_lin C25670 2410 2411 DML_lin C26040 898 899 DML_lin C26156 2111 2112 DML_lin C26256 553 554 DML_lin C26536 2616 2617 DML_lin C26646 2199 2200 DML_lin C26812 1897 1898 DML_lin
In [441]:
!wc -l cglarv_lineage_differ.bed
483 cglarv_lineage_differ.bed
Intersect bed¶
In [ ]:
Download oyster genome tracks
In [ ]:
mkdir genome_tracks
In [394]:
for i in ("exon","intron","TE","gene","1k5p_gene_promoter","CG"):
!wget -q -P genome_tracks http://eagle.fish.washington.edu/trilobite/Crassostrea_gigas_v9_tracks/Cgigas_v9_{i}.gff
In [442]:
for i in ("exon","intron","TE","gene","1k5p_gene_promoter"):
!intersectbed \
-wb \
-a cglarv_lineage_differ.bed \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_DML_lin_wb.txt
!intersectbed \
-u \
-a cglarv_lineage_differ.bed \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_DML_lin_u.txt
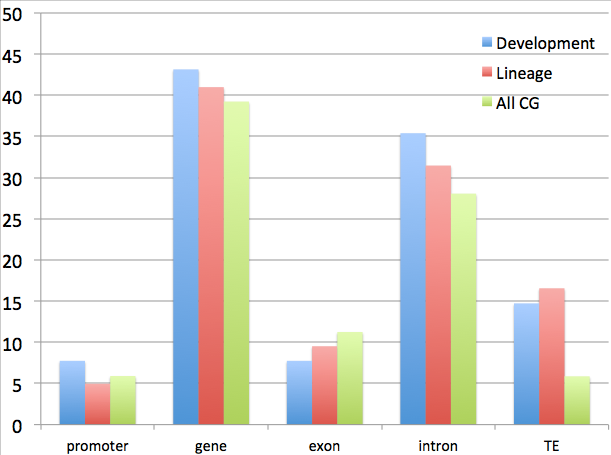
!wc -l {i}_intersect_DML_lin_u.txt
46 exon_intersect_DML_lin_u.txt
152 intron_intersect_DML_lin_u.txt
80 TE_intersect_DML_lin_u.txt
198 gene_intersect_DML_lin_u.txt
24 1k5p_gene_promoter_intersect_DML_lin_u.txt
In [20]:
!wc -l ./genome_tracks/Cgigas_v9_CG.gff
10035701 ./genome_tracks/Cgigas_v9_CG.gff
In [22]:
for i in ("exon","intron","TE","gene","1k5p_gene_promoter"):
!intersectbed \
-wb \
-a ./genome_tracks/Cgigas_v9_CG.gff \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_CG_wb.txt
!intersectbed \
-u \
-a ./genome_tracks/Cgigas_v9_CG.gff \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_CG_u.txt
!wc -l {i}_intersect_CG_u.txt
1129658 exon_intersect_CG_u.txt 2815997 intron_intersect_CG_u.txt 589509 TE_intersect_CG_u.txt 3938356 gene_intersect_CG_u.txt 593081 1k5p_gene_promoter_intersect_CG_u.txt
Developmental Differences¶
In [ ]:
ID similarities in Biological Reps
In [466]:
out="sperm"
r1="M1"
r2="M3"
In [468]:
!python {pt}fetchdata.py -s "SELECT *, ([{r2}ratio]+[{r1}ratio])/2 as mean_{out} FROM [sr320@washington.edu].[sqlr_{r1}.txt]{r1} inner join [sr320@washington.edu].[sqlr_{r2}.txt]{r2} on {r1}.loci={r2}.loci where [{r1}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2) and [{r2}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2)" -f tsv -o br_cglarv_{out}.txt
!wc -l br_cglarv_{out}.txt
!head br_cglarv_{out}.txt
!python {pt}singleupload.py br_cglarv_{out}.txt
421561 br_cglarv_sperm.txt processing chunk line 0 to 421561 (0.26978302002 s elapsed) pushing br_cglarv_sperm.txt... parsing 2396B0BE... finished br_cglarv_sperm.txt
In [469]:
out="day3"
r1="T1D3"
r2="T3D3"
In [470]:
!python {pt}fetchdata.py -s "SELECT *, ([{r2}ratio]+[{r1}ratio])/2 as mean_{out} FROM [sr320@washington.edu].[sqlr_{r1}.txt]{r1} inner join [sr320@washington.edu].[sqlr_{r2}.txt]{r2} on {r1}.loci={r2}.loci where [{r1}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2) and [{r2}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2)" -f tsv -o br_cglarv_{out}.txt
!wc -l br_cglarv_{out}.txt
!head br_cglarv_{out}.txt
!python {pt}singleupload.py br_cglarv_{out}.txt
108870 br_cglarv_day3.txt processing chunk line 0 to 108870 (0.104866027832 s elapsed) pushing br_cglarv_day3.txt... parsing 876A9D68... finished br_cglarv_day3.txt
In [471]:
out="day5"
r1="T1D5"
r2="T3D5"
In [477]:
!python {pt}fetchdata.py -s "SELECT *, ([{r2}ratio]+[{r1}ratio])/2 as mean_{out} FROM [sr320@washington.edu].[sqlr_{r1}.txt]{r1} inner join [sr320@washington.edu].[sqlr_{r2}.txt]{r2} on {r1}.loci={r2}.loci where [{r1}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2) and [{r2}ratio]-(([{r2}ratio]+[{r1}ratio])/2) < abs(.2)" -f tsv -o br_cglarv_{out}.txt
!wc -l br_cglarv_{out}.txt
!head br_cglarv_{out}.txt
!python {pt}singleupload.py br_cglarv_{out}.txt
302492 br_cglarv_day5.txt processing chunk line 0 to 302492 (0.181667089462 s elapsed) pushing br_cglarv_day5.txt... parsing A941BD6C... finished br_cglarv_day5.txt
In [473]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[br_cglarv_sperm.txt]sperm inner join [sr320@washington.edu].[br_cglarv_day3.txt]d3 on sperm.loci=d3.loci inner join [sr320@washington.edu].[br_cglarv_day5.txt]d5 on sperm.loci=d5.loci" -f tsv -o br_joinall.txt
In [476]:
!head -2 br_joinall.txt
In [475]:
!wc -l br_joinall.txt
36670 br_joinall.txt
In [478]:
!python {pt}fetchdata.py -s "SELECT *, sperm.mean_sperm-d3.mean_day3 as 's-3', sperm.mean_sperm-d5.mean_day5 as 's-5', d3.mean_day3-d5.mean_day5 as '3-5' FROM [sr320@washington.edu].[br_cglarv_sperm.txt]sperm inner join [sr320@washington.edu].[br_cglarv_day3.txt]d3 on sperm.loci=d3.loci inner join [sr320@washington.edu].[br_cglarv_day5.txt]d5 on sperm.loci=d5.loci where abs(sperm.mean_sperm-d3.mean_day3) > 0.4 or abs(sperm.mean_sperm-d5.mean_day5) > 0.4 or abs(d3.mean_day3-d5.mean_day5) > 0.4" -f tsv -o DML_development.txt
In [479]:
!head DML_development.txt
In [481]:
!wc -l DML_development.txt
388 DML_development.txt
In [482]:
!awk '{print $1}' DML_development.txt > cglarv_dd_a
!tail -n +2 cglarv_dd_a > cglarv_dd_b
!tr '.' "\t" <cglarv_dd_b> cglarv_dd_c
!awk '{print $1, $2, ($2+1), "DML_dev" }' cglarv_dd_c > cglarv_dd_d
!tr ' ' "\t" <cglarv_dd_d> cglarv_development_differ.bed
!head cglarv_development_differ.bed
!wc -l cglarv_development_differ.bed
C22666 609 610 DML_dev
C22666 624 625 DML_dev
C24478 2098 2099 DML_dev
C24478 2120 2121 DML_dev
C24628 2236 2237 DML_dev
C24628 473 474 DML_dev
C26614 3295 3296 DML_dev
C26938 1051 1052 DML_dev
C27500 1245 1246 DML_dev
C27928 1457 1458 DML_dev
387 cglarv_development_differ.bed
In [483]:
for i in ("exon","intron","TE","gene","1k5p_gene_promoter"):
!intersectbed \
-wb \
-a cglarv_development_differ.bed \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_DML_dev_wb.txt
!intersectbed \
-u \
-a cglarv_development_differ.bed \
-b ./genome_tracks/Cgigas_v9_{i}.gff \
> {i}_intersect_DML_dev_u.txt
!wc -l {i}_intersect_DML_dev_u.txt
30 exon_intersect_DML_dev_u.txt
137 intron_intersect_DML_dev_u.txt
57 TE_intersect_DML_dev_u.txt
167 gene_intersect_DML_dev_u.txt
18 1k5p_gene_promoter_intersect_DML_dev_u.txt
In [6]:
!head TE_intersect_DML_dev_wb.txt
C30526 2573 2574 DML_dev C30526 WUBlastX DNA_TcMar-Tc2 2452 2679 91 - . . C30526 2573 2574 DML_dev C30526 WUBlastX DNA_TcMar-Tc2 2452 2640 106 - . . C34310 1254 1255 DML_dev C34310 WUBlastX DNA_IS4EU 775 1458 20 - . . scaffold1171 48436 48437 DML_dev scaffold1171 WUBlastX DNA_hAT-hATw 47463 48491 36 + . . scaffold1171 48436 48437 DML_dev scaffold1171 WUBlastX DNA_hAT-hATw 47790 48476 102 + . . scaffold1171 48436 48437 DML_dev scaffold1171 WUBlastX DNA_hAT-hATw 48375 49889 213 + . . scaffold1188 80972 80973 DML_dev scaffold1188 WUBlastX LINE_R2-Hero 78711 81929 2619 - . . scaffold1188 81327 81328 DML_dev scaffold1188 WUBlastX LINE_R2-Hero 78711 81929 2619 - . . scaffold1227 137921 137922 DML_dev scaffold1227 TRF Tandem_Repeat 137882 138064 330 + . . scaffold1232 318833 318834 DML_dev scaffold1232 WUBlastX LTR_Pao 318601 319908 47 - . .
In [119]:
!wc -l TE_intersect_DML_lin_wb.txt
112 TE_intersect_DML_lin_wb.txt
In [18]:
!awk '{print $7}' TE_intersect_DML_dev_wb.txt > TE_intersect_DML_dev_wb_TEonly.txt
!<TE_intersect_DML_dev_wb_TEonly.txt cut -d' ' -f1 | sort -n | uniq -c
4 DNA_En-Spm 1 DNA_IS4EU 1 DNA_MuDR 1 DNA_Sola 2 DNA_TcMar-Tc2 1 DNA_hAT-Ac 3 DNA_hAT-hATw 1 LINE_CR1-L2 3 LINE_L1 1 LINE_L1-Tx1 9 LINE_L2 2 LINE_R2-Hero 1 LINE_RTE-X 3 LTR_Copia 5 LTR_DIRS 16 LTR_Gypsy 6 LTR_Gypsy-Cigr 4 LTR_Ngaro 10 LTR_Pao 1 RC_Helitron 13 Tandem_Repeat
In [19]:
!awk '{print $7}' TE_intersect_DML_lin_wb.txt > TE_intersect_DML_lin_wb_TEonly.txt
!<TE_intersect_DML_lin_wb_TEonly.txt cut -d' ' -f1 | sort -n | uniq -c
2 DNA_Helitron 1 DNA_IS 2 DNA_IS4EU 3 DNA_Kolobok 3 DNA_Kolobok-Hyd 1 DNA_Kolobok-IS4 3 DNA_MuDR 1 DNA_NOF 1 DNA_TcMar-Marin 1 DNA_TcMar-Tc2 2 DNA_hAT-Blackja 3 DNA_hAT-hATw 2 LINE_L1 5 LINE_L1-Tx1 1 LTR_Copia 4 LTR_DIRS 20 LTR_Gypsy 5 LTR_Gypsy-Cigr 16 LTR_Pao 14 RC_Helitron 22 Tandem_Repeat
In [24]:
!head TE_intersect_CG_wb.txt
scaffold350 fuzznuc nucleotide_motif 965 965 2 + . ID=scaffold350.20;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 965 1123 27 - . . scaffold350 fuzznuc nucleotide_motif 986 987 2 + . ID=scaffold350.21;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 965 1123 27 - . . scaffold350 fuzznuc nucleotide_motif 1015 1016 2 + . ID=scaffold350.22;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 965 1123 27 - . . scaffold350 fuzznuc nucleotide_motif 1042 1043 2 + . ID=scaffold350.23;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 965 1123 27 - . . scaffold350 fuzznuc nucleotide_motif 1042 1043 2 + . ID=scaffold350.23;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 1019 1123 33 - . . scaffold350 fuzznuc nucleotide_motif 1095 1096 2 + . ID=scaffold350.24;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 965 1123 27 - . . scaffold350 fuzznuc nucleotide_motif 1095 1096 2 + . ID=scaffold350.24;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 1019 1123 33 - . . scaffold350 fuzznuc nucleotide_motif 1161 1162 2 + . ID=scaffold350.25;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 1141 1863 356 - . . scaffold350 fuzznuc nucleotide_motif 1183 1184 2 + . ID=scaffold350.26;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 1141 1863 356 - . . scaffold350 fuzznuc nucleotide_motif 1200 1201 2 + . ID=scaffold350.27;note=*pat pattern:CG scaffold350 WUBlastX DNA_Academ 1141 1863 356 - . .
In [26]:
!awk '{print $13}' TE_intersect_CG_wb.txt > TE_intersect_CG_wb_TEonly.txt
!<TE_intersect_CG_wb_TEonly.txt cut -d' ' -f1 | sort -n | uniq -c
104 DNA 21 DNA?_Crypton? 2855 DNA_Academ 312 DNA_Chapaev 28 DNA_Chapaev-Cha 5217 DNA_Crypton 2834 DNA_En-Spm 14127 DNA_Harbinger 1006 DNA_Helitron 1579 DNA_IS 10257 DNA_IS4EU 2810 DNA_Kolobok 832 DNA_Kolobok-Hyd 2794 DNA_Kolobok-IS4 18792 DNA_Maverick 1177 DNA_Merlin 4480 DNA_MuDR 739 DNA_NOF 230 DNA_NOF? 10 DNA_Novosib 1422 DNA_P 4238 DNA_PiggyBac 6521 DNA_Sola 12 DNA_TcMar 137 DNA_TcMar-Ant1 3606 DNA_TcMar-Fot1 15 DNA_TcMar-IS630 9 DNA_TcMar-IS885 410 DNA_TcMar-ISRm1 7 DNA_TcMar-M44 6880 DNA_TcMar-Marin 1660 DNA_TcMar-Pogo 33 DNA_TcMar-Stowa 23935 DNA_TcMar-Tc1 3429 DNA_TcMar-Tc2 328 DNA_TcMar-Tigge 449 DNA_TcMar-m44 4 DNA_Transib 1173 DNA_Zator 3081 DNA_hAT 1930 DNA_hAT-Ac 286 DNA_hAT-Blackja 63 DNA_hAT-Charlie 23 DNA_hAT-Hobo 7 DNA_hAT-Restles 32 DNA_hAT-Tag1 5341 DNA_hAT-Tip100 35 DNA_hAT-Tip100? 10 DNA_hAT-Tol2 25 DNA_hAT-hAT5 1264 DNA_hAT-hATw 34 DNA_hAT-hATx 14609 LINE_CR1 9901 LINE_CR1-L2 113 LINE_DRE 36 LINE_Dong-R4 1667 LINE_I 344 LINE_I-Nimb 1668 LINE_Jockey 17973 LINE_L1 30803 LINE_L1-Tx1 10 LINE_L1? 32261 LINE_L2 614 LINE_L2-Hydra 94 LINE_LOA 8 LINE_Odin 17175 LINE_Penelope 2392 LINE_Proto2 156 LINE_R1 1103 LINE_R2 2508 LINE_R2-Hero 92 LINE_RTE 14138 LINE_RTE-BovB 338 LINE_RTE-RTE 56175 LINE_RTE-X 482 LINE_Rex-Babar 106 LINE_Tad1? 36 LINE_Telomeric 17 LINE_Zorro 89 LTR_Caulimoviru 3310 LTR_Copia 20656 LTR_DIRS 593 LTR_DIRS? 44 LTR_Delta 62 LTR_ERV-Foamy 3 LTR_ERV-Lenti 1640 LTR_ERV1 205 LTR_ERVK 48 LTR_ERVL 97 LTR_Foamy 132084 LTR_Gypsy 19891 LTR_Gypsy-Cigr 205 LTR_Gypsy-Gmr1 3352 LTR_Gypsy-Troyk 61 LTR_Gypsy? 27 LTR_Lenti 25073 LTR_Ngaro 38903 LTR_Pao 9607 RC_Helitron 235190 Tandem_Repeat 7 Unknown
Checking Gene Function¶
In [6]:
!head -3 gene_intersect_DML_dev_wb.txt
!wc -l gene_intersect_DML_dev_wb.txt
C30046 1305 1306 DML_dev C30046 GLEAN mRNA 202 6592 0.480335 + . ID=CGI_10000591;
C30046 522 523 DML_dev C30046 GLEAN mRNA 202 6592 0.480335 + . ID=CGI_10000591;
scaffold101 283044 283045 DML_dev scaffold101 GLEAN mRNA 269476 308543 0.957882 + . ID=CGI_10021644;
175 gene_intersect_DML_dev_wb.txt
In [7]:
!head -3 gene_intersect_DML_lin_wb.txt
!wc -l gene_intersect_DML_lin_wb.txt
C30046 1952 1953 DML_lin C30046 GLEAN mRNA 202 6592 0.480335 + . ID=CGI_10000591;
C33148 7907 7908 DML_lin C33148 GLEAN mRNA 5142 11739 0.999994 - . ID=CGI_10001003;
C33148 7925 7926 DML_lin C33148 GLEAN mRNA 5142 11739 0.999994 - . ID=CGI_10001003;
208 gene_intersect_DML_lin_wb.txt
In [8]:
!head -3 gene_intersect_CG_wb.txt
!wc -l gene_intersect_CG_wb.txt
scaffold350 fuzznuc nucleotide_motif 1161 1162 2 + . ID=scaffold350.25;note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . ID=CGI_10000780; scaffold350 fuzznuc nucleotide_motif 1183 1184 2 + . ID=scaffold350.26;note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . ID=CGI_10000780; scaffold350 fuzznuc nucleotide_motif 1200 1201 2 + . ID=scaffold350.27;note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . ID=CGI_10000780; 4046625 gene_intersect_CG_wb.txt
In [32]:
In [10]:
for i in ("DML_dev","DML_lin","CG"):
#!sed 's/ID=C/C/g' <gene_intersect_{i}_wb.txt> gene_intersect_{i}_wb_a.txt
#!sed 's/;//g' <gene_intersect_{i}_wb_a.txt> gene_intersect_{i}_wb_b.txt
!head -3 gene_intersect_{i}_wb_b.txt
!wc -l gene_intersect_{i}_wb_b.txt
#!python {pt}multiupload.py gene_intersect_{i}_wb_b.txt
C30046 1305 1306 DML_dev C30046 GLEAN mRNA 202 6592 0.480335 + . CGI_10000591
C30046 522 523 DML_dev C30046 GLEAN mRNA 202 6592 0.480335 + . CGI_10000591
scaffold101 283044 283045 DML_dev scaffold101 GLEAN mRNA 269476 308543 0.957882 + . CGI_10021644
175 gene_intersect_DML_dev_wb_b.txt
C30046 1952 1953 DML_lin C30046 GLEAN mRNA 202 6592 0.480335 + . CGI_10000591
C33148 7907 7908 DML_lin C33148 GLEAN mRNA 5142 11739 0.999994 - . CGI_10001003
C33148 7925 7926 DML_lin C33148 GLEAN mRNA 5142 11739 0.999994 - . CGI_10001003
208 gene_intersect_DML_lin_wb_b.txt
scaffold350 fuzznuc nucleotide_motif 1161 1162 2 + . ID=scaffold350.25note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . CGI_10000780
scaffold350 fuzznuc nucleotide_motif 1183 1184 2 + . ID=scaffold350.26note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . CGI_10000780
scaffold350 fuzznuc nucleotide_motif 1200 1201 2 + . ID=scaffold350.27note=*pat pattern:CG scaffold350 GLEAN mRNA 1105 3206 0.996578 - . CGI_10000780
4046625 gene_intersect_CG_wb_b.txt
In [30]:
!python {pt}multiupload.py gene_intersect_CG_wb_b.txt
uploading gene_intersect_CG_wb_b.txt
uploading gene_intersect_CG_wb_b.txt into ['gene_intersect_CG_wb_b.txt']
processing chunk line 0 to 658002 (2.16350102425 s elapsed)
pushing gene_intersect_CG_wb_b.txt...
parsing 1E6747F5...
processing chunk line 658002 to 1315922 (190.930948973 s elapsed)
pushing gene_intersect_CG_wb_b.txt...
parsing C2CCD007...
processing chunk line 1315922 to 1974653 (384.863775969 s elapsed)
pushing gene_intersect_CG_wb_b.txt...
parsing 3D4A4B12...
processing chunk line 1974653 to 2632485 (569.311421871 s elapsed)
pushing gene_intersect_CG_wb_b.txt...
parsing 4897A41D...
Traceback (most recent call last):
File "/Users/sr320/sqlshare-pythonclient/tools/multiupload.py", line 40, in <module>
main()
File "/Users/sr320/sqlshare-pythonclient/tools/multiupload.py", line 37, in main
multiupload(args, options.username, options.password)
File "/Users/sr320/sqlshare-pythonclient/tools/multiupload.py", line 24, in multiupload
for response in conn.upload(globexpr):
File "build/bdist.macosx-10.5-x86_64/egg/sqlshare/__init__.py", line 158, in upload
File "build/bdist.macosx-10.5-x86_64/egg/sqlshare/__init__.py", line 183, in uploadone
File "build/bdist.macosx-10.5-x86_64/egg/sqlshare/__init__.py", line 202, in upload_chunk
File "build/bdist.macosx-10.5-x86_64/egg/sqlshare/__init__.py", line 244, in poll_selector
File "/Users/sr320/anaconda/lib/python2.7/httplib.py", line 1045, in getresponse
response.begin()
File "/Users/sr320/anaconda/lib/python2.7/httplib.py", line 409, in begin
version, status, reason = self._read_status()
File "/Users/sr320/anaconda/lib/python2.7/httplib.py", line 365, in _read_status
line = self.fp.readline(_MAXLINE + 1)
File "/Users/sr320/anaconda/lib/python2.7/socket.py", line 476, in readline
data = self._sock.recv(self._rbufsize)
File "/Users/sr320/anaconda/lib/python2.7/ssl.py", line 241, in recv
return self.read(buflen)
File "/Users/sr320/anaconda/lib/python2.7/ssl.py", line 160, in read
return self._sslobj.read(len)
socket.error: [Errno 54] Connection reset by peer
In [14]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[gene_intersect_DML_dev_wb_b.txt]dev left join [sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]des on dev.Column13=des.CGI_ID" -f tsv -o DML_dev_SPID.txt
In [16]:
!head -2 DML_dev_SPID.txt
!wc -l DML_dev_SPID.txt
176 DML_dev_SPID.txt
In [18]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[gene_intersect_DML_lin_wb_b.txt]lin left join [sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]des on lin.Column13=des.CGI_ID" -f tsv -o DML_lin_SPID.txt
In [19]:
!head -2 DML_lin_SPID.txt
!wc -l DML_lin_SPID.txt
209 DML_lin_SPID.txt
In [52]:
!python {pt}fetchdata.py -d "[sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]" -f tsv -o qdod_sp.txt
In [17]:
!wc -l qdod_sp.txt
18159 qdod_sp.txt
In [27]:
!awk '{print $2}' qdod_sp.txt | sort | uniq | pbcopy
In [28]:
!awk '{print $15}' DML_lin_SPID.txt | sort | uniq | pbcopy
In [29]:
!awk '{print $15}' DML_dev_SPID.txt | sort | uniq | pbcopy
DAVID¶
In [ ]:
Lineage¶
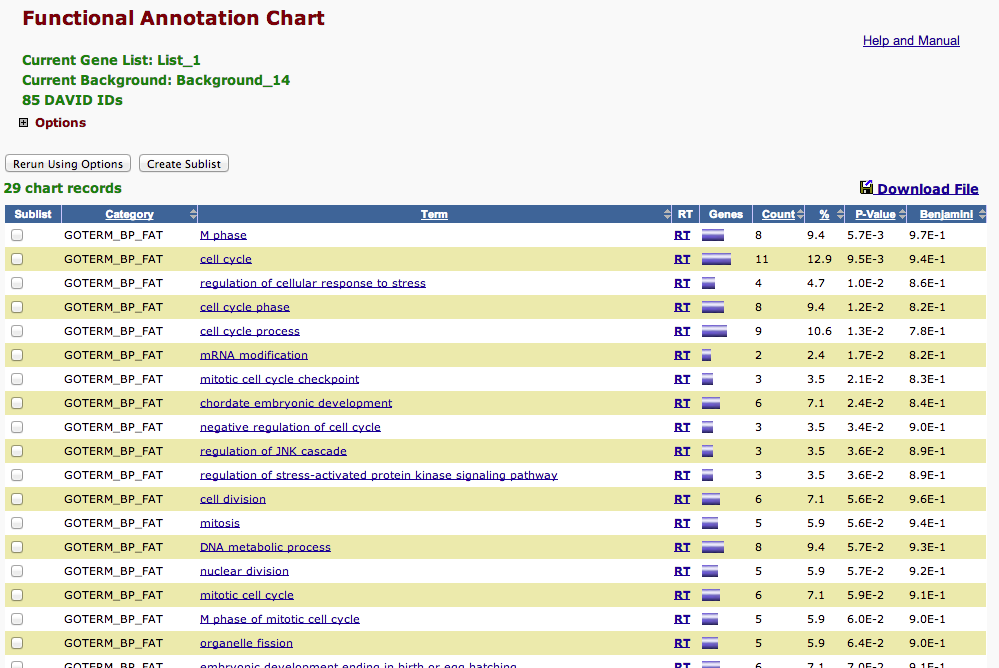
In [69]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953792021.txt
--2014-07-21 07:44:52-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953792021.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 6284 (6.1K) [text/plain] Saving to: `chart_E961EBC140A01405953792021.txt' 100%[======================================>] 6,284 --.-K/s in 0.02s 2014-07-21 07:44:52 (267 KB/s) - `chart_E961EBC140A01405953792021.txt' saved [6284/6284]
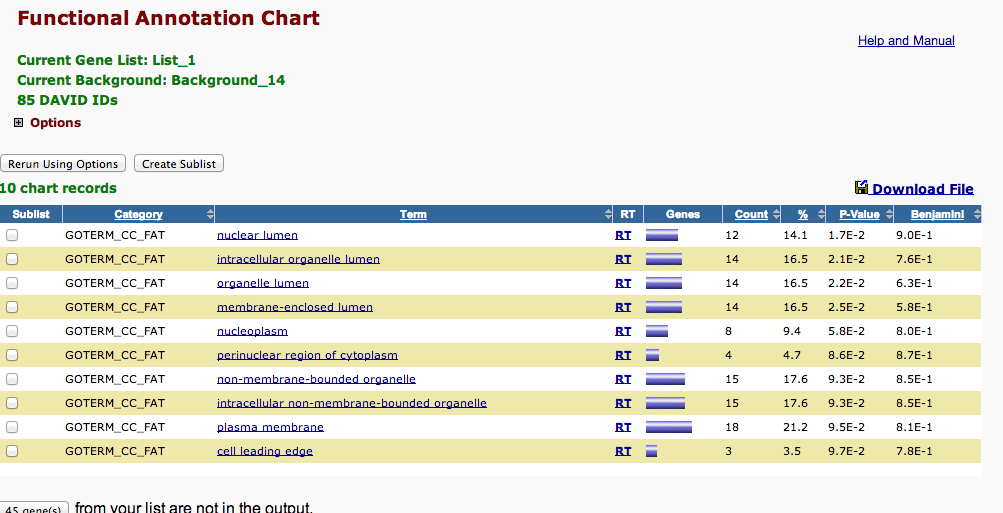
In [70]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953960577.txt
--2014-07-21 07:46:12-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953960577.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 2804 (2.7K) [text/plain] Saving to: `chart_E961EBC140A01405953960577.txt' 100%[======================================>] 2,804 --.-K/s in 0s 2014-07-21 07:46:12 (149 MB/s) - `chart_E961EBC140A01405953960577.txt' saved [2804/2804]
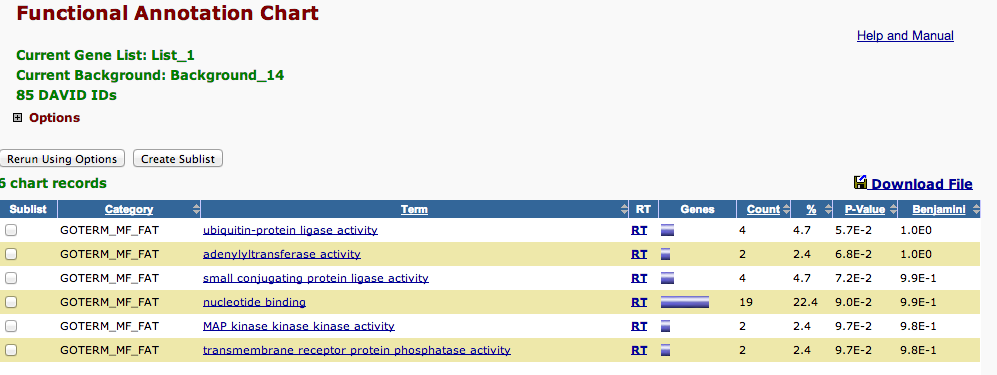
In [72]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953986581.txt
--2014-07-21 07:47:02-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405953986581.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 1480 (1.4K) [text/plain] Saving to: `chart_E961EBC140A01405953986581.txt' 100%[======================================>] 1,480 --.-K/s in 0s 2014-07-21 07:47:03 (54.3 MB/s) - `chart_E961EBC140A01405953986581.txt' saved [1480/1480]
Development¶
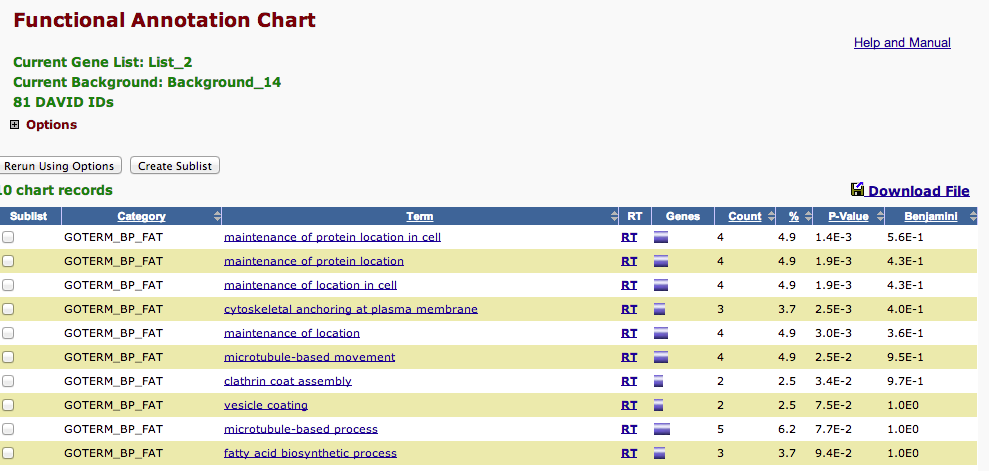
In [73]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954167593.txt
--2014-07-21 07:50:44-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954167593.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 2147 (2.1K) [text/plain] Saving to: `chart_E961EBC140A01405954167593.txt' 100%[======================================>] 2,147 --.-K/s in 0s 2014-07-21 07:50:44 (56.9 MB/s) - `chart_E961EBC140A01405954167593.txt' saved [2147/2147]
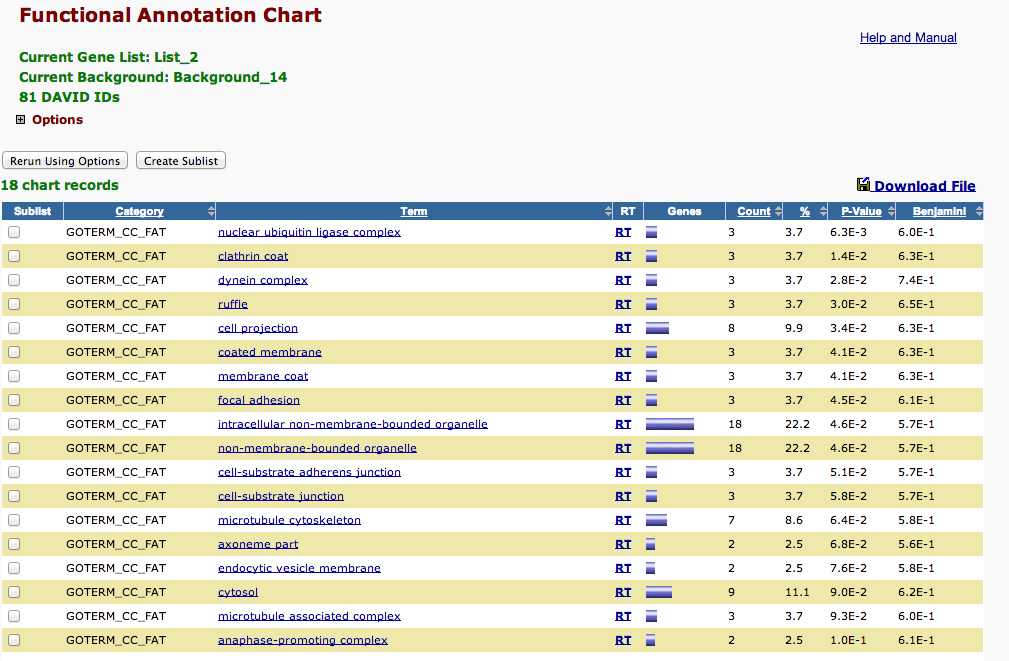
In [74]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954304119.txt
--2014-07-21 07:51:55-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954304119.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 3968 (3.9K) [text/plain] Saving to: `chart_E961EBC140A01405954304119.txt' 100%[======================================>] 3,968 --.-K/s in 0s 2014-07-21 07:51:55 (84.1 MB/s) - `chart_E961EBC140A01405954304119.txt' saved [3968/3968]
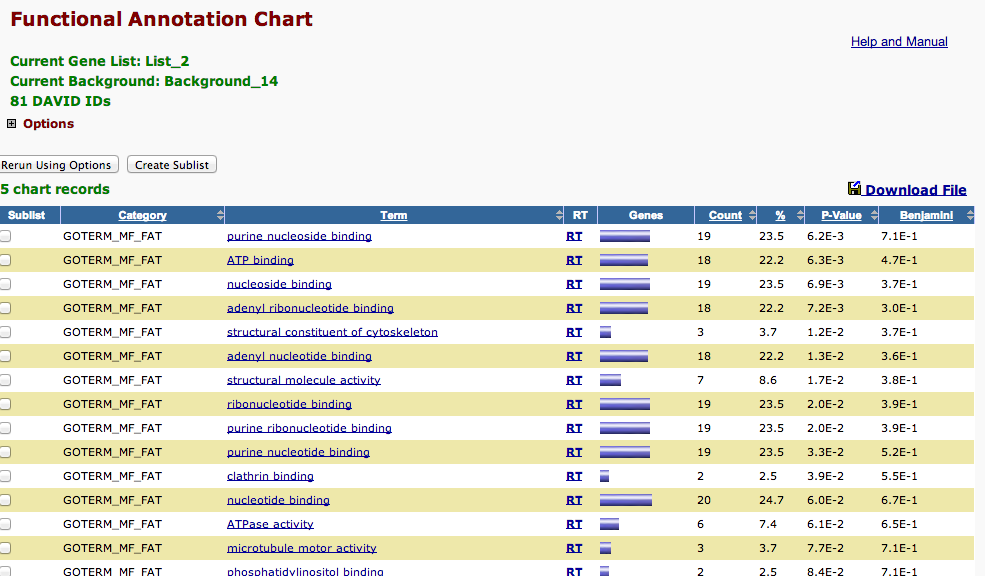
In [75]:
!wget http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954319262.txt
--2014-07-21 07:52:35-- http://david.abcc.ncifcrf.gov/data/download/chart_E961EBC140A01405954319262.txt Resolving david.abcc.ncifcrf.gov... 129.43.1.164 Connecting to david.abcc.ncifcrf.gov|129.43.1.164|:80... connected. HTTP request sent, awaiting response... 200 OK Length: 4310 (4.2K) [text/plain] Saving to: `chart_E961EBC140A01405954319262.txt' 100%[======================================>] 4,310 --.-K/s in 0.07s 2014-07-21 07:52:35 (62.3 KB/s) - `chart_E961EBC140A01405954319262.txt' saved [4310/4310]
Are TEs likely to be found in certain classes of genes?¶
In [ ]:
intersect bed genes on TEs
In [96]:
!intersectbed \
-wb \
-a ./genome_tracks/Cgigas_v9_gene.gff \
-b ./genome_tracks/Cgigas_v9_TE.gff \
> gene_intersect_TE_wb.txt
In [97]:
!head gene_intersect_TE_wb.txt
C22430 GLEAN mRNA 263 493 0.555898 + . ID=CGI_10000081; C22430 WUBlastX DNA_TcMar-Tc2 230 493 60 + . . C23316 GLEAN mRNA 929 1111 0.999999 - . ID=CGI_10000116; C23316 WUBlastX LTR_Ngaro 929 1675 117 - . . C23316 GLEAN mRNA 953 1111 0.999999 - . ID=CGI_10000116; C23316 WUBlastX LTR_Ngaro 953 1489 157 - . . C23316 GLEAN mRNA 644 660 0.999999 - . ID=CGI_10000116; C23316 WUBlastX LTR_Ngaro 547 660 32 - . . C23734 GLEAN mRNA 512 712 1 - . ID=CGI_10000149; C23734 WUBlastX LTR_Gypsy 23 724 246 - . . C23734 GLEAN mRNA 512 712 1 - . ID=CGI_10000149; C23734 WUBlastX LTR_Gypsy 26 964 217 - . . C24320 GLEAN mRNA 1545 1904 0.999372 - . ID=CGI_10000190; C24320 WUBlastX LTR_ERV1 1545 1904 27 - . . C24604 GLEAN mRNA 792 989 0.997918 - . ID=CGI_10000200; C24604 WUBlastX DNA_MuDR 792 989 42 - . . C25496 GLEAN mRNA 2006 2074 0.998748 + . ID=CGI_10000243; C25496 WUBlastX LINE_CR1 2006 2536 27 + . . C26320 GLEAN mRNA 2740 2964 0.481188 + . ID=CGI_10000297; C26320 WUBlastX LINE_L2 2710 2964 75 + . .
In [99]:
!intersectbed \
-c \
-a ./genome_tracks/Cgigas_v9_gene.gff \
-b ./genome_tracks/Cgigas_v9_TE.gff \
> gene_intersect_TE_c.txt
!head gene_intersect_TE_c.txt
C16582 GLEAN mRNA 35 385 0.555898 - . ID=CGI_10000001; 0 C17212 GLEAN mRNA 31 363 0.999572 + . ID=CGI_10000002; 0 C17316 GLEAN mRNA 30 257 0.555898 + . ID=CGI_10000003; 0 C17476 GLEAN mRNA 34 257 0.998947 - . ID=CGI_10000004; 0 C17998 GLEAN mRNA 196 387 1 - . ID=CGI_10000005; 0 C18346 GLEAN mRNA 174 551 1 + . ID=CGI_10000009; 0 C18428 GLEAN mRNA 286 546 0.555898 - . ID=CGI_10000010; 0 C18964 GLEAN mRNA 203 658 0.999572 - . ID=CGI_10000011; 0 C18980 GLEAN mRNA 30 674 0.555898 + . ID=CGI_10000012; 0 C19100 GLEAN mRNA 160 681 0.999955 - . ID=CGI_10000013; 0
In [111]:
!awk '{print $9,$10}' gene_intersect_TE_c.txt > gene_intersect_TE_c2.txt
!sed 's/ID=C/C/g' <gene_intersect_TE_c2.txt> gene_intersect_TE_c2c.txt
!sed 's/;//g' <gene_intersect_TE_c2c.txt> gene_intersect_TE_c2ge.txt
!echo "CGI_ID Num_TEs" >> gTE.head
!cat gTE.head gene_intersect_TE_c2ge.txt > gene_intersect_TE_c2g.txt
!head -3 gene_intersect_TE_c2g.txt
!python {pt}singleupload.py gene_intersect_TE_c2g.txt
CGI_ID Num_TEs CGI_10000001 0 CGI_10000002 0 processing chunk line 0 to 28028 (0.00581192970276 s elapsed) pushing gene_intersect_TE_c2g.txt... parsing 4AD03B75... finished gene_intersect_TE_c2g.txt
In [116]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[gene_intersect_TE_c2g.txt]te left join [sr320@washington.edu].[qDOD_Cgigas_GOslim_DISTINCT]slim on te.CGI_ID=slim.CGI_ID where [aspect] = 'P'" -f tsv -o gene_i_TE_geneGOslim.txt
In [117]:
!head gene_i_TE_geneGOslim.txt
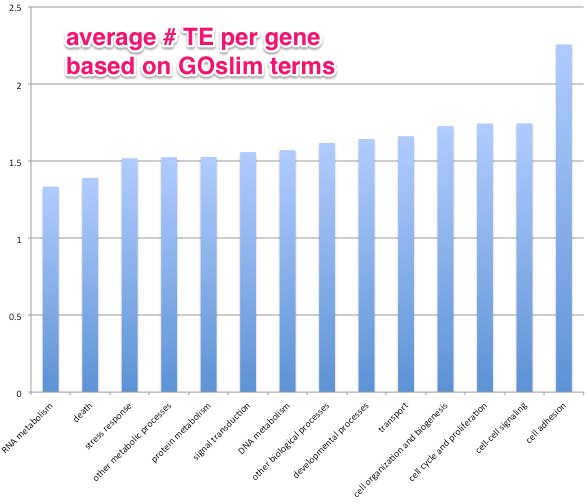
GO Slim - DML¶
In [120]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[gene_intersect_DML_lin_wb_b.txt]lin left join [sr320@washington.edu].[qDOD_Cgigas_GOslim_DISTINCT]slim on lin.Column13=slim.CGI_ID where [aspect] = 'P'" -f tsv -o DML_lin_geneGOslimDistinct.txt
In [122]:
!head -2 DML_lin_geneGOslimDistinct.txt
In [123]:
!wc -l DML_lin_geneGOslimDistinct.txt
358 DML_lin_geneGOslimDistinct.txt
In [124]:
!awk '{print $15}' DML_lin_geneGOslimDistinct.txt > DML_lin_geneGOslimDistinct_15.txt
!<DML_lin_geneGOslimDistinct_15.txt cut -d' ' -f1 | sort -n | uniq -c
11 DNA 1 GOslim_bin 18 RNA 75 cell 5 cell-cell 13 death 26 developmental 101 other 41 protein 21 signal 21 stress 25 transport
In [125]:
!python {pt}fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[gene_intersect_DML_dev_wb_b.txt]dev left join [sr320@washington.edu].[qDOD_Cgigas_GOslim_DISTINCT]slim on dev.Column13=slim.CGI_ID where [aspect] = 'P'" -f tsv -o DML_dev_geneGOslimDistinct.txt
In [127]:
!wc -l DML_dev_geneGOslimDistinct.txt
284 DML_dev_geneGOslimDistinct.txt
In [126]:
!awk '{print $15}' DML_dev_geneGOslimDistinct.txt > DML_dev_geneGOslimDistinct_15.txt
!<DML_dev_geneGOslimDistinct_15.txt cut -d' ' -f1 | sort -n | uniq -c
8 DNA 1 GOslim_bin 15 RNA 52 cell 6 cell-cell 6 death 25 developmental 78 other 30 protein 16 signal 17 stress 30 transport
In [ ]: