BSMAP-methatio-feature track¶
In [81]:
#trying to get clean version
!date
Sun Mar 16 04:32:55 PDT 2014
Concatenate technical replicates¶
September is known as FCD2CA8: November Core ID FCC39EM
M1¶
In [11]:
#Will go September then Novemeber
!cat /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgM1/filtered_BS_CgM1_ACTTGA_L007_R1.fastq \
/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgM1/filtered_BS_CgM1_ACTTGA_L004_R1.fastq > \
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_R1a.fastq \
In [12]:
!cat /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgM1/filtered_BS_CgM1_ACTTGA_L007_R2.fastq \
/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgM1/filtered_BS_CgM1_ACTTGA_L004_R2.fastq > \
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_R2a.fastq \
M3¶
In [20]:
R1_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgM3/filtered_BS_CgM3_GATCAG_L007_R1.fastq"
R1_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgM3/filtered_BS_CgM3_GATCAG_L004_R1.fastq"
R2_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgM3/filtered_BS_CgM3_GATCAG_L007_R2.fastq"
R2_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgM3/filtered_BS_CgM3_GATCAG_L004_R2.fastq"
In [21]:
!cat {R1_A8} \
{R1_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_R1a.fastq \
In [22]:
!cat {R2_A8} \
{R2_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_R2a.fastq \
T1D3¶
In [23]:
sample="T1D3"
In [13]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L004_R1.fastq.gz
In [14]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L004_R2.fastq.gz
In [24]:
R1_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L007_R1.fastq"
R1_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L004_R1.fastq"
R2_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L007_R2.fastq"
R2_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D3/filtered_BS_CgLarv_T1D3_TGACCA_L004_R2.fastq"
In [25]:
!cat {R1_A8} \
{R1_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R1a.fastq \
In [26]:
!cat {R2_A8} \
{R2_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R2a.fastq \
T1D5¶
In [ ]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R1.fastq.gz
In [29]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R2.fastq.gz
In [31]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L004_R1.fastq.gz
In [32]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L004_R2.fastq.gz
In [33]:
R1_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R1.fastq"
R1_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L004_R1.fastq"
R2_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L007_R2.fastq"
R2_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T1D5/filtered_BS_CgLarv_T1D5_ACAGTG_L004_R2.fastq"
In [34]:
sample="T1D5"
In [35]:
!cat {R1_A8} \
{R1_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R1a.fastq \
In [36]:
!cat {R2_A8} \
{R2_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R2a.fastq \
T3D3¶
In [51]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L007*
In [ ]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L004*
In [58]:
R1_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R1.fastq"
R1_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L004_R1.fastq"
R2_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L007_R2.fastq"
R2_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_Bs_CgLarve_T3D3/filtered_Bs_CgLarve_T3D3_GCCAAT_L004_R2.fastq"
In [59]:
sample="T3D3"
In [60]:
!cat {R1_A8} \
{R1_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R1a.fastq \
In [61]:
!cat {R2_A8} \
{R2_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R2a.fastq \
T3D5¶
In [38]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T3D5/filtered_*
In [39]:
!gunzip /Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T3D5/filtered_*
In [40]:
R1_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T3D5/filtered_BS_CgLarv_T3D5_CAGATC_L007_R1.fastq"
R1_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T3D5/filtered_BS_CgLarv_T3D5_CAGATC_L004_R1.fastq"
R2_A8="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCD2CA8_2/Sample_BS_CgLarv_T3D5/filtered_BS_CgLarv_T3D5_CAGATC_L007_R2.fastq"
R2_EM="/Volumes/web/trilobite/Crassostrea_gigas_HTSdata/batterbox/FCC39EM_02/FCC39EM/Sample_BS_CgLarv_T3D5/filtered_BS_CgLarv_T3D5_CAGATC_L004_R2.fastq"
In [41]:
sample="T3D5"
In [42]:
!cat {R1_A8} \
{R1_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R1a.fastq \
In [ ]:
!cat {R2_A8} \
{R2_EM} > \
/Volumes/web/cnidarian/BiGo_larvae_merge/Cg{sample}_R2a.fastq \
BSMAP¶
In [1]:
#file ID
fid="CgM1"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd='/Volumes/web/cnidarian/BiGo_larvae_merge/'
#where is bsmap
#bsmap="/Users/Shared/Apps/bsmap-2.73/"
bsmap="/Volumes/Bay3/Software/BSMAP/bsmap-2.74/"
#genome file
genome="/Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa"
In [2]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [3]:
mkdir {fid}_{date}
In [4]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]
In [5]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 09:54:25 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 13 secs passed total_kmers: 43046721 Create seed table. 55 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #0: 150000 read pairs finished. 98 secs passed Thread #3: 200000 read pairs finished. 98 secs passed Thread #1: 100000 read pairs finished. 98 secs passed Thread #2: 50000 read pairs finished. 99 secs passed Thread #0: 250000 read pairs finished. 137 secs passed Thread #3: 300000 read pairs finished. 138 secs passed Thread #1: 350000 read pairs finished. 139 secs passed Thread #2: 400000 read pairs finished. 140 secs passed Thread #0: 450000 read pairs finished. 178 secs passed Thread #3: 500000 read pairs finished. 179 secs passed Thread #1: 550000 read pairs finished. 180 secs passed Thread #2: 600000 read pairs finished. 181 secs passed Thread #0: 650000 read pairs finished. 219 secs passed Thread #3: 700000 read pairs finished. 220 secs passed Thread #1: 750000 read pairs finished. 223 secs passed Thread #2: 800000 read pairs finished. 224 secs passed Thread #0: 850000 read pairs finished. 262 secs passed Thread #3: 900000 read pairs finished. 263 secs passed Thread #1: 950000 read pairs finished. 271 secs passed Thread #2: 1000000 read pairs finished. 273 secs passed Thread #3: 1100000 read pairs finished. 308 secs passed Thread #0: 1050000 read pairs finished. 309 secs passed Thread #1: 1150000 read pairs finished. 313 secs passed Thread #2: 1200000 read pairs finished. 316 secs passed Thread #3: 1250000 read pairs finished. 352 secs passed Thread #0: 1300000 read pairs finished. 353 secs passed Thread #1: 1350000 read pairs finished. 358 secs passed Thread #2: 1400000 read pairs finished. 359 secs passed Thread #3: 1450000 read pairs finished. 393 secs passed Thread #0: 1500000 read pairs finished. 395 secs passed Thread #1: 1550000 read pairs finished. 399 secs passed Thread #2: 1600000 read pairs finished. 405 secs passed Thread #3: 1650000 read pairs finished. 438 secs passed Thread #0: 1700000 read pairs finished. 442 secs passed Thread #1: 1750000 read pairs finished. 444 secs passed Thread #2: 1800000 read pairs finished. 455 secs passed Thread #3: 1850000 read pairs finished. 491 secs passed Thread #0: 1900000 read pairs finished. 496 secs passed Thread #1: 1950000 read pairs finished. 498 secs passed Thread #2: 2000000 read pairs finished. 508 secs passed Thread #3: 2050000 read pairs finished. 541 secs passed Thread #0: 2100000 read pairs finished. 545 secs passed Thread #1: 2150000 read pairs finished. 547 secs passed Thread #2: 2200000 read pairs finished. 556 secs passed Thread #3: 2250000 read pairs finished. 585 secs passed Thread #0: 2300000 read pairs finished. 589 secs passed Thread #1: 2350000 read pairs finished. 590 secs passed Thread #2: 2400000 read pairs finished. 597 secs passed Thread #3: 2450000 read pairs finished. 621 secs passed Thread #0: 2500000 read pairs finished. 625 secs passed Thread #1: 2550000 read pairs finished. 625 secs passed Thread #2: 2600000 read pairs finished. 632 secs passed Thread #3: 2650000 read pairs finished. 658 secs passed Thread #0: 2700000 read pairs finished. 663 secs passed Thread #1: 2750000 read pairs finished. 664 secs passed Thread #2: 2800000 read pairs finished. 670 secs passed Thread #3: 2850000 read pairs finished. 694 secs passed Thread #0: 2900000 read pairs finished. 699 secs passed Thread #1: 2950000 read pairs finished. 700 secs passed Thread #2: 3000000 read pairs finished. 707 secs passed Thread #3: 3050000 read pairs finished. 730 secs passed Thread #0: 3100000 read pairs finished. 736 secs passed Thread #1: 3150000 read pairs finished. 737 secs passed Thread #2: 3200000 read pairs finished. 743 secs passed Thread #3: 3250000 read pairs finished. 766 secs passed Thread #0: 3300000 read pairs finished. 773 secs passed Thread #1: 3350000 read pairs finished. 774 secs passed Thread #2: 3400000 read pairs finished. 781 secs passed Thread #3: 3450000 read pairs finished. 807 secs passed Thread #0: 3500000 read pairs finished. 815 secs passed Thread #1: 3550000 read pairs finished. 816 secs passed Thread #2: 3600000 read pairs finished. 821 secs passed Thread #3: 3650000 read pairs finished. 844 secs passed Thread #0: 3700000 read pairs finished. 854 secs passed Thread #1: 3750000 read pairs finished. 858 secs passed Thread #2: 3800000 read pairs finished. 860 secs passed Thread #3: 3850000 read pairs finished. 881 secs passed Thread #0: 3900000 read pairs finished. 891 secs passed Thread #1: 3950000 read pairs finished. 894 secs passed Thread #2: 4000000 read pairs finished. 896 secs passed Thread #3: 4050000 read pairs finished. 916 secs passed Thread #0: 4100000 read pairs finished. 928 secs passed Thread #1: 4150000 read pairs finished. 930 secs passed Thread #2: 4200000 read pairs finished. 931 secs passed Thread #3: 4250000 read pairs finished. 953 secs passed Thread #0: 4300000 read pairs finished. 966 secs passed Thread #1: 4350000 read pairs finished. 968 secs passed Thread #2: 4400000 read pairs finished. 969 secs passed Thread #3: 4450000 read pairs finished. 989 secs passed Thread #0: 4500000 read pairs finished. 1002 secs passed Thread #1: 4550000 read pairs finished. 1003 secs passed Thread #2: 4600000 read pairs finished. 1006 secs passed Thread #3: 4650000 read pairs finished. 1026 secs passed Thread #0: 4700000 read pairs finished. 1038 secs passed Thread #1: 4750000 read pairs finished. 1038 secs passed Thread #2: 4800000 read pairs finished. 1041 secs passed Thread #3: 4850000 read pairs finished. 1062 secs passed Thread #0: 4900000 read pairs finished. 1074 secs passed Thread #1: 4950000 read pairs finished. 1074 secs passed Thread #2: 5000000 read pairs finished. 1076 secs passed Thread #3: 5050000 read pairs finished. 1098 secs passed Thread #0: 5100000 read pairs finished. 1111 secs passed Thread #1: 5150000 read pairs finished. 1112 secs passed Thread #2: 5200000 read pairs finished. 1113 secs passed Thread #3: 5250000 read pairs finished. 1134 secs passed Thread #0: 5300000 read pairs finished. 1147 secs passed Thread #1: 5350000 read pairs finished. 1148 secs passed Thread #2: 5400000 read pairs finished. 1150 secs passed Thread #3: 5450000 read pairs finished. 1171 secs passed Thread #0: 5500000 read pairs finished. 1184 secs passed Thread #1: 5550000 read pairs finished. 1185 secs passed Thread #2: 5600000 read pairs finished. 1187 secs passed Thread #3: 5650000 read pairs finished. 1208 secs passed Thread #0: 5700000 read pairs finished. 1221 secs passed Thread #1: 5750000 read pairs finished. 1222 secs passed Thread #2: 5800000 read pairs finished. 1223 secs passed Thread #3: 5850000 read pairs finished. 1244 secs passed Thread #0: 5900000 read pairs finished. 1259 secs passed Thread #1: 5950000 read pairs finished. 1260 secs passed Thread #2: 6000000 read pairs finished. 1261 secs passed Thread #3: 6050000 read pairs finished. 1281 secs passed Thread #0: 6100000 read pairs finished. 1297 secs passed Thread #1: 6150000 read pairs finished. 1298 secs passed Thread #2: 6200000 read pairs finished. 1299 secs passed Thread #3: 6250000 read pairs finished. 1318 secs passed Thread #0: 6300000 read pairs finished. 1334 secs passed Thread #1: 6350000 read pairs finished. 1335 secs passed Thread #2: 6400000 read pairs finished. 1336 secs passed Thread #3: 6450000 read pairs finished. 1355 secs passed Thread #0: 6500000 read pairs finished. 1371 secs passed Thread #1: 6550000 read pairs finished. 1371 secs passed Thread #2: 6600000 read pairs finished. 1373 secs passed Thread #3: 6650000 read pairs finished. 1392 secs passed Thread #0: 6700000 read pairs finished. 1407 secs passed Thread #1: 6750000 read pairs finished. 1408 secs passed Thread #2: 6800000 read pairs finished. 1410 secs passed Thread #3: 6850000 read pairs finished. 1428 secs passed Thread #0: 6900000 read pairs finished. 1445 secs passed Thread #1: 6950000 read pairs finished. 1445 secs passed Thread #2: 7000000 read pairs finished. 1447 secs passed Thread #3: 7050000 read pairs finished. 1465 secs passed Thread #0: 7100000 read pairs finished. 1482 secs passed Thread #1: 7150000 read pairs finished. 1482 secs passed Thread #2: 7200000 read pairs finished. 1483 secs passed Thread #3: 7250000 read pairs finished. 1501 secs passed Thread #0: 7300000 read pairs finished. 1520 secs passed Thread #1: 7350000 read pairs finished. 1521 secs passed Thread #2: 7400000 read pairs finished. 1522 secs passed Thread #3: 7450000 read pairs finished. 1538 secs passed Thread #0: 7500000 read pairs finished. 1557 secs passed Thread #1: 7550000 read pairs finished. 1558 secs passed Thread #2: 7600000 read pairs finished. 1559 secs passed Thread #3: 7650000 read pairs finished. 1575 secs passed Thread #0: 7700000 read pairs finished. 1595 secs passed Thread #1: 7750000 read pairs finished. 1595 secs passed Thread #2: 7800000 read pairs finished. 1597 secs passed Thread #3: 7850000 read pairs finished. 1611 secs passed Thread #0: 7900000 read pairs finished. 1632 secs passed Thread #1: 7950000 read pairs finished. 1633 secs passed Thread #2: 8000000 read pairs finished. 1634 secs passed Thread #3: 8050000 read pairs finished. 1648 secs passed Thread #0: 8100000 read pairs finished. 1669 secs passed Thread #1: 8150000 read pairs finished. 1670 secs passed Thread #2: 8200000 read pairs finished. 1671 secs passed Thread #3: 8250000 read pairs finished. 1679 secs passed Thread #0: 8297202 read pairs finished. 1689 secs passed Total number of aligned reads: pairs: 3832546 (46%) single a: 1591687 (19%) single b: 1349271 (16%) Done. Finished at Sun Mar 16 10:22:35 2014 Total time consumed: 1690 secs @ Sun Mar 16 10:22:36 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 10:23:28 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 10:31:20 2014: read 10000000 lines @ Sun Mar 16 10:31:49 2014: combining CpG methylation from both strands ... @ Sun Mar 16 10:32:31 2014: writing methratio_out.txt ... @ Sun Mar 16 10:47:08 2014: done. total 8562355 valid mappings, 48968967 covered cytosines, average coverage: 1.80 fold.
In [6]:
!head filt_methratio_{fid}.igv
C12768 102 104 CpG 0.167 C12806 141 143 CpG 0.333 C12924 29 31 CpG 0.000 C12924 37 39 CpG 0.000 C12924 51 53 CpG 0.000 C12924 59 61 CpG 0.000 C12924 126 128 CpG 0.000 C12924 135 137 CpG 0.000 C13128 86 88 CpG 0.000 C13208 82 84 CpG 0.400
In [8]:
#file ID
fid="CgM3"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd='/Volumes/web/cnidarian/BiGo_larvae_merge/'
In [10]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [12]:
mkdir {fid}_{date}
In [13]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]
In [14]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 12:08:52 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 22 secs passed total_kmers: 43046721 Create seed table. 63 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #3: 200000 read pairs finished. 98 secs passed Thread #1: 100000 read pairs finished. 98 secs passed Thread #0: 50000 read pairs finished. 99 secs passed Thread #2: 150000 read pairs finished. 99 secs passed Thread #3: 250000 read pairs finished. 131 secs passed Thread #1: 300000 read pairs finished. 132 secs passed Thread #2: 400000 read pairs finished. 133 secs passed Thread #0: 350000 read pairs finished. 133 secs passed Thread #3: 450000 read pairs finished. 163 secs passed Thread #1: 500000 read pairs finished. 165 secs passed Thread #2: 550000 read pairs finished. 167 secs passed Thread #0: 600000 read pairs finished. 168 secs passed Thread #3: 650000 read pairs finished. 199 secs passed Thread #1: 700000 read pairs finished. 201 secs passed Thread #2: 750000 read pairs finished. 203 secs passed Thread #0: 800000 read pairs finished. 205 secs passed Thread #3: 850000 read pairs finished. 237 secs passed Thread #1: 900000 read pairs finished. 240 secs passed Thread #2: 950000 read pairs finished. 246 secs passed Thread #0: 1000000 read pairs finished. 248 secs passed Thread #3: 1050000 read pairs finished. 274 secs passed Thread #1: 1100000 read pairs finished. 276 secs passed Thread #2: 1150000 read pairs finished. 280 secs passed Thread #0: 1200000 read pairs finished. 283 secs passed Thread #3: 1250000 read pairs finished. 308 secs passed Thread #1: 1300000 read pairs finished. 309 secs passed Thread #2: 1350000 read pairs finished. 313 secs passed Thread #0: 1400000 read pairs finished. 317 secs passed Thread #3: 1450000 read pairs finished. 341 secs passed Thread #1: 1500000 read pairs finished. 342 secs passed Thread #2: 1550000 read pairs finished. 346 secs passed Thread #0: 1600000 read pairs finished. 350 secs passed Thread #3: 1650000 read pairs finished. 374 secs passed Thread #1: 1700000 read pairs finished. 376 secs passed Thread #2: 1750000 read pairs finished. 380 secs passed Thread #0: 1800000 read pairs finished. 384 secs passed Thread #3: 1850000 read pairs finished. 408 secs passed Thread #1: 1900000 read pairs finished. 409 secs passed Thread #2: 1950000 read pairs finished. 413 secs passed Thread #0: 2000000 read pairs finished. 417 secs passed Thread #3: 2050000 read pairs finished. 440 secs passed Thread #1: 2100000 read pairs finished. 444 secs passed Thread #2: 2150000 read pairs finished. 446 secs passed Thread #0: 2200000 read pairs finished. 451 secs passed Thread #3: 2250000 read pairs finished. 473 secs passed Thread #1: 2300000 read pairs finished. 477 secs passed Thread #2: 2350000 read pairs finished. 479 secs passed Thread #0: 2400000 read pairs finished. 484 secs passed Thread #3: 2450000 read pairs finished. 507 secs passed Thread #1: 2500000 read pairs finished. 510 secs passed Thread #2: 2550000 read pairs finished. 512 secs passed Thread #0: 2600000 read pairs finished. 518 secs passed Thread #3: 2650000 read pairs finished. 540 secs passed Thread #1: 2700000 read pairs finished. 544 secs passed Thread #2: 2750000 read pairs finished. 546 secs passed Thread #0: 2800000 read pairs finished. 553 secs passed Thread #3: 2850000 read pairs finished. 573 secs passed Thread #1: 2900000 read pairs finished. 577 secs passed Thread #2: 2950000 read pairs finished. 580 secs passed Thread #0: 3000000 read pairs finished. 586 secs passed Thread #3: 3050000 read pairs finished. 607 secs passed Thread #1: 3100000 read pairs finished. 611 secs passed Thread #2: 3150000 read pairs finished. 614 secs passed Thread #0: 3200000 read pairs finished. 620 secs passed Thread #3: 3250000 read pairs finished. 640 secs passed Thread #1: 3300000 read pairs finished. 644 secs passed Thread #2: 3350000 read pairs finished. 647 secs passed Thread #0: 3400000 read pairs finished. 654 secs passed Thread #3: 3450000 read pairs finished. 674 secs passed Thread #1: 3500000 read pairs finished. 678 secs passed Thread #2: 3550000 read pairs finished. 681 secs passed Thread #0: 3600000 read pairs finished. 688 secs passed Thread #3: 3650000 read pairs finished. 708 secs passed Thread #1: 3700000 read pairs finished. 714 secs passed Thread #2: 3750000 read pairs finished. 719 secs passed Thread #0: 3800000 read pairs finished. 726 secs passed Thread #3: 3850000 read pairs finished. 743 secs passed Thread #1: 3900000 read pairs finished. 749 secs passed Thread #2: 3950000 read pairs finished. 754 secs passed Thread #0: 4000000 read pairs finished. 759 secs passed Thread #3: 4050000 read pairs finished. 776 secs passed Thread #1: 4100000 read pairs finished. 783 secs passed Thread #2: 4150000 read pairs finished. 788 secs passed Thread #0: 4200000 read pairs finished. 793 secs passed Thread #3: 4250000 read pairs finished. 810 secs passed Thread #1: 4300000 read pairs finished. 816 secs passed Thread #2: 4350000 read pairs finished. 821 secs passed Thread #0: 4400000 read pairs finished. 827 secs passed Thread #3: 4450000 read pairs finished. 843 secs passed Thread #1: 4500000 read pairs finished. 849 secs passed Thread #2: 4550000 read pairs finished. 853 secs passed Thread #0: 4600000 read pairs finished. 860 secs passed Thread #3: 4650000 read pairs finished. 877 secs passed Thread #1: 4700000 read pairs finished. 883 secs passed Thread #2: 4750000 read pairs finished. 886 secs passed Thread #0: 4800000 read pairs finished. 894 secs passed Thread #3: 4850000 read pairs finished. 910 secs passed Thread #1: 4900000 read pairs finished. 917 secs passed Thread #2: 4950000 read pairs finished. 921 secs passed Thread #0: 5000000 read pairs finished. 928 secs passed Thread #3: 5050000 read pairs finished. 945 secs passed Thread #1: 5100000 read pairs finished. 951 secs passed Thread #2: 5150000 read pairs finished. 954 secs passed Thread #0: 5200000 read pairs finished. 962 secs passed Thread #3: 5250000 read pairs finished. 980 secs passed Thread #1: 5300000 read pairs finished. 986 secs passed Thread #2: 5350000 read pairs finished. 988 secs passed Thread #0: 5400000 read pairs finished. 996 secs passed Thread #3: 5450000 read pairs finished. 1013 secs passed Thread #1: 5500000 read pairs finished. 1020 secs passed Thread #2: 5550000 read pairs finished. 1022 secs passed Thread #0: 5600000 read pairs finished. 1031 secs passed Thread #3: 5650000 read pairs finished. 1047 secs passed Thread #1: 5700000 read pairs finished. 1054 secs passed Thread #2: 5750000 read pairs finished. 1056 secs passed Thread #0: 5800000 read pairs finished. 1066 secs passed Thread #3: 5850000 read pairs finished. 1081 secs passed Thread #1: 5900000 read pairs finished. 1088 secs passed Thread #2: 5950000 read pairs finished. 1090 secs passed Thread #0: 6000000 read pairs finished. 1101 secs passed Thread #3: 6050000 read pairs finished. 1115 secs passed Thread #1: 6100000 read pairs finished. 1123 secs passed Thread #2: 6150000 read pairs finished. 1124 secs passed Thread #0: 6200000 read pairs finished. 1134 secs passed Thread #3: 6250000 read pairs finished. 1150 secs passed Thread #1: 6300000 read pairs finished. 1159 secs passed Thread #2: 6350000 read pairs finished. 1160 secs passed Thread #0: 6400000 read pairs finished. 1173 secs passed Thread #3: 6450000 read pairs finished. 1190 secs passed Thread #1: 6500000 read pairs finished. 1199 secs passed Thread #2: 6550000 read pairs finished. 1200 secs passed Thread #0: 6600000 read pairs finished. 1212 secs passed Thread #3: 6650000 read pairs finished. 1228 secs passed Thread #1: 6700000 read pairs finished. 1237 secs passed Thread #2: 6750000 read pairs finished. 1238 secs passed Thread #0: 6800000 read pairs finished. 1250 secs passed Thread #3: 6850000 read pairs finished. 1265 secs passed Thread #1: 6900000 read pairs finished. 1275 secs passed Thread #2: 6950000 read pairs finished. 1276 secs passed Thread #0: 7000000 read pairs finished. 1286 secs passed Thread #3: 7050000 read pairs finished. 1300 secs passed Thread #1: 7100000 read pairs finished. 1309 secs passed Thread #2: 7150000 read pairs finished. 1309 secs passed Thread #0: 7200000 read pairs finished. 1319 secs passed Thread #3: 7250000 read pairs finished. 1333 secs passed Thread #1: 7300000 read pairs finished. 1344 secs passed Thread #2: 7350000 read pairs finished. 1344 secs passed Thread #0: 7400000 read pairs finished. 1354 secs passed Thread #3: 7450000 read pairs finished. 1367 secs passed Thread #1: 7500000 read pairs finished. 1378 secs passed Thread #2: 7550000 read pairs finished. 1380 secs passed Thread #0: 7600000 read pairs finished. 1388 secs passed Thread #3: 7650000 read pairs finished. 1400 secs passed Thread #1: 7700000 read pairs finished. 1412 secs passed Thread #2: 7750000 read pairs finished. 1414 secs passed Thread #0: 7800000 read pairs finished. 1423 secs passed Thread #3: 7850000 read pairs finished. 1435 secs passed Thread #1: 7900000 read pairs finished. 1448 secs passed Thread #2: 7950000 read pairs finished. 1449 secs passed Thread #0: 8000000 read pairs finished. 1456 secs passed Thread #1: 8078115 read pairs finished. 1461 secs passed Thread #3: 8050000 read pairs finished. 1463 secs passed Total number of aligned reads: pairs: 4380128 (54%) single a: 1426879 (18%) single b: 1168161 (14%) Done. Finished at Sun Mar 16 12:33:15 2014 Total time consumed: 1463 secs @ Sun Mar 16 12:33:17 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 12:34:09 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 12:43:14 2014: read 10000000 lines @ Sun Mar 16 12:44:27 2014: combining CpG methylation from both strands ... @ Sun Mar 16 12:45:10 2014: writing methratio_out.txt ... @ Sun Mar 16 13:01:19 2014: done. total 9597901 valid mappings, 53820491 covered cytosines, average coverage: 1.79 fold.
In [15]:
#file ID
fid="CgT1D3"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd="/Volumes/web/cnidarian/BiGo_larvae_merge/"
In [16]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [17]:
mkdir {fid}_{date}
In [18]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_[0316_1306]
In [19]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 13:06:42 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 22 secs passed total_kmers: 43046721 Create seed table. 63 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #1: 100000 read pairs finished. 101 secs passed Thread #0: 50000 read pairs finished. 101 secs passed Thread #3: 150000 read pairs finished. 101 secs passed Thread #2: 200000 read pairs finished. 102 secs passed Thread #1: 250000 read pairs finished. 134 secs passed Thread #0: 300000 read pairs finished. 135 secs passed Thread #3: 350000 read pairs finished. 136 secs passed Thread #2: 400000 read pairs finished. 138 secs passed Thread #1: 450000 read pairs finished. 168 secs passed Thread #0: 500000 read pairs finished. 170 secs passed Thread #3: 550000 read pairs finished. 171 secs passed Thread #2: 600000 read pairs finished. 177 secs passed Thread #1: 650000 read pairs finished. 205 secs passed Thread #0: 700000 read pairs finished. 206 secs passed Thread #3: 750000 read pairs finished. 206 secs passed Thread #2: 800000 read pairs finished. 212 secs passed Thread #1: 850000 read pairs finished. 240 secs passed Thread #0: 900000 read pairs finished. 241 secs passed Thread #3: 950000 read pairs finished. 242 secs passed Thread #2: 1000000 read pairs finished. 246 secs passed Thread #1: 1050000 read pairs finished. 274 secs passed Thread #0: 1100000 read pairs finished. 276 secs passed Thread #3: 1150000 read pairs finished. 278 secs passed Thread #2: 1200000 read pairs finished. 281 secs passed Thread #1: 1250000 read pairs finished. 309 secs passed Thread #0: 1300000 read pairs finished. 311 secs passed Thread #3: 1350000 read pairs finished. 313 secs passed Thread #2: 1400000 read pairs finished. 316 secs passed Thread #1: 1450000 read pairs finished. 344 secs passed Thread #0: 1500000 read pairs finished. 346 secs passed Thread #3: 1550000 read pairs finished. 349 secs passed Thread #2: 1600000 read pairs finished. 352 secs passed Thread #1: 1650000 read pairs finished. 380 secs passed Thread #0: 1700000 read pairs finished. 383 secs passed Thread #3: 1750000 read pairs finished. 384 secs passed Thread #2: 1800000 read pairs finished. 387 secs passed Thread #1: 1850000 read pairs finished. 415 secs passed Thread #0: 1900000 read pairs finished. 418 secs passed Thread #3: 1950000 read pairs finished. 419 secs passed Thread #2: 2000000 read pairs finished. 423 secs passed Thread #1: 2050000 read pairs finished. 450 secs passed Thread #0: 2100000 read pairs finished. 453 secs passed Thread #3: 2150000 read pairs finished. 455 secs passed Thread #2: 2200000 read pairs finished. 457 secs passed Thread #1: 2250000 read pairs finished. 488 secs passed Thread #0: 2300000 read pairs finished. 491 secs passed Thread #3: 2350000 read pairs finished. 492 secs passed Thread #2: 2400000 read pairs finished. 493 secs passed Thread #1: 2450000 read pairs finished. 525 secs passed Thread #0: 2500000 read pairs finished. 529 secs passed Thread #3: 2550000 read pairs finished. 531 secs passed Thread #2: 2600000 read pairs finished. 532 secs passed Thread #1: 2650000 read pairs finished. 565 secs passed Thread #0: 2700000 read pairs finished. 568 secs passed Thread #3: 2750000 read pairs finished. 570 secs passed Thread #2: 2800000 read pairs finished. 572 secs passed Thread #1: 2850000 read pairs finished. 601 secs passed Thread #0: 2900000 read pairs finished. 604 secs passed Thread #3: 2950000 read pairs finished. 605 secs passed Thread #2: 3000000 read pairs finished. 607 secs passed Thread #1: 3050000 read pairs finished. 638 secs passed Thread #0: 3100000 read pairs finished. 641 secs passed Thread #3: 3150000 read pairs finished. 643 secs passed Thread #2: 3200000 read pairs finished. 645 secs passed Thread #1: 3250000 read pairs finished. 673 secs passed Thread #0: 3300000 read pairs finished. 676 secs passed Thread #3: 3350000 read pairs finished. 679 secs passed Thread #2: 3400000 read pairs finished. 682 secs passed Thread #1: 3450000 read pairs finished. 709 secs passed Thread #0: 3500000 read pairs finished. 713 secs passed Thread #3: 3550000 read pairs finished. 716 secs passed Thread #2: 3600000 read pairs finished. 718 secs passed Thread #1: 3650000 read pairs finished. 745 secs passed Thread #0: 3700000 read pairs finished. 748 secs passed Thread #3: 3750000 read pairs finished. 752 secs passed Thread #2: 3800000 read pairs finished. 755 secs passed Thread #1: 3850000 read pairs finished. 781 secs passed Thread #0: 3900000 read pairs finished. 784 secs passed Thread #3: 3950000 read pairs finished. 788 secs passed Thread #2: 4000000 read pairs finished. 792 secs passed Thread #1: 4050000 read pairs finished. 817 secs passed Thread #0: 4100000 read pairs finished. 821 secs passed Thread #3: 4150000 read pairs finished. 825 secs passed Thread #2: 4200000 read pairs finished. 829 secs passed Thread #1: 4250000 read pairs finished. 854 secs passed Thread #0: 4300000 read pairs finished. 858 secs passed Thread #3: 4350000 read pairs finished. 863 secs passed Thread #2: 4400000 read pairs finished. 866 secs passed Thread #1: 4450000 read pairs finished. 891 secs passed Thread #0: 4500000 read pairs finished. 895 secs passed Thread #3: 4550000 read pairs finished. 899 secs passed Thread #2: 4600000 read pairs finished. 902 secs passed Thread #1: 4650000 read pairs finished. 928 secs passed Thread #0: 4700000 read pairs finished. 931 secs passed Thread #3: 4750000 read pairs finished. 936 secs passed Thread #2: 4800000 read pairs finished. 939 secs passed Thread #1: 4850000 read pairs finished. 965 secs passed Thread #0: 4900000 read pairs finished. 968 secs passed Thread #3: 4950000 read pairs finished. 973 secs passed Thread #2: 5000000 read pairs finished. 977 secs passed Thread #1: 5050000 read pairs finished. 1001 secs passed Thread #0: 5100000 read pairs finished. 1004 secs passed Thread #3: 5150000 read pairs finished. 1010 secs passed Thread #2: 5200000 read pairs finished. 1014 secs passed Thread #1: 5250000 read pairs finished. 1036 secs passed Thread #0: 5300000 read pairs finished. 1040 secs passed Thread #3: 5350000 read pairs finished. 1046 secs passed Thread #2: 5400000 read pairs finished. 1050 secs passed Thread #0: 5465974 read pairs finished. 1050 secs passed Thread #1: 5450000 read pairs finished. 1061 secs passed Total number of aligned reads: pairs: 2625402 (48%) single a: 1124227 (21%) single b: 885757 (16%) Done. Finished at Sun Mar 16 13:24:24 2014 Total time consumed: 1062 secs @ Sun Mar 16 13:24:25 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 13:25:16 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 13:30:53 2014: combining CpG methylation from both strands ... @ Sun Mar 16 13:31:31 2014: writing methratio_out.txt ... @ Sun Mar 16 13:44:03 2014: done. total 5964269 valid mappings, 39695564 covered cytosines, average coverage: 1.51 fold.
In [20]:
#file ID
fid="CgT1D5"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd="/Volumes/web/cnidarian/BiGo_larvae_merge/"
In [21]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [22]:
mkdir {fid}_{date}
In [23]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_[0316_1346]
In [24]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 13:46:58 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 22 secs passed total_kmers: 43046721 Create seed table. 68 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #2: 100000 read pairs finished. 112 secs passed Thread #1: 150000 read pairs finished. 113 secs passed Thread #3: 50000 read pairs finished. 114 secs passed Thread #0: 200000 read pairs finished. 114 secs passed Thread #2: 250000 read pairs finished. 156 secs passed Thread #1: 300000 read pairs finished. 157 secs passed Thread #3: 350000 read pairs finished. 158 secs passed Thread #0: 400000 read pairs finished. 159 secs passed Thread #2: 450000 read pairs finished. 201 secs passed Thread #0: 600000 read pairs finished. 206 secs passed Thread #3: 550000 read pairs finished. 207 secs passed Thread #1: 500000 read pairs finished. 209 secs passed Thread #2: 650000 read pairs finished. 246 secs passed Thread #0: 700000 read pairs finished. 251 secs passed Thread #3: 750000 read pairs finished. 252 secs passed Thread #1: 800000 read pairs finished. 255 secs passed Thread #2: 850000 read pairs finished. 291 secs passed Thread #0: 900000 read pairs finished. 297 secs passed Thread #3: 950000 read pairs finished. 298 secs passed Thread #1: 1000000 read pairs finished. 299 secs passed Thread #2: 1050000 read pairs finished. 336 secs passed Thread #0: 1100000 read pairs finished. 343 secs passed Thread #3: 1150000 read pairs finished. 344 secs passed Thread #1: 1200000 read pairs finished. 346 secs passed Thread #2: 1250000 read pairs finished. 381 secs passed Thread #0: 1300000 read pairs finished. 388 secs passed Thread #3: 1350000 read pairs finished. 390 secs passed Thread #1: 1400000 read pairs finished. 393 secs passed Thread #2: 1450000 read pairs finished. 426 secs passed Thread #0: 1500000 read pairs finished. 433 secs passed Thread #3: 1550000 read pairs finished. 435 secs passed Thread #1: 1600000 read pairs finished. 438 secs passed Thread #2: 1650000 read pairs finished. 470 secs passed Thread #0: 1700000 read pairs finished. 476 secs passed Thread #3: 1750000 read pairs finished. 478 secs passed Thread #1: 1800000 read pairs finished. 481 secs passed Thread #2: 1850000 read pairs finished. 514 secs passed Thread #3: 1950000 read pairs finished. 518 secs passed Thread #0: 1900000 read pairs finished. 519 secs passed Thread #1: 2000000 read pairs finished. 521 secs passed Thread #2: 2050000 read pairs finished. 553 secs passed Thread #3: 2100000 read pairs finished. 558 secs passed Thread #0: 2150000 read pairs finished. 560 secs passed Thread #1: 2200000 read pairs finished. 561 secs passed Thread #2: 2250000 read pairs finished. 595 secs passed Thread #3: 2300000 read pairs finished. 600 secs passed Thread #0: 2350000 read pairs finished. 601 secs passed Thread #1: 2400000 read pairs finished. 603 secs passed Thread #2: 2450000 read pairs finished. 636 secs passed Thread #3: 2500000 read pairs finished. 643 secs passed Thread #0: 2550000 read pairs finished. 644 secs passed Thread #1: 2600000 read pairs finished. 645 secs passed Thread #2: 2650000 read pairs finished. 678 secs passed Thread #3: 2700000 read pairs finished. 684 secs passed Thread #0: 2750000 read pairs finished. 686 secs passed Thread #1: 2800000 read pairs finished. 688 secs passed Thread #2: 2850000 read pairs finished. 719 secs passed Thread #3: 2900000 read pairs finished. 726 secs passed Thread #0: 2950000 read pairs finished. 727 secs passed Thread #1: 3000000 read pairs finished. 730 secs passed Thread #2: 3050000 read pairs finished. 761 secs passed Thread #3: 3100000 read pairs finished. 768 secs passed Thread #0: 3150000 read pairs finished. 769 secs passed Thread #1: 3200000 read pairs finished. 772 secs passed Thread #2: 3250000 read pairs finished. 804 secs passed Thread #3: 3300000 read pairs finished. 810 secs passed Thread #0: 3350000 read pairs finished. 811 secs passed Thread #1: 3400000 read pairs finished. 814 secs passed Thread #2: 3450000 read pairs finished. 848 secs passed Thread #3: 3500000 read pairs finished. 854 secs passed Thread #0: 3550000 read pairs finished. 856 secs passed Thread #1: 3600000 read pairs finished. 858 secs passed Thread #2: 3650000 read pairs finished. 893 secs passed Thread #3: 3700000 read pairs finished. 899 secs passed Thread #0: 3750000 read pairs finished. 900 secs passed Thread #1: 3800000 read pairs finished. 903 secs passed Thread #2: 3850000 read pairs finished. 939 secs passed Thread #3: 3900000 read pairs finished. 945 secs passed Thread #0: 3950000 read pairs finished. 947 secs passed Thread #1: 4000000 read pairs finished. 949 secs passed Thread #2: 4050000 read pairs finished. 985 secs passed Thread #3: 4100000 read pairs finished. 991 secs passed Thread #0: 4150000 read pairs finished. 992 secs passed Thread #1: 4200000 read pairs finished. 995 secs passed Thread #2: 4250000 read pairs finished. 1028 secs passed Thread #3: 4300000 read pairs finished. 1035 secs passed Thread #0: 4350000 read pairs finished. 1037 secs passed Thread #1: 4400000 read pairs finished. 1040 secs passed Thread #2: 4450000 read pairs finished. 1073 secs passed Thread #3: 4500000 read pairs finished. 1080 secs passed Thread #0: 4550000 read pairs finished. 1081 secs passed Thread #1: 4600000 read pairs finished. 1085 secs passed Thread #2: 4650000 read pairs finished. 1119 secs passed Thread #3: 4700000 read pairs finished. 1127 secs passed Thread #0: 4750000 read pairs finished. 1128 secs passed Thread #1: 4800000 read pairs finished. 1131 secs passed Thread #2: 4850000 read pairs finished. 1160 secs passed Thread #3: 4900000 read pairs finished. 1167 secs passed Thread #0: 4950000 read pairs finished. 1168 secs passed Thread #1: 5000000 read pairs finished. 1171 secs passed Thread #2: 5050000 read pairs finished. 1197 secs passed Thread #3: 5100000 read pairs finished. 1204 secs passed Thread #0: 5150000 read pairs finished. 1206 secs passed Thread #1: 5200000 read pairs finished. 1207 secs passed Thread #2: 5250000 read pairs finished. 1234 secs passed Thread #3: 5300000 read pairs finished. 1241 secs passed Thread #0: 5350000 read pairs finished. 1242 secs passed Thread #1: 5400000 read pairs finished. 1245 secs passed Thread #2: 5450000 read pairs finished. 1270 secs passed Thread #3: 5500000 read pairs finished. 1278 secs passed Thread #0: 5550000 read pairs finished. 1279 secs passed Thread #1: 5600000 read pairs finished. 1282 secs passed Thread #2: 5650000 read pairs finished. 1307 secs passed Thread #3: 5700000 read pairs finished. 1314 secs passed Thread #0: 5750000 read pairs finished. 1316 secs passed Thread #1: 5800000 read pairs finished. 1319 secs passed Thread #2: 5850000 read pairs finished. 1345 secs passed Thread #3: 5900000 read pairs finished. 1352 secs passed Thread #0: 5950000 read pairs finished. 1354 secs passed Thread #1: 6000000 read pairs finished. 1357 secs passed Thread #2: 6050000 read pairs finished. 1383 secs passed Thread #3: 6100000 read pairs finished. 1390 secs passed Thread #0: 6150000 read pairs finished. 1391 secs passed Thread #1: 6200000 read pairs finished. 1395 secs passed Thread #0: 6327160 read pairs finished. 1408 secs passed Thread #2: 6250000 read pairs finished. 1413 secs passed Thread #3: 6300000 read pairs finished. 1417 secs passed Total number of aligned reads: pairs: 2608088 (41%) single a: 1736286 (27%) single b: 1343344 (21%) Done. Finished at Sun Mar 16 14:10:36 2014 Total time consumed: 1418 secs @ Sun Mar 16 14:10:37 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 14:11:29 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 14:18:41 2014: combining CpG methylation from both strands ... @ Sun Mar 16 14:19:18 2014: writing methratio_out.txt ... @ Sun Mar 16 14:33:02 2014: done. total 6801985 valid mappings, 45492773 covered cytosines, average coverage: 1.54 fold.
In [25]:
#file ID
fid="CgT3D3"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd="/Volumes/web/cnidarian/BiGo_larvae_merge/"
In [26]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [27]:
mkdir {fid}_{date}
In [28]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_[0316_1436]
In [29]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 14:36:38 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 22 secs passed total_kmers: 43046721 Create seed table. 63 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #2: 150000 read pairs finished. 100 secs passed Thread #3: 50000 read pairs finished. 100 secs passed Thread #1: 100000 read pairs finished. 101 secs passed Thread #0: 200000 read pairs finished. 101 secs passed Thread #2: 250000 read pairs finished. 133 secs passed Thread #3: 300000 read pairs finished. 134 secs passed Thread #1: 350000 read pairs finished. 136 secs passed Thread #0: 400000 read pairs finished. 137 secs passed Thread #2: 450000 read pairs finished. 176 secs passed Thread #3: 500000 read pairs finished. 179 secs passed Thread #1: 550000 read pairs finished. 181 secs passed Thread #0: 600000 read pairs finished. 182 secs passed Thread #2: 650000 read pairs finished. 221 secs passed Thread #3: 700000 read pairs finished. 224 secs passed Thread #1: 750000 read pairs finished. 227 secs passed Thread #0: 800000 read pairs finished. 229 secs passed Thread #2: 850000 read pairs finished. 259 secs passed Thread #3: 900000 read pairs finished. 261 secs passed Thread #1: 950000 read pairs finished. 263 secs passed Thread #0: 1000000 read pairs finished. 270 secs passed Thread #3: 1100000 read pairs finished. 298 secs passed Thread #2: 1050000 read pairs finished. 299 secs passed Thread #1: 1150000 read pairs finished. 299 secs passed Thread #0: 1200000 read pairs finished. 305 secs passed Thread #3: 1250000 read pairs finished. 332 secs passed Thread #2: 1300000 read pairs finished. 334 secs passed Thread #1: 1350000 read pairs finished. 335 secs passed Thread #0: 1400000 read pairs finished. 339 secs passed Thread #3: 1450000 read pairs finished. 367 secs passed Thread #2: 1500000 read pairs finished. 368 secs passed Thread #1: 1550000 read pairs finished. 370 secs passed Thread #0: 1600000 read pairs finished. 374 secs passed Thread #3: 1650000 read pairs finished. 401 secs passed Thread #2: 1700000 read pairs finished. 402 secs passed Thread #1: 1750000 read pairs finished. 405 secs passed Thread #0: 1800000 read pairs finished. 408 secs passed Thread #3: 1850000 read pairs finished. 435 secs passed Thread #2: 1900000 read pairs finished. 438 secs passed Thread #1: 1950000 read pairs finished. 440 secs passed Thread #0: 2000000 read pairs finished. 442 secs passed Thread #3: 2050000 read pairs finished. 469 secs passed Thread #2: 2100000 read pairs finished. 472 secs passed Thread #1: 2150000 read pairs finished. 474 secs passed Thread #0: 2200000 read pairs finished. 476 secs passed Thread #3: 2250000 read pairs finished. 503 secs passed Thread #2: 2300000 read pairs finished. 507 secs passed Thread #1: 2350000 read pairs finished. 510 secs passed Thread #0: 2400000 read pairs finished. 512 secs passed Thread #3: 2450000 read pairs finished. 539 secs passed Thread #2: 2500000 read pairs finished. 543 secs passed Thread #1: 2550000 read pairs finished. 545 secs passed Thread #0: 2600000 read pairs finished. 548 secs passed Thread #3: 2650000 read pairs finished. 573 secs passed Thread #2: 2700000 read pairs finished. 578 secs passed Thread #1: 2750000 read pairs finished. 581 secs passed Thread #0: 2800000 read pairs finished. 584 secs passed Thread #3: 2850000 read pairs finished. 609 secs passed Thread #2: 2900000 read pairs finished. 615 secs passed Thread #1: 2950000 read pairs finished. 617 secs passed Thread #0: 3000000 read pairs finished. 620 secs passed Thread #3: 3050000 read pairs finished. 643 secs passed Thread #2: 3100000 read pairs finished. 650 secs passed Thread #1: 3150000 read pairs finished. 652 secs passed Thread #0: 3200000 read pairs finished. 655 secs passed Thread #3: 3250000 read pairs finished. 678 secs passed Thread #2: 3300000 read pairs finished. 686 secs passed Thread #1: 3350000 read pairs finished. 688 secs passed Thread #0: 3400000 read pairs finished. 691 secs passed Thread #3: 3450000 read pairs finished. 713 secs passed Thread #2: 3500000 read pairs finished. 722 secs passed Thread #1: 3550000 read pairs finished. 725 secs passed Thread #0: 3600000 read pairs finished. 727 secs passed Thread #3: 3650000 read pairs finished. 749 secs passed Thread #2: 3700000 read pairs finished. 757 secs passed Thread #1: 3750000 read pairs finished. 760 secs passed Thread #0: 3800000 read pairs finished. 762 secs passed Thread #3: 3850000 read pairs finished. 785 secs passed Thread #2: 3900000 read pairs finished. 799 secs passed Thread #1: 3950000 read pairs finished. 804 secs passed Thread #0: 4000000 read pairs finished. 805 secs passed Thread #3: 4050000 read pairs finished. 826 secs passed Thread #2: 4100000 read pairs finished. 841 secs passed Thread #1: 4150000 read pairs finished. 845 secs passed Thread #0: 4200000 read pairs finished. 846 secs passed Thread #3: 4250000 read pairs finished. 865 secs passed Thread #2: 4300000 read pairs finished. 876 secs passed Thread #1: 4350000 read pairs finished. 882 secs passed Thread #0: 4400000 read pairs finished. 883 secs passed Thread #3: 4450000 read pairs finished. 900 secs passed Thread #2: 4500000 read pairs finished. 911 secs passed Thread #1: 4550000 read pairs finished. 918 secs passed Thread #0: 4600000 read pairs finished. 919 secs passed Thread #3: 4650000 read pairs finished. 934 secs passed Thread #2: 4700000 read pairs finished. 946 secs passed Thread #1: 4750000 read pairs finished. 953 secs passed Thread #0: 4800000 read pairs finished. 956 secs passed Thread #3: 4850000 read pairs finished. 970 secs passed Thread #2: 4900000 read pairs finished. 981 secs passed Thread #1: 4950000 read pairs finished. 988 secs passed Thread #0: 5000000 read pairs finished. 990 secs passed Thread #3: 5050000 read pairs finished. 1004 secs passed Thread #2: 5100000 read pairs finished. 1017 secs passed Thread #1: 5150000 read pairs finished. 1024 secs passed Thread #0: 5200000 read pairs finished. 1027 secs passed Thread #3: 5250000 read pairs finished. 1040 secs passed Thread #2: 5300000 read pairs finished. 1053 secs passed Thread #1: 5350000 read pairs finished. 1061 secs passed Thread #0: 5400000 read pairs finished. 1063 secs passed Thread #3: 5450000 read pairs finished. 1076 secs passed Thread #2: 5500000 read pairs finished. 1089 secs passed Thread #1: 5550000 read pairs finished. 1097 secs passed Thread #0: 5600000 read pairs finished. 1099 secs passed Thread #3: 5650000 read pairs finished. 1112 secs passed Thread #2: 5700000 read pairs finished. 1125 secs passed Thread #1: 5750000 read pairs finished. 1133 secs passed Thread #0: 5800000 read pairs finished. 1135 secs passed Thread #3: 5850000 read pairs finished. 1148 secs passed Thread #2: 5900000 read pairs finished. 1162 secs passed Thread #1: 5950000 read pairs finished. 1170 secs passed Thread #0: 6000000 read pairs finished. 1172 secs passed Thread #3: 6050000 read pairs finished. 1185 secs passed Thread #2: 6100000 read pairs finished. 1198 secs passed Thread #1: 6150000 read pairs finished. 1206 secs passed Thread #0: 6200000 read pairs finished. 1209 secs passed Thread #3: 6250000 read pairs finished. 1222 secs passed Thread #2: 6300000 read pairs finished. 1234 secs passed Thread #1: 6350000 read pairs finished. 1243 secs passed Thread #0: 6400000 read pairs finished. 1245 secs passed Thread #3: 6450000 read pairs finished. 1258 secs passed Thread #2: 6500000 read pairs finished. 1270 secs passed Thread #1: 6550000 read pairs finished. 1279 secs passed Thread #0: 6600000 read pairs finished. 1282 secs passed Thread #3: 6650000 read pairs finished. 1293 secs passed Thread #2: 6700000 read pairs finished. 1306 secs passed Thread #1: 6750000 read pairs finished. 1316 secs passed Thread #0: 6800000 read pairs finished. 1319 secs passed Thread #3: 6850000 read pairs finished. 1330 secs passed Thread #2: 6900000 read pairs finished. 1342 secs passed Thread #1: 6950000 read pairs finished. 1352 secs passed Thread #0: 7000000 read pairs finished. 1355 secs passed Thread #3: 7050000 read pairs finished. 1366 secs passed Thread #2: 7100000 read pairs finished. 1378 secs passed Thread #1: 7150000 read pairs finished. 1389 secs passed Thread #0: 7200000 read pairs finished. 1392 secs passed Thread #3: 7250000 read pairs finished. 1402 secs passed Thread #0: 7380649 read pairs finished. 1414 secs passed Thread #2: 7300000 read pairs finished. 1415 secs passed Thread #1: 7350000 read pairs finished. 1421 secs passed Total number of aligned reads: pairs: 3906464 (53%) single a: 1418844 (19%) single b: 1116765 (15%) Done. Finished at Sun Mar 16 15:00:20 2014 Total time consumed: 1422 secs @ Sun Mar 16 15:00:22 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 15:01:23 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 15:10:55 2014: read 10000000 lines @ Sun Mar 16 15:11:15 2014: combining CpG methylation from both strands ... @ Sun Mar 16 15:12:07 2014: writing methratio_out.txt ... @ Sun Mar 16 15:28:02 2014: done. total 8688793 valid mappings, 52876971 covered cytosines, average coverage: 1.67 fold.
In [30]:
#file ID
fid="CgT3D5"
#TIMESTAMP
date=!date +%m%d_%H%M
#working directory (parent)
wd="/Volumes/web/cnidarian/BiGo_larvae_merge/"
In [31]:
cd {wd}
/Volumes/web/cnidarian/BiGo_larvae_merge
In [32]:
mkdir {fid}_{date}
In [33]:
cd {fid}_{date}
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_[0316_1532]
In [34]:
#option - number of processes
!{bsmap}bsmap -a {wd}{fid}_R1a.fastq -b {wd}{fid}_R2a.fastq -d {genome} -o bsmap_out.sam -p 4
!python {bsmap}methratio.py -d {genome} -u -z -g -o methratio_out.txt -s {bsmap}samtools bsmap_out.sam
#command for only obtaining the context '__CG_'
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> methratio_out_CG.txt
#5x coverage
!awk '{if ($8 >= 5) print $1,$2-1,$2+1,"CpG",$5}' <methratio_out_CG.txt> filt_methratio_out_CG.igv
!tr ' ' "\t" <filt_methratio_out_CG.igv> filt_methratio_{fid}.igv
BSMAP v2.74 Start at: Sun Mar 16 15:32:31 2014 Input reference file: /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa (format: FASTA) Load in 7658 db seqs, total size 557717710 bp. 22 secs passed total_kmers: 43046721 Create seed table. 74 secs passed max number of mismatches: read_length * 8% max gap size: 0 kmer cut-off ratio: 5e-07 max multi-hits: 100 max Ns: 5 seed size: 16 index interval: 4 quality cutoff: 0 base quality char: '!' min fragment size:28 max fragemt size:500 start from read #1 end at read #4294967295 additional alignment: T in reads => C in reference mapping strand (read_1): ++,-+ mapping strand (read_2): +-,-- Pair-end alignment(4 threads) Input read file #1: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_R1a.fastq (format: FASTQ) Input read file #2: /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_R2a.fastq (format: FASTQ) Output file: bsmap_out.sam (format: SAM) Thread #2: 150000 read pairs finished. 130 secs passed Thread #0: 100000 read pairs finished. 131 secs passed Thread #1: 50000 read pairs finished. 132 secs passed Thread #3: 200000 read pairs finished. 132 secs passed Thread #2: 250000 read pairs finished. 181 secs passed Thread #0: 300000 read pairs finished. 184 secs passed Thread #1: 350000 read pairs finished. 185 secs passed Thread #3: 400000 read pairs finished. 187 secs passed Thread #2: 450000 read pairs finished. 230 secs passed Thread #0: 500000 read pairs finished. 233 secs passed Thread #1: 550000 read pairs finished. 236 secs passed Thread #3: 600000 read pairs finished. 236 secs passed Thread #2: 650000 read pairs finished. 285 secs passed Thread #0: 700000 read pairs finished. 289 secs passed Thread #1: 750000 read pairs finished. 294 secs passed Thread #3: 800000 read pairs finished. 300 secs passed Thread #2: 850000 read pairs finished. 346 secs passed Thread #0: 900000 read pairs finished. 346 secs passed Thread #1: 950000 read pairs finished. 351 secs passed Thread #3: 1000000 read pairs finished. 358 secs passed Thread #2: 1050000 read pairs finished. 401 secs passed Thread #0: 1100000 read pairs finished. 403 secs passed Thread #1: 1150000 read pairs finished. 406 secs passed Thread #3: 1200000 read pairs finished. 412 secs passed Thread #2: 1250000 read pairs finished. 456 secs passed Thread #0: 1300000 read pairs finished. 458 secs passed Thread #1: 1350000 read pairs finished. 462 secs passed Thread #3: 1400000 read pairs finished. 467 secs passed Thread #2: 1450000 read pairs finished. 509 secs passed Thread #0: 1500000 read pairs finished. 511 secs passed Thread #1: 1550000 read pairs finished. 514 secs passed Thread #3: 1600000 read pairs finished. 519 secs passed Thread #2: 1650000 read pairs finished. 563 secs passed Thread #0: 1700000 read pairs finished. 566 secs passed Thread #1: 1750000 read pairs finished. 569 secs passed Thread #3: 1800000 read pairs finished. 576 secs passed Thread #2: 1850000 read pairs finished. 619 secs passed Thread #0: 1900000 read pairs finished. 623 secs passed Thread #1: 1950000 read pairs finished. 625 secs passed Thread #3: 2000000 read pairs finished. 631 secs passed Thread #2: 2050000 read pairs finished. 676 secs passed Thread #0: 2100000 read pairs finished. 680 secs passed Thread #1: 2150000 read pairs finished. 683 secs passed Thread #3: 2200000 read pairs finished. 691 secs passed Thread #2: 2250000 read pairs finished. 733 secs passed Thread #0: 2300000 read pairs finished. 737 secs passed Thread #1: 2350000 read pairs finished. 739 secs passed Thread #3: 2400000 read pairs finished. 747 secs passed Thread #2: 2450000 read pairs finished. 788 secs passed Thread #0: 2500000 read pairs finished. 792 secs passed Thread #1: 2550000 read pairs finished. 795 secs passed Thread #3: 2600000 read pairs finished. 800 secs passed Thread #2: 2650000 read pairs finished. 838 secs passed Thread #0: 2700000 read pairs finished. 844 secs passed Thread #1: 2750000 read pairs finished. 846 secs passed Thread #3: 2800000 read pairs finished. 853 secs passed Thread #2: 2850000 read pairs finished. 891 secs passed Thread #0: 2900000 read pairs finished. 896 secs passed Thread #1: 2950000 read pairs finished. 903 secs passed Thread #3: 3000000 read pairs finished. 913 secs passed Thread #2: 3050000 read pairs finished. 951 secs passed Thread #0: 3100000 read pairs finished. 954 secs passed Thread #1: 3150000 read pairs finished. 961 secs passed Thread #3: 3200000 read pairs finished. 971 secs passed Thread #2: 3250000 read pairs finished. 1009 secs passed Thread #0: 3300000 read pairs finished. 1013 secs passed Thread #1: 3350000 read pairs finished. 1018 secs passed Thread #3: 3400000 read pairs finished. 1028 secs passed Thread #2: 3450000 read pairs finished. 1067 secs passed Thread #0: 3500000 read pairs finished. 1072 secs passed Thread #1: 3550000 read pairs finished. 1076 secs passed Thread #3: 3600000 read pairs finished. 1085 secs passed Thread #2: 3650000 read pairs finished. 1121 secs passed Thread #0: 3700000 read pairs finished. 1127 secs passed Thread #1: 3750000 read pairs finished. 1130 secs passed Thread #3: 3800000 read pairs finished. 1139 secs passed Thread #2: 3850000 read pairs finished. 1177 secs passed Thread #0: 3900000 read pairs finished. 1184 secs passed Thread #1: 3950000 read pairs finished. 1186 secs passed Thread #3: 4000000 read pairs finished. 1195 secs passed Thread #2: 4050000 read pairs finished. 1231 secs passed Thread #0: 4100000 read pairs finished. 1239 secs passed Thread #1: 4150000 read pairs finished. 1242 secs passed Thread #3: 4200000 read pairs finished. 1250 secs passed Thread #2: 4250000 read pairs finished. 1281 secs passed Thread #0: 4300000 read pairs finished. 1289 secs passed Thread #1: 4350000 read pairs finished. 1292 secs passed Thread #3: 4400000 read pairs finished. 1298 secs passed Thread #2: 4450000 read pairs finished. 1335 secs passed Thread #0: 4500000 read pairs finished. 1344 secs passed Thread #1: 4550000 read pairs finished. 1347 secs passed Thread #3: 4600000 read pairs finished. 1358 secs passed Thread #2: 4650000 read pairs finished. 1406 secs passed Thread #0: 4700000 read pairs finished. 1417 secs passed Thread #1: 4750000 read pairs finished. 1420 secs passed Thread #3: 4800000 read pairs finished. 1431 secs passed Thread #2: 4850000 read pairs finished. 1479 secs passed Thread #0: 4900000 read pairs finished. 1491 secs passed Thread #1: 4950000 read pairs finished. 1494 secs passed Thread #3: 5000000 read pairs finished. 1505 secs passed Thread #2: 5050000 read pairs finished. 1549 secs passed Thread #0: 5100000 read pairs finished. 1561 secs passed Thread #1: 5150000 read pairs finished. 1564 secs passed Thread #3: 5200000 read pairs finished. 1574 secs passed Thread #2: 5250000 read pairs finished. 1620 secs passed Thread #0: 5300000 read pairs finished. 1632 secs passed Thread #1: 5350000 read pairs finished. 1634 secs passed Thread #3: 5400000 read pairs finished. 1646 secs passed Thread #2: 5450000 read pairs finished. 1685 secs passed Thread #0: 5500000 read pairs finished. 1697 secs passed Thread #1: 5550000 read pairs finished. 1699 secs passed Thread #3: 5600000 read pairs finished. 1709 secs passed Thread #2: 5650000 read pairs finished. 1751 secs passed Thread #0: 5700000 read pairs finished. 1769 secs passed Thread #1: 5750000 read pairs finished. 1771 secs passed Thread #3: 5800000 read pairs finished. 1779 secs passed Thread #2: 5850000 read pairs finished. 1820 secs passed Thread #0: 5900000 read pairs finished. 1838 secs passed Thread #1: 5950000 read pairs finished. 1840 secs passed Thread #3: 6000000 read pairs finished. 1848 secs passed Thread #2: 6050000 read pairs finished. 1886 secs passed Thread #0: 6100000 read pairs finished. 1904 secs passed Thread #1: 6150000 read pairs finished. 1906 secs passed Thread #3: 6200000 read pairs finished. 1911 secs passed Thread #2: 6250000 read pairs finished. 1941 secs passed Thread #0: 6300000 read pairs finished. 1957 secs passed Thread #1: 6350000 read pairs finished. 1959 secs passed Thread #3: 6400000 read pairs finished. 1964 secs passed Thread #2: 6450000 read pairs finished. 1993 secs passed Thread #0: 6500000 read pairs finished. 2013 secs passed Thread #1: 6550000 read pairs finished. 2015 secs passed Thread #3: 6600000 read pairs finished. 2020 secs passed Thread #2: 6650000 read pairs finished. 2047 secs passed Thread #0: 6700000 read pairs finished. 2066 secs passed Thread #1: 6750000 read pairs finished. 2068 secs passed Thread #3: 6800000 read pairs finished. 2076 secs passed Thread #2: 6850000 read pairs finished. 2096 secs passed Thread #0: 6900000 read pairs finished. 2113 secs passed Thread #1: 6950000 read pairs finished. 2114 secs passed Thread #3: 7000000 read pairs finished. 2120 secs passed Thread #2: 7050000 read pairs finished. 2136 secs passed Thread #0: 7100000 read pairs finished. 2149 secs passed Thread #1: 7150000 read pairs finished. 2151 secs passed Thread #3: 7200000 read pairs finished. 2155 secs passed Thread #3: 7350645 read pairs finished. 2156 secs passed Thread #2: 7250000 read pairs finished. 2169 secs passed Thread #0: 7300000 read pairs finished. 2177 secs passed Thread #1: 7350000 read pairs finished. 2178 secs passed Total number of aligned reads: pairs: 3946673 (54%) single a: 1329262 (18%) single b: 1076148 (15%) Done. Finished at Sun Mar 16 16:08:49 2014 Total time consumed: 2178 secs @ Sun Mar 16 16:08:50 2014: reading reference /Volumes/web/whale/ensembl/ftp.ensemblgenomes.org/pub/release-21/metazoa/fasta/crassostrea_gigas/dna/Crassostrea_gigas.GCA_000297895.1.21.dna_sm.genome.fa ... @ Sun Mar 16 16:09:51 2014: reading bsmap_out.sam ... [samopen] SAM header is present: 7658 sequences. @ Sun Mar 16 16:19:00 2014: read 10000000 lines @ Sun Mar 16 16:19:17 2014: combining CpG methylation from both strands ... @ Sun Mar 16 16:19:59 2014: writing methratio_out.txt ... @ Sun Mar 16 16:35:26 2014: done. total 8650035 valid mappings, 52138971 covered cytosines, average coverage: 1.70 fold.
In [35]:
!head filt_methratio_{fid}.igv
C13220 109 111 CpG 0.000 C13220 163 165 CpG 0.000 C13490 82 84 CpG 0.000 C13546 83 85 CpG 0.000 C14220 95 97 CpG 0.000 C14450 59 61 CpG 0.000 C14450 93 95 CpG 0.000 C14450 105 107 CpG 0.000 C14450 117 119 CpG 0.000 C14450 186 188 CpG 0.000
Comparing two sperm samples¶
In [1]:
!head /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1.igv
C12768 102 104 CpG 0.167 C12806 141 143 CpG 0.333 C12924 29 31 CpG 0.000 C12924 37 39 CpG 0.000 C12924 51 53 CpG 0.000 C12924 59 61 CpG 0.000 C12924 126 128 CpG 0.000 C12924 135 137 CpG 0.000 C13128 86 88 CpG 0.000 C13208 82 84 CpG 0.400
In [10]:
!awk '{print ($1"_"$2),$1,$2,$3,$4,$5}' </Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1.igv> \
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1_id.tab
In [11]:
!head /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1_id.tab
C12768_102 C12768 102 104 CpG 0.167 C12806_141 C12806 141 143 CpG 0.333 C12924_29 C12924 29 31 CpG 0.000 C12924_37 C12924 37 39 CpG 0.000 C12924_51 C12924 51 53 CpG 0.000 C12924_59 C12924 59 61 CpG 0.000 C12924_126 C12924 126 128 CpG 0.000 C12924_135 C12924 135 137 CpG 0.000 C13128_86 C13128 86 88 CpG 0.000 C13208_82 C13208 82 84 CpG 0.400
In [ ]:
!python {spd}singleupload.py -d filt_methratio_CgM1 /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1.igv
In [2]:
!head /Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]/filt_methratio_CgM3.igv
C13128 86 88 CpG 0.000 C13208 82 84 CpG 0.429 C13442 64 66 CpG 0.000 C13442 78 80 CpG 0.000 C13612 88 90 CpG 0.000 C13992 53 55 CpG 0.000 C14180 157 159 CpG 0.000 C14180 192 194 CpG 0.000 C14220 95 97 CpG 0.000 C14220 142 144 CpG 0.000
Try SQLShare¶
In [3]:
#Location of SQLShare python tools: you can empty ("") if tools are in PATH
spd="/Users/sr320/sqlshare-pythonclient/tools/"
In [4]:
!python {spd}singleupload.py -d filt_methratio_CgM1 /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1.igv
processing chunk line 0 to 380223 (18.4622421265 s elapsed) pushing /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/filt_methratio_CgM1.igv... parsing D588E61B... finished filt_methratio_CgM1
In [5]:
!python {spd}singleupload.py -d filt_methratio_CgM3 /Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]/filt_methratio_CgM3.igv
processing chunk line 0 to 404025 (15.2012460232 s elapsed) pushing /Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]/filt_methratio_CgM3.igv... parsing A7780515... finished filt_methratio_CgM3
Coverting files for Methylkit¶
In [ ]:
target
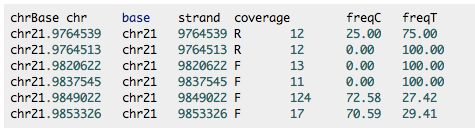
In [7]:
cat /Volumes/web/Mollusk/bs_larvae_exp/methratio.awk.sh
#!/usr/bin/awk -f
BEGIN{ print "chr.Base\tchr\tbase\tstrand\tcoverage\tfreqC\tfreqT" }
{
if ($3 == "+") {
strand="F"
} else {
strand="R"
}
FC=$7/$8
FT=1-FC
chrbase=$1"."$2
printf "%s\t%s\t%s\t%s\t%d\t%.2f\t%.2f\n",
chrbase, $1, $2, strand, $8, FC, FT
}
In [9]:
!head /Volumes/web/cnidarian/CgM1_[0316_0954]mk.txt
chr.Base chr base strand coverage freqC freqT C12706.69 C12706 69 F 1 0.00 1.00 C12706.71 C12706 71 F 1 0.00 1.00 C12722.104 C12722 104 F 2 0.50 0.50 C12722.134 C12722 134 F 2 0.50 0.50 C12768.103 C12768 103 F 6 0.17 0.83 C12768.119 C12768 119 F 4 0.25 0.75 C12768.145 C12768 145 F 3 0.00 1.00 C12768.161 C12768 161 F 1 0.00 1.00 C12778.114 C12778 114 F 1 0.00 1.00
List of methratio files¶
In [11]:
!head -2 /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/methratio_out.txt
chr pos strand context ratio eff_CT_count C_count CT_count rev_G_count rev_GA_count CI_lower CI_upper C12706 51 - CAGAG 0.000 1.00 0 1 0 0 0.000 0.793
In [26]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]
In [2]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [34]:
!awk '{if ($8 >= 5) print $1,$2,$2+1,$3,$8,($7/$8),(1-($7/$8))}' <methratio_out.txt> mk_methratio_out_CG5x.txt
awk: division by zero input record number 1, file source line number 1
In [33]:
!head mk_methratio_out_CG5x.txt
C12706 69 + TCCGC 0.000 1.00 0 1 1 1 0.000 0.793 C12706 71 + CGCGT 0.000 1.00 0 1 1 1 0.000 0.793 C12722 104 + TACGT 0.500 2.00 1 2 1 1 0.095 0.905 C12722 134 + TGCGG 0.500 2.00 1 2 2 2 0.095 0.905 C12768 103 + TTCGT 0.167 6.00 1 6 6 6 0.030 0.564 C12768 119 + TTCGT 0.250 4.00 1 4 4 4 0.046 0.699 C12768 145 + TTCGT 0.000 3.00 0 3 3 3 0.000 0.562 C12768 161 + TTCGT 0.000 1.00 0 1 1 1 0.000 0.793 C12778 114 + AGCGG 0.000 1.00 0 1 1 1 0.000 0.793 C12782 75 + CGCGT 0.000 1.00 0 1 0 0 0.000 0.793
In [5]:
cd
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]
In [6]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [7]:
!awk '$8 >= 5' <methratio_out_CG.txt> mk_methratio_out_CG5x.txt
In [8]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_[0316_1306]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_[0316_1306]
In [9]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [10]:
!awk '$8 >= 5' <methratio_out_CG.txt> mk_methratio_out_CG5x.txt
In [11]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_[0316_1346]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_[0316_1346]
In [12]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [13]:
!awk '$8 >= 5' <methratio_out_CG.txt> mk_methratio_out_CG5x.txt
In [14]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_[0316_1436]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_[0316_1436]
In [15]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [16]:
!awk '$8 >= 5' <methratio_out_CG.txt> mk_methratio_out_CG5x.txt
In [17]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_[0316_1532]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_[0316_1532]
In [18]:
!grep "[A-Z][A-Z]CG[A-Z]" <methratio_out.txt> mk_methratio_out_CG.txt
In [19]:
!awk '$8 >= 5' <methratio_out_CG.txt> mk_methratio_out_CG5x.txt
In [20]:
!head mk_methratio_out_CG5x.txt
C12722 104 + TACGT 1.000 1.00 1 1 1 1 0.207 1.000 C12722 134 + TGCGG 1.000 1.00 1 1 1 1 0.207 1.000 C12768 103 + TTCGT 1.000 1.00 1 1 1 1 0.207 1.000 C12768 119 + TTCGT 1.000 1.00 1 1 1 1 0.207 1.000 C12778 34 + CTCGC 0.000 1.00 0 1 1 1 0.000 0.793 C12778 52 + TTCGA 0.000 2.00 0 2 2 2 0.000 0.658 C12778 114 + AGCGG 0.000 1.00 0 1 1 1 0.000 0.793 C12780 100 + GACGT 0.000 1.00 0 1 1 1 0.000 0.793 C12780 144 + ATCGA 0.000 2.00 0 2 2 2 0.000 0.658 C12788 86 + CTCGA 0.000 1.00 0 1 1 1 0.000 0.793
methylkit it¶
In [ ]:
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM3_[0316_1207]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D3_[0316_1306]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT1D5_[0316_1346]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D3_[0316_1436]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgT3D5_[0316_1532]/
In [21]:
cd /Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]/
/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]
In [22]:
!/Volumes/web/Mollusk/bs_larvae_exp/methratio.awk.sh \
mk_methratio_out_CG5x.txt > mk_CgM1.txt
In [25]:
pwd
Out[25]:
u'/Volumes/web/cnidarian/BiGo_larvae_merge/CgM1_[0316_0954]'
In [24]:
!head mk_methratio_out_CG5x.txt
C12706 69 + TCCGC 0.000 1.00 0 1 1 1 0.000 0.793 C12706 71 + CGCGT 0.000 1.00 0 1 1 1 0.000 0.793 C12722 104 + TACGT 0.500 2.00 1 2 1 1 0.095 0.905 C12722 134 + TGCGG 0.500 2.00 1 2 2 2 0.095 0.905 C12768 103 + TTCGT 0.167 6.00 1 6 6 6 0.030 0.564 C12768 119 + TTCGT 0.250 4.00 1 4 4 4 0.046 0.699 C12768 145 + TTCGT 0.000 3.00 0 3 3 3 0.000 0.562 C12768 161 + TTCGT 0.000 1.00 0 1 1 1 0.000 0.793 C12778 114 + AGCGG 0.000 1.00 0 1 1 1 0.000 0.793 C12782 75 + CGCGT 0.000 1.00 0 1 0 0 0.000 0.793
In [ ]: