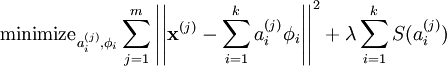Data Driven Modeling¶
### PhD seminar series at Chair for Computer Aided Architectural Design (CAAD), ETH Zurich
Topics to be discussed¶
- Self Organizing Maps
In [1]:
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
pd.__version__
import sys
from scipy import stats
%matplotlib inline
In [2]:
# A two dimensional example
fig = plt.figure()
N = 100
d0 = 1.6*np.random.randn(N,2)
d0[:,0]= d0[:,0] - 3
plt.plot(d0[:,0],d0[:,1],'.b')
d1 = 1.6*np.random.randn(N,2)+7.6
plt.plot(d1[:,0],d1[:,1],'.b')
d2 = 1.6*np.random.randn(N,2)
d2[:,0]= d2[:,0] + 5
d2[:,1]= d2[:,1] + 1
plt.plot(d2[:,0],d2[:,1],'.b')
d3 = 1.6*np.random.randn(N,2)
d3[:,0]= d3[:,0] - 5
d3[:,1]= d3[:,1] + 4
plt.plot(d3[:,0],d3[:,1],'.b')
d4 = 1.8*np.random.randn(N,2)
d4[:,0]= d4[:,0] - 15
d4[:,1]= d4[:,1] + 14
plt.plot(d4[:,0],d4[:,1],'.b')
d5 = 1.8*np.random.randn(N,2)
d5[:,0]= d5[:,0] + 25
d5[:,1]= d5[:,1] + 14
plt.plot(d5[:,0],d5[:,1],'.b')
Data1 = np.concatenate((d0,d1,d2,d3,d4,d5))
fig.set_size_inches(7,7)

The Original Algorithm¶
Online learning¶

0- Assume we have a grid (e.g. 2D), where each node has an n-dimensional vector w, and a 2-D index, (x,y)¶
1- initialize ws with random vectors¶
2- for several runs repeat the following steps¶
- ** project each data point to the grid and find the most similar node (using the selected similarity measure) to that data point. This node is called Best Matching Unit (BMU)**
- ** considering the BMU as the center, BMU and its neighboring nodes adapt their Ws to be similar to the data vector based on a selected learning rate**
In [14]:
def normalize_by(data_raw, data, method='var'):
me = np.mean(data_raw, axis = 0)
st = np.std(data_raw, axis = 0)
if method == 'var':
n_data = (data-me)/st
return n_data
def denormalize_by(data_by, n_vect, n_method = 'var'):
#based on the normalization
if n_method == 'var':
me = np.mean(data_by, axis = 0)
st = np.std(data_by, axis = 0)
vect = n_vect* st + me
return vect
else:
print 'data is not normalized before'
return n_vect
def random_initialize(mapszie,data):
nnodes = mapsize[0]*mapsize[1]
dim = data.shape[1]
#It produces random values in the range of min- max of each dimension based on a uniform distribution
mn = np.tile(np.min(data, axis =0)-1.5, (nnodes,1))
mx = np.tile(np.max(data, axis =0)+1.5, (nnodes,1))
init_W = mn + (mx-mn)*(np.random.rand(nnodes,dim))
return init_W
def rect_dist(mapsize,bmu):
#the way we consider the list of nodes in a planar grid is that node0 is on top left corner,
#nodemapsz[1]-1 is top right corner and then it goes to the second row.
#no. of rows is map_size[0] and no. of cols is map_size[1]
rows = mapsize[0]
cols = mapsize[1]
#bmu should be an integer between 0 to no_nodes
if 0<=bmu<=(rows*cols):
c_bmu = int(bmu%cols)
r_bmu = int(bmu/cols)
else:
print 'wrong bmu'
#calculating the grid distance
if np.logical_and(rows>0 , cols>0):
r,c = np.arange(0, rows, 1)[:,np.newaxis] , np.arange(0,cols, 1)
dist2 = (r-r_bmu)**2 + (c-c_bmu)**2
return dist2.ravel()
else:
print 'please consider the above mentioned errors'
return np.zeros((rows,cols)).ravel()
def calculate_UD2(mapsize):
nnodes = mapsize[0]*mapsize[1]
UD2 = np.zeros((nnodes, nnodes))
for i in range(nnodes):
UD2[i,:] = rect_dist(mapsize,i).reshape(1,nnodes)
return UD2
def calculate_neigh_activation(UD2,bmu,radius):
# 'Guassian'
return np.exp(-1.0*UD2[bmu]/(2.0*radius**2))
def find_bmu(data,W):
import scipy.spatial.distance as DIST
Dists = DIST.cdist(data,W)
ind_sets = np.argmin(Dists,axis=1)
return Dists,ind_sets
#Learning rate adaptation
def calc_alpha(t,alpha0,T,method='linear'):
if method =='linear':
return alpha0*(1-t/float(T))
if method =='power':
return alpha0*(.005/alpha0)**(float(t)/T)
if method =='inv':
return alpha0/(1+100*t/float(T))
In [57]:
def train_som_online(data,mapsize,trainlen):
# mapsize = [10,10]
dlen = len(data)
#normalize the data
data_n = normalize_by(data, data, method='var');
#Initialize the map
W = random_initialize(mapsize,data_n);
UD2 = calculate_UD2(mapsize);
alpha0 = .5
if min(mapsize)==1:
radiusin = max(mapsize)/float(2)
else:
radiusin = max(mapsize)/float(6)
radiusfin = max([1,radiusin/float(4)])
radiusfin = radiusin/float(10)
print radiusin,radiusfin
# trainlen= 200*nnodes/float(dlen)
# trainlen = int(np.ceil(trainlen))
radius = np.linspace(radiusin, radiusfin, trainlen)
qe = []
Ws = []
W_denorm = denormalize_by(data, W.copy(), n_method = 'var')
Ws.append(W_denorm.copy())
for i in range(trainlen):
#all or a sample
data_e = np.random.permutation(data_n)[:]
qe_t = np.zeros((data_e.shape[0],1))
for j, d in enumerate(data_e):
#project using a distance function
#find bmu
#Competition between nodes
Dists,bmu = find_bmu(d[np.newaxis,:],W)
qe_t[j] = np.min(Dists)
# Adapt nodes
#distance to bmu in the lower dimensional space
neiboring_effect = calculate_neigh_activation(UD2,bmu,radius[i])
#learning rate
alpha = calc_alpha(i,alpha0,trainlen,method='inv')
#Adapatation
W[:] = W[:] + neiboring_effect.T * alpha * (d-W[:])
W_denorm = denormalize_by(data, W.copy(), n_method = 'var')
Ws.append(W_denorm.copy())
if i%10==0:
print 'iteration {}: alpha: {}, raduis: {} and quantization error is {}'.format(i,alpha,radius[i],np.mean(qe_t))
qe.append(np.mean(qe_t))
Ws = np.asarray(Ws)
return W, qe,Ws
In [434]:
data = Data1
Data = np.random.randint(0,2,size = (500,3))
data = Data1
In [435]:
data = Data1
mapsize = [40,40]
trainlen = 200
W, qe,Ws = train_som_online(data,mapsize,trainlen)
codebook = denormalize_by(data, W, n_method = 'var')
6.66666666667 0.666666666667 iteration 0: alpha: 0.5, raduis: 6.66666666667 and quantization error is 0.118186899882 iteration 10: alpha: 0.0833333333333, raduis: 6.36515912898 and quantization error is 0.0978878664771 iteration 20: alpha: 0.0454545454545, raduis: 6.06365159129 and quantization error is 0.0906245838952 iteration 30: alpha: 0.03125, raduis: 5.7621440536 and quantization error is 0.08590481532 iteration 40: alpha: 0.0238095238095, raduis: 5.46063651591 and quantization error is 0.0820396115341 iteration 50: alpha: 0.0192307692308, raduis: 5.15912897822 and quantization error is 0.0748734499173 iteration 60: alpha: 0.0161290322581, raduis: 4.85762144054 and quantization error is 0.0700469994541 iteration 70: alpha: 0.0138888888889, raduis: 4.55611390285 and quantization error is 0.0647056462594 iteration 80: alpha: 0.0121951219512, raduis: 4.25460636516 and quantization error is 0.0611252921953 iteration 90: alpha: 0.0108695652174, raduis: 3.95309882747 and quantization error is 0.0556555269744 iteration 100: alpha: 0.00980392156863, raduis: 3.65159128978 and quantization error is 0.0517942198458 iteration 110: alpha: 0.00892857142857, raduis: 3.35008375209 and quantization error is 0.0480460081038 iteration 120: alpha: 0.00819672131148, raduis: 3.04857621441 and quantization error is 0.0442862072837 iteration 130: alpha: 0.00757575757576, raduis: 2.74706867672 and quantization error is 0.0414536197213 iteration 140: alpha: 0.00704225352113, raduis: 2.44556113903 and quantization error is 0.0387689534516 iteration 150: alpha: 0.00657894736842, raduis: 2.14405360134 and quantization error is 0.036193964836 iteration 160: alpha: 0.00617283950617, raduis: 1.84254606365 and quantization error is 0.0337516643049 iteration 170: alpha: 0.00581395348837, raduis: 1.54103852596 and quantization error is 0.0317958630539 iteration 180: alpha: 0.00549450549451, raduis: 1.23953098827 and quantization error is 0.0301403734082 iteration 190: alpha: 0.00520833333333, raduis: 0.938023450586 and quantization error is 0.0284984199562
In [436]:
plt.plot(qe)
Out[436]:
[<matplotlib.lines.Line2D at 0x147049050>]
In [437]:
# Double pendulum formula translated from the C code at
# http://www.physics.usyd.edu.au/~wheat/dpend_html/solve_dpend.c
from numpy import sin, cos
import numpy as np
import matplotlib.pyplot as plt
import scipy.integrate as integrate
import matplotlib.animation as animation
# data_n = normalize_by(data, data, method='var');
fig = plt.figure()
ax = fig.add_subplot(111, autoscale_on=False, xlim=(data.min(axis=0)[0],data.max(axis=0)[0]), ylim=(data.min(axis=0)[1],data.max(axis=0)[1]))
# ax.grid()
dataplt, = ax.plot([], [], '.k',markersize=3)
somplt, = ax.plot([], [], 'o',markersize=3,alpha=.5,markerfacecolor='None',markeredgecolor='green')
time_template = 'time = %.1fs'
time_text = ax.text(0.05, 0.9, '', transform=ax.transAxes)
def init():
dataplt.set_data([], [])
somplt.set_data([], [])
time_text.set_text('')
return dataplt,somplt, time_text
def animate(i):
dataplt.set_data(data[:,0],data[:,1])
somplt.set_data(Ws[i][:,0],Ws[i][:,1])
time_text.set_text(time_template % (i))
return dataplt,somplt, time_text
ani = animation.FuncAnimation(fig, animate, np.arange(0, trainlen),
interval=25, blit=True, init_func=init)
# ani.save('./Images/double_pendulum.mp4', fps=15)
ani.save('./Images/SOM.mp4', fps=15, extra_args=['-vcodec', 'libx264'],dpi=200)
plt.close()
In [1]:
from IPython.display import HTML
HTML("""
<video width="600" height="400" controls>
<source src="files/Images/SOM.mp4" type="video/mp4">
</video>
""")
Out[1]:
In [439]:
import sompylib.sompy as SOM
msz11 =40
msz10 = 40
X = Data1
som1 = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
som1.init_map()
# som1.train(trainlen=None,n_job = 1, shared_memory = 'no',verbose='final')
som1.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook1 = som1.codebook[:]
codebook1_n = SOM.denormalize_by(som1.data_raw, codebook1, n_method = 'var')
Total time elapsed: 9.754000 secodns final quantization error: 0.015409
In [440]:
fig = plt.figure()
plt.plot(data[:,0],data[:,1],'.')
plt.plot(codebook1_n[:,0],codebook1_n[:,1],'.r')
fig.set_size_inches(7,7)
Applications of SOM¶
- ** Dimensionality Reduction and Visualization of High Dimensional Spaces"**
- ** Feature selection**
- ** Multi-criteria Optimization**
- ** Clustering**
- ** Classification**
- ** Space transmorformation similar to PCA, K-means, Sparse Coding,...**
- ** Function approximation**
- ** Transfer function : Multidirectional function approximantion**
- ** Non parametric Probability density estimation**
- ** filling missing values**
- resampling
- ** Data Reduction and Abstraction similar to K-means with a large K**
In [441]:
som1.hit_map()
In [94]:
som1.view_map(text_size=8)
An example with high dimensional data¶
Feature extraction¶
In many real engineering applications, we are intersted to know what are the important factors in an experiment.¶
Imagine, each experiment takes few hours and considering the combinations of all values of different variable explodes the experimental space!!!¶
in the following case, we have 5 parameters and one label which says the label of the experiment¶
In [442]:
path = "./Data/CFD.csv"
D = pd.read_csv(path)
# D = D[['Rent','ZIP','Year built','Living space','lng','lat']]
D.head()
Out[442]:
| P1 | P2 | P3 | P4 | P5 | Label | |
|---|---|---|---|---|---|---|
| 0 | 11.00 | 7.50 | 5.75 | 0.001750 | 0.000500 | 1 |
| 1 | 5.75 | 11.00 | 9.25 | 0.001125 | 0.001750 | 1 |
| 2 | 9.25 | 9.25 | 11.00 | 0.001750 | 0.003000 | 1 |
| 3 | 9.25 | 11.00 | 11.00 | 0.001750 | 0.002375 | 1 |
| 4 | 11.00 | 11.00 | 11.00 | 0.000500 | 0.002375 | 1 |
If we use one to one plots and with colors show the labels, usually we don't get any thing¶
In [443]:
c = 1
fig = plt.figure(figsize=(20,20))
for i in range(0,4):
for j in range(i,5):
if i != j:
plt.subplot(5,4,c);
c = c +1;
plt.scatter(D.values[:,i],D.values[:,j],marker='o',s=70,edgecolors=None,alpha=1.,vmin=0,vmax=1,c=plt.cm.RdYlBu_r(D.Label.values[:].astype(float)));
plt.xlim((D.values[:,i].min(),D.values[:,i].max()));
plt.ylim((D.values[:,j].min(),D.values[:,j].max()));
plt.xlabel(D.columns[i]);
plt.ylabel(D.columns[j]);
if i == j:
plt.subplot(5,4,c);
c = c +1;
plt.scatter(D.values[:,i],D.values[:,j],marker='o',s=70,edgecolors=None,alpha=1.,vmin=0,vmax=1,c=plt.cm.RdYlBu_r(D.Label.values[:].astype(float)));
plt.xlim((D.values[:,i].min(),D.values[:,i].max()));
plt.ylim((D.values[:,j].min(),D.values[:,j].max()));
plt.xlabel(D.columns[i]);
plt.ylabel(D.columns[j]);
if we take the usual correlations¶
In [444]:
D.corr()
Out[444]:
| P1 | P2 | P3 | P4 | P5 | Label | |
|---|---|---|---|---|---|---|
| P1 | 1.000000 | -0.020883 | -0.089509 | -0.036360 | 0.031264 | 0.383061 |
| P2 | -0.020883 | 1.000000 | 0.082728 | -0.084491 | 0.042166 | 0.168023 |
| P3 | -0.089509 | 0.082728 | 1.000000 | -0.023677 | 0.012584 | 0.144121 |
| P4 | -0.036360 | -0.084491 | -0.023677 | 1.000000 | -0.118082 | -0.220599 |
| P5 | 0.031264 | 0.042166 | 0.012584 | -0.118082 | 1.000000 | -0.020029 |
| Label | 0.383061 | 0.168023 | 0.144121 | -0.220599 | -0.020029 | 1.000000 |
But now if we train a SOM with the parameters and visualize their labels¶
In [630]:
import sompylib1.sompy as SOM1
msz11 =70
msz10 = 70
X = D.values[:,:-1]
som2 = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
# som1 = SOM1.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
som2.init_map()
som2.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook1 = som2.codebook[:]
codebook1_n = SOM.denormalize_by(som2.data_raw, codebook1, n_method = 'var')
som2.compname= [D.columns[:-1]]
Total time elapsed: 231.507000 secodns final quantization error: 0.116649
In [ ]:
In [631]:
som2.view_map(text_size=8)
In [632]:
# We project data
som2.cluster_labels = np.asarray(["" for i in range(som2.nnodes)])
bmus = som2.project_data(X)
Df = pd.DataFrame(data=bmus,columns=['bmu'])
Df['dlabel'] = D['Label']
Df.head()
gb = Df.groupby(by='bmu').median()
sel_bmus = gb.index.values[:]
print sel_bmus.shape
sel_bmus_label = gb.dlabel.values[:]
som2.cluster_labels[sel_bmus]= sel_bmus_label
(725,)
In [633]:
def viewmap_withlabel(som,data=None):
if hasattr(som, 'cluster_labels'):
codebook = getattr(som, 'cluster_labels')
else:
print 'clustering based on default parameters...'
codebook = som.cluster()
msz = getattr(som, 'mapsize')
fig = plt.figure(figsize=(20,14))
dim = som.codebook.shape[1]
no_row_in_plot = dim/3 + 1 #6 is arbitrarily selected
if no_row_in_plot <=1:
no_col_in_plot = dim
else:
no_col_in_plot = 3
for k in range(dim):
ax = plt.subplot(no_row_in_plot,no_col_in_plot,k+1)
if data == None:
# data_tr = getattr(som, 'data_raw')
# proj = som.project_data(data_tr)
# coord = som.ind_to_xy(proj)
cents = som.ind_to_xy(np.arange(0,msz[0]*msz[1]))
for i, txt in enumerate(codebook):
ax.annotate(txt, (cents[i,1],cents[i,0]),size=8, va="center")
if data != None:
proj = som.project_data(data)
cents = som.ind_to_xy(proj)
label = codebook[proj]
for i, txt in enumerate(label):
ax.annotate(txt, (cents[i,1],cents[i,0]),size=8, va="center")
ax.set_yticklabels([])
ax.set_xticklabels([])
ax.imshow(som.codebook[:,k].reshape(msz[0],msz[1])[::],alpha=1,cmap=plt.cm.RdYlBu_r)
# ax.imshow(som.codebook[:,k].reshape(msz[0],msz[1])[::],alpha=1,cmap=plt.cm.Accent_r)
plt.subplots_adjust(hspace = .001,wspace=.01)
# plt.tight_layout()
In [634]:
viewmap_withlabel(som2)
In [647]:
import sompylib.sompy as SOM
msz11 =1
msz10 = 10
X = Data1
somS = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
somS.init_map()
# som1.train(trainlen=None,n_job = 1, shared_memory = 'no',verbose='final')
somS.train(n_job = 1, shared_memory = 'no',verbose='final')
codebookS = somS.codebook[:]
codebookS_n = SOM.denormalize_by(somS.data_raw, codebookS, n_method = 'var')
Total time elapsed: 0.014000 secodns final quantization error: 0.332617
In [648]:
fig = plt.figure()
plt.plot(X[:,0],X[:,1],'.')
plt.plot(codebookS_n[:,0],codebookS_n[:,1],'or')
fig.set_size_inches(7,7)
In [649]:
import sompylib.sompy as SOM
msz11 =10
msz10 = 10
X = Data1
somM = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
somM.init_map()
# som1.train(trainlen=None,n_job = 1, shared_memory = 'no',verbose='final')
somM.train(n_job = 1, shared_memory = 'no',verbose='final')
codebookM = somM.codebook[:]
codebookM_n = SOM.denormalize_by(somM.data_raw, codebookM, n_method = 'var')
Total time elapsed: 0.023000 secodns final quantization error: 0.099163
In [666]:
somM.hit_map_cluster_number();
In [667]:
K = 8
cl = somM.cluster(method='Kmeans', n_clusters=K)
fig = plt.figure()
plt.plot(X[:,0],X[:,1],'.',alpha=.4)
plt.scatter(codebookM_n[:,0],codebookM_n[:,1],s=60,marker='o',alpha=1.,c=plt.cm.RdYlBu_r(np.asarray(cl)/float(K)));
fig.set_size_inches(7,7);
In [668]:
import sompylib.sompy as SOM
msz11 =30
msz10 = 30
X = Data1
somL = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
somL.init_map()
# som1.train(trainlen=None,n_job = 1, shared_memory = 'no',verbose='final')
somL.train(n_job = 1, shared_memory = 'no',verbose='final')
codebookL = somL.codebook[:]
codebookL_n = SOM.denormalize_by(somL.data_raw, codebookL, n_method = 'var')
Total time elapsed: 1.589000 secodns final quantization error: 0.023740
In [672]:
somL.view_map(text_size=8)
In [673]:
U = somL.view_U_matrix(distance2=2, row_normalized='No', show_data='Yes', contooor='Yes', blob='No', save='NO', save_dir='')
In [265]:
dlen = 700
tetha = np.random.uniform(low=0,high=2*np.pi,size=dlen)[:,np.newaxis]
X1 = 3*np.cos(tetha)+ .22*np.random.rand(dlen,1)
Y1 = 3*np.sin(tetha)+ .22*np.random.rand(dlen,1)
D1 = np.concatenate((X1,Y1),axis=1)
X2 = 1*np.cos(tetha)+ .22*np.random.rand(dlen,1)
Y2 = 1*np.sin(tetha)+ .22*np.random.rand(dlen,1)
D2 = np.concatenate((X2,Y2),axis=1)
X3 = 5*np.cos(tetha)+ .22*np.random.rand(dlen,1)
Y3 = 5*np.sin(tetha)+ .22*np.random.rand(dlen,1)
D3 = np.concatenate((X3,Y3),axis=1)
X4 = 8*np.cos(tetha)+ .22*np.random.rand(dlen,1)
Y4 = 8*np.sin(tetha)+ .22*np.random.rand(dlen,1)
D4 = np.concatenate((X4,Y4),axis=1)
Data3 = np.concatenate((D1,D2,D3,D4),axis=0)
fig = plt.figure()
plt.plot(Data3[:,0],Data3[:,1],'ob',alpha=0.2, markersize=4)
fig.set_size_inches(7,7)
In [675]:
import sompylib.sompy as SOM
msz11 =30
msz10 = 30
X = Data3
somL = SOM.SOM('', X, mapsize = [msz10, msz11],norm_method = 'var',initmethod='pca')
somL.init_map()
# som1.train(trainlen=None,n_job = 1, shared_memory = 'no',verbose='final')
somL.train(n_job = 1, shared_memory = 'no',verbose='final')
codebookL = somL.codebook[:]
codebookL_n = SOM.denormalize_by(somL.data_raw, codebookL, n_method = 'var')
Total time elapsed: 0.804000 secodns final quantization error: 0.027553
In [676]:
U = somL.view_U_matrix(distance2=2, row_normalized='No', show_data='Yes', contooor='Yes', blob='No', save='NO', save_dir='')
Applications of SOM¶
- ** Classification**
- ** Function approximation**
- ** Transfer function : Multidirectional function approximantion**
SOM is an unsupervised learning algorithm and it tries to learn the probabilistic distribution of the data¶
Then, in principle, we can bias it toward certain target variable or after training we ask conditional questions.¶
For classification there are two main approaches¶
to train one SOM with/without labels. Then, we project the new data to SOM and look what are the labels around the selected node. using this method, it is possible to come up with probabilistic classifications
** To train one SOM for each class/labels. Then, project a new data points to all of these SOMs and see the activation of each SOM. This also can be used for probabilistic classification**
** For function approximation/prediction we have similar set up, but comparing to other function approximation methods, here there is no out put/response variable. Therefore, we can ask questions from any directions**
We will discuss these issues on session of classification/function approximation¶
How to interpret SOM algorithm?¶
Two types of space transformations¶
PCA as a global geometric space transformation¶
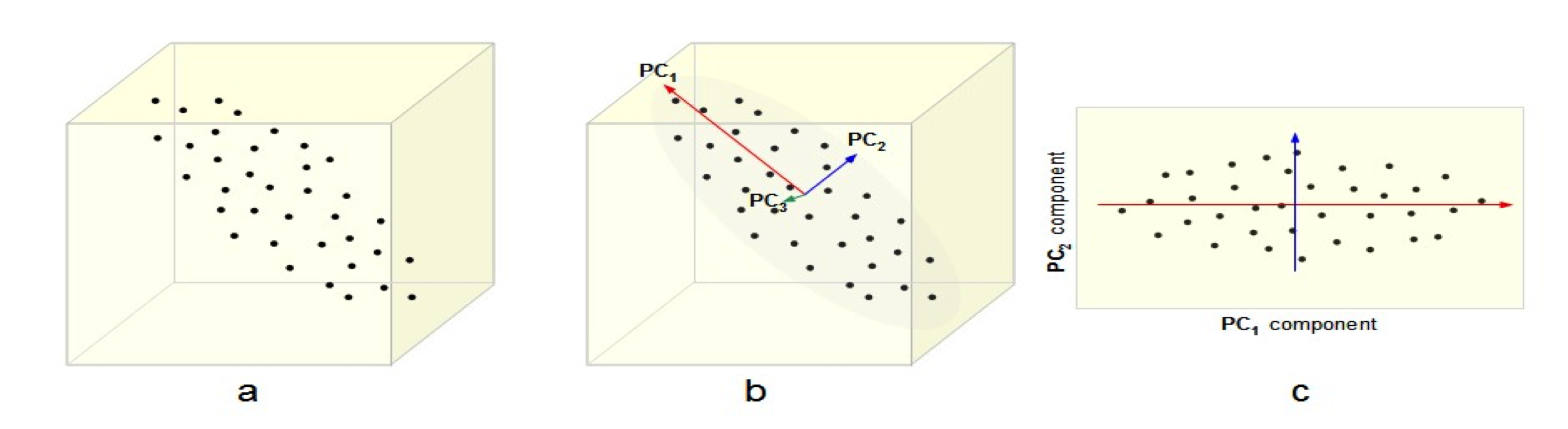
Local toology preserving transformations¶

Picture is for LLE by the way¶
SOM is more toward the second approach¶
In that sense, usually it is considered as dimensionality reduction method¶
However, in addition to dimensionality reduction method, SOM has data abstraction in the nodes. In that sense, similar to PCA, Sparse Coding or K-means with large K, each node of the SOM can be seen as a new dimension.¶
In this way, we can look at SOM in a different way, using dot products.¶
In [233]:
# We use MNIST DATA
from sklearn.datasets import fetch_mldata
DATA_PATH = './Data/MNIST/'
mnist = fetch_mldata('MNIST original', data_home=DATA_PATH)
In [234]:
train = mnist.data[:60000]
# train[train>0]=1
test = mnist.data[60000:]
# test[test>0]=1
Data_tr = train + 1e-32*np.random.randn(train.shape[0],train.shape[1])
# Data_tr = train
labels = mnist.target[:]
test_labels = labels[60000:]
train_labels = labels[:60000]
In [235]:
fig = plt.imshow(train[20000].reshape(28,28),cmap='gray')
In [236]:
tr_low = 0
tr_up = 60 *1000
msz1 = 50
msz0 = 50
In [712]:
Tr_data = Data_tr[tr_low:tr_up].astype(float)
perm = np.random.permutation(Tr_data.shape[0])
somMNIST = SOM.SOM('som1', Tr_data[perm], mapsize = [msz0, msz1],norm_method = 'var',initmethod='random')
# som1 = SOM.SOM('som1', D, mapsize = [1, 100],norm_method = 'var',initmethod='pca')
somMNIST.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook_MNIST = somMNIST.codebook[:]
Total time elapsed: 19.729000 secodns final quantization error: 20.612293
In [704]:
codebook_MNIST_n = denormalize_by(somMNIST.data_raw, codebook_MNIST, n_method = 'var')
In [705]:
c = 0;
fig = plt.figure(figsize=(30,30))
for i in range(msz0):
for j in range(msz1):
plt.subplot(msz0,msz1,c+1);
plt.imshow(codebook_MNIST_n[c].reshape(28,28),cmap='gray')
c = c +1
plt.xticks([])
plt.yticks([])
fname = path = "./Data/MINISTSOM.jpeg"
fig.savefig(fname, dpi=200)
PCA¶
In [810]:
from sklearn.decomposition import PCA
K = 400
pca = PCA(n_components=K)
pca.fit(Tr_data)
Out[810]:
PCA(copy=True, n_components=400, whiten=False)
In [807]:
fig = plt.figure(figsize=(30,30))
for i in range(400):
plt.subplot(20,20,i+1)
plt.imshow(pca.components_[i,:].reshape(28,28),cmap=plt.cm.gray);
plt.xticks([]);
plt.yticks([]);
plt.tight_layout();
In [815]:
# Some test data
import random
ind_row_test = random.sample(range(test.shape[0]),9)
ind_row_test = range(5000,5009)
x_to_proj = test[ind_row_test]
labels_to_proj = test_labels[ind_row_test]
print x_to_proj.shape
print labels_to_proj
(9, 784) [ 4. 4. 4. 4. 4. 4. 4. 4. 4.]
In [297]:
# Adding nonlinearities and normalizing the results
def sigmoid(x):
return 1/(1+np.exp(-.03*x))
def tanh(x):
return sigmoid(2*x) - sigmoid(-2*x)
With SOM¶
In [817]:
activation = x_to_proj.dot(codebook_MNIST_n.T)
fig = plt.figure(figsize=(10,10));
for i in range(x_to_proj.shape[0]):
plt.subplot(3,3,i+1);
sig_acivatio = sigmoid(activation[i])
sig_acivatio = tanh(activation[i])
sig_acivatio = activation[i]
plt.imshow(sig_acivatio.reshape(msz0,msz1),cmap=plt.cm.RdYlBu_r);
plt.title(labels_to_proj[i]);
plt.xticks([]);
plt.yticks([]);
With PCA¶
In [818]:
activation = pca.transform(x_to_proj)
# activation = x_to_proj.dot(pca.components_.T)
fig = plt.figure(figsize=(10,10));
for i in range(x_to_proj.shape[0]):
plt.subplot(3,3,i+1);
# sig_acivatio = sigmoid(activation[i])
sig_acivatio = tanh(activation[i])
# sig_acivatio = activation[i]
plt.imshow(sig_acivatio.reshape(20,20),cmap=plt.cm.RdYlBu_r);
plt.title(labels_to_proj[i]);
plt.xticks([]);
plt.yticks([]);
Another way would be train one SOM for each class¶
In [412]:
tr_low = 0
tr_up = 5500
msz1 = 20
msz0 = 20
Tr_data = Data_tr[tr_low:tr_up].astype(float)
perm = np.random.permutation(Tr_data.shape[0])
somMNIST_0 = SOM.SOM('som1', Tr_data[perm], mapsize = [msz0, msz1],norm_method = 'var',initmethod='random')
# som1 = SOM.SOM('som1', D, mapsize = [1, 100],norm_method = 'var',initmethod='pca')
somMNIST_0.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook_MNIST_0 = somMNIST_0.codebook[:]
codebook_MNIST_n_0 = denormalize_by(somMNIST_0.data_raw, codebook_MNIST_0, n_method = 'var')
Total time elapsed: 0.555000 secodns final quantization error: 22.073439
In [413]:
c = 0;
fig = plt.figure(figsize=(20,20))
for i in range(msz0):
for j in range(msz1):
plt.subplot(msz0,msz1,c+1);
plt.imshow(codebook_MNIST_n_0[c].reshape(28,28),cmap='gray')
c = c +1
plt.xticks([])
plt.yticks([])
In [416]:
tr_low = 6000
tr_up = 10000
msz1 = 20
msz0 = 20
Tr_data = Data_tr[tr_low:tr_up].astype(float)
perm = np.random.permutation(Tr_data.shape[0])
somMNIST_1 = SOM.SOM('som1', Tr_data[perm], mapsize = [msz0, msz1],norm_method = 'var',initmethod='random')
# som1 = SOM.SOM('som1', D, mapsize = [1, 100],norm_method = 'var',initmethod='pca')
somMNIST_1.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook_MNIST_1 = somMNIST_1.codebook[:]
codebook_MNIST_n_1 = denormalize_by(somMNIST_1.data_raw, codebook_MNIST_1, n_method = 'var')
Total time elapsed: 0.454000 secodns final quantization error: 20.015139
In [417]:
c = 0;
fig = plt.figure(figsize=(20,20))
for i in range(msz0):
for j in range(msz1):
plt.subplot(msz0,msz1,c+1);
plt.imshow(codebook_MNIST_n_1[c].reshape(28,28),cmap='gray')
c = c +1
plt.xticks([])
plt.yticks([])
In [445]:
# Now project new data to both SOMs
import random
ind_row_test = random.sample(range(test.shape[0]),9)
ind_row_test = range(1000,1009)
x_to_proj = test[ind_row_test]
labels_to_proj = test_labels[ind_row_test]
print x_to_proj.shape
print labels_to_proj
(9, 784) [ 1. 1. 1. 1. 1. 1. 1. 1. 1.]
In [446]:
# If we project new data into som trained with label 0
activation_0 = x_to_proj.dot(codebook_MNIST_n_0.T)
# If we project new data into som trained with label 1
activation_1 = x_to_proj.dot(codebook_MNIST_n_1.T)
fig = plt.figure(figsize=(10,10));
for i in range(x_to_proj.shape[0]):
plt.subplot(3,3,i+1);
# Activation with label 0
sig_acivatio = activation_0[i]
plt.plot(range(sig_acivatio.shape[0]),sig_acivatio,'g')
# Activation with label 1
sig_acivatio = activation_1[i]
plt.plot(range(sig_acivatio.shape[0]),sig_acivatio,'r')
plt.title(labels_to_proj[i]);
plt.xticks([]);
plt.yticks([]);
Different modifications to the original SOM¶
- Topology of low dimensional space
- ** planar**
- ** One-dimensional grid**
- ** Two-dimensional grid**
- cylinder
- Toroid
- Spherical SOM
- ** planar**
- ** Growing SOM**
- ** Hierarchical SOM **
- ** Growing Hierarchical SOM**
- ** Neural Gas**
- ** Generative Topographic Map**
- ** Similarity Measures**
- ** Relational SOM**
- ** Median SOM**
- Vectorial representation or Non-vectorial SOM
In [116]:
# Using the Weight vectors from the first example
Ws.shape
j = 120
All_diffs = np.zeros((Ws.shape[1],Ws.shape[0]))
for j in range(Ws.shape[1]):
diffs = []
for i in range(0,Ws.shape[0]):
diff = (Ws[i,j,:] -Ws[i-1,j,:]).sum()
diffs.append(diff)
All_diffs[j] = diffs
plt.plot(range(2,Ws.shape[0]),diffs[2:],'-b');
In [117]:
msz1 =20
msz0 = 20
somConverge= SOM.SOM('som1', All_diffs[:,2:], mapsize = [msz0, msz1],norm_method = 'var',initmethod='random')
# som1 = SOM.SOM('som1', D, mapsize = [1, 100],norm_method = 'var',initmethod='pca')
somConverge.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook_Converge = somConverge.codebook[:]
Total time elapsed: 0.390000 secodns final quantization error: 5.734004
In [118]:
c = 0;
fig = plt.figure(figsize=(30,30))
for i in range(msz0):
for j in range(msz1):
plt.subplot(msz0,msz1,c+1);
plt.plot(range(codebook_Converge.shape[1]),codebook_Converge[c],'-k',linewidth=.8)
c = c +1
plt.xticks([])
plt.yticks([])
In [119]:
somConverge.hit_map()
Another extension to original SOM¶
Since all the nodes have similar dimesnions, they can only be in between training data¶
What if each node looks at the world in a different way?¶
We implement this like a multi-agent system, where each node is an agent with a different weight vector¶
In order to do so, we introduce a mask in the BMU selection step. In fact, now we need a weighted distance.¶
In [188]:
import numpy as np
def fast_wdist(A, B, IM):
"""
Compute the weighted euclidean distance between two arrays of points:
D{i,j} =
sqrt( ((A{0,i}-B{0,j})/W{0,i})^2 + ... + ((A{k,i}-B{k,j})/W{k,i})^2 )
inputs:
A is an (k, m) array of coordinates
B is an (k, n) array of coordinates
IM is an (k, m) array of weights
returns:
D is an (m, n) array of weighted euclidean distances
"""
# compute the differences and apply the weights in one go using
# broadcasting jujitsu. the result is (n, k, m)
wdiff = (A[np.newaxis,...] - B[np.newaxis,...].T) * IM[np.newaxis,...]
# square and sum over the second axis, take the sqrt and transpose. the
# result is an (m, n) array of weighted euclidean distances
D = np.sqrt((wdiff*wdiff).sum(1))
return D
def find_bmu_Weighted(W,data,IM):
import scipy.spatial.distance as DIST
Dists = fast_wdist(W.T, data.T, IM.T)
ind_sets = np.argmin(Dists,axis=1)
return Dists,ind_sets
In [282]:
def train_som_online_Weighted(data,mapsize,trainlen,IM):
# mapsize = [10,10]
dlen = len(data)
#normalize the data
data_n = normalize_by(data, data, method='var');
#Initialize the map
W = random_initialize(mapsize,data_n);
UD2 = calculate_UD2(mapsize);
alpha0 = .5
if min(mapsize)==1:
radiusin = max(mapsize)/float(2)
else:
radiusin = max(mapsize)/float(6)
radiusfin = max([1,radiusin/float(4)])
radiusfin = radiusin/float(10)
print radiusin,radiusfin
# trainlen= 200*nnodes/float(dlen)
# trainlen = int(np.ceil(trainlen))
radius = np.linspace(radiusin, radiusfin, trainlen)
qe = []
Ws = []
W_denorm = denormalize_by(data, W.copy(), n_method = 'var')
Ws.append(W_denorm.copy())
for i in range(trainlen):
#all or a sample
data_e = np.random.permutation(data_n)[:]
qe_t = np.zeros((data_e.shape[0],1))
for j, d in enumerate(data_e):
#project using a distance function
#find bmu
#Competition between nodes now based on the importance in each dimensions
Dists,bmu = find_bmu_Weighted(W,d[np.newaxis,:],IM)
# Dists,bmu = find_bmu(d[np.newaxis,:],W)
qe_t[j] = np.min(Dists)
# Adapt nodes
#distance to bmu in the lower dimensional space
neiboring_effect = calculate_neigh_activation(UD2,bmu,radius[i])
#learning rate
alpha = calc_alpha(i,alpha0,trainlen,method='inv')
#Adapatation
W[:] = W[:] + neiboring_effect.T * alpha * np.multiply(IM,(d-W[:]))
W_denorm = denormalize_by(data, W.copy(), n_method = 'var')
Ws.append(W_denorm.copy())
if i%10==0:
print 'iteration {}: alpha: {}, raduis: {} and quantization error is {}'.format(i,alpha,radius[i],np.mean(qe_t))
qe.append(np.mean(qe_t))
Ws = np.asarray(Ws)
return W, qe,Ws
In [303]:
data = Data1
# data = D.values[:,:-1].astype(float)
mapsize = [40,40]
trainlen = 200
# just repeat the dimensions
Wdata = np.tile(data,(1,4))
nnodes = mapsize[0]*mapsize[1]
IM = np.random.rand(nnodes,Wdata.shape[1])
IM = tanh(IM)
# IM = np.ones((nnodes,Wdata.shape[1]))
S = IM.sum(axis=1)[:,np.newaxis]
# IM = IM/S
# IM[IM>=.25]=1
# IM[IM<.25]=.01
# Now IM says the importance of each dimension for each node
W_IM, qe_IM,Ws_IM = train_som_online_Weighted(Wdata,mapsize,trainlen,IM)
# W, qe,Ws = train_som_online(data,mapsize,trainlen)
codebook_IM = denormalize_by(Wdata, W_IM, n_method = 'var')
6.66666666667 0.666666666667 iteration 0: alpha: 0.5, raduis: 6.66666666667 and quantization error is 0.0163359281554 iteration 10: alpha: 0.0833333333333, raduis: 6.36515912898 and quantization error is 0.00654020351724 iteration 20: alpha: 0.0454545454545, raduis: 6.06365159129 and quantization error is 0.00581916974755 iteration 30: alpha: 0.03125, raduis: 5.7621440536 and quantization error is 0.00541026595449 iteration 40: alpha: 0.0238095238095, raduis: 5.46063651591 and quantization error is 0.00511596247526 iteration 50: alpha: 0.0192307692308, raduis: 5.15912897822 and quantization error is 0.00488666566941 iteration 60: alpha: 0.0161290322581, raduis: 4.85762144054 and quantization error is 0.00470234541166 iteration 70: alpha: 0.0138888888889, raduis: 4.55611390285 and quantization error is 0.00454548278697 iteration 80: alpha: 0.0121951219512, raduis: 4.25460636516 and quantization error is 0.00441129912534 iteration 90: alpha: 0.0108695652174, raduis: 3.95309882747 and quantization error is 0.00429692392467 iteration 100: alpha: 0.00980392156863, raduis: 3.65159128978 and quantization error is 0.00420410797779 iteration 110: alpha: 0.00892857142857, raduis: 3.35008375209 and quantization error is 0.00413043762322 iteration 120: alpha: 0.00819672131148, raduis: 3.04857621441 and quantization error is 0.00406840683725 iteration 130: alpha: 0.00757575757576, raduis: 2.74706867672 and quantization error is 0.00401764613451 iteration 140: alpha: 0.00704225352113, raduis: 2.44556113903 and quantization error is 0.00397515596563 iteration 150: alpha: 0.00657894736842, raduis: 2.14405360134 and quantization error is 0.00393811763255 iteration 160: alpha: 0.00617283950617, raduis: 1.84254606365 and quantization error is 0.00390761151723 iteration 170: alpha: 0.00581395348837, raduis: 1.54103852596 and quantization error is 0.00388199421278 iteration 180: alpha: 0.00549450549451, raduis: 1.23953098827 and quantization error is 0.00386081008274 iteration 190: alpha: 0.00520833333333, raduis: 0.938023450586 and quantization error is 0.00384333993677
In [316]:
fig = plt.figure()
plt.plot(data[:,0],data[:,1],'.')
plt.plot(codebook_IM[:,0],codebook_IM[:,1],'.r')
plt.plot(codebook_IM[:,2],codebook_IM[:,3],'.g')
plt.plot(codebook_IM[:,4],codebook_IM[:,5],'.k')
# plt.plot(codebook[:,0],codebook[:,1],'.k')
fig.set_size_inches(7,7)
In [313]:
def viewmap(codebook,msz):
fig = plt.figure(figsize=(20,14))
dim = codebook.shape[1]
no_row_in_plot = dim/6 + 1 #6 is arbitrarily selected
if no_row_in_plot <=1:
no_col_in_plot = dim
else:
no_col_in_plot = 6
for k in range(dim):
ax = plt.subplot(no_row_in_plot,no_col_in_plot,k+1)
ax.set_yticklabels([])
ax.set_xticklabels([])
ax.imshow(codebook[:,k].reshape(msz[0],msz[1])[::],alpha=1,cmap=plt.cm.RdYlBu_r)
# ax.imshow(som.codebook[:,k].reshape(msz[0],msz[1])[::],alpha=1,cmap=plt.cm.Accent_r)
plt.subplots_adjust(hspace = .001,wspace=.01)
# plt.tight_layout()
In [314]:
viewmap(codebook_IM,mapsize)
Contextual Numbers¶
How to index high dimensional spaces in to one dimensional index space, like a number space.¶
- Some related works:
- 1- Vahid, Moosavi, Computing With Contextual Numbers. arXiv preprint arXiv:1408.0889. (2014).
- ** 2- Vahid Moosavi, Contextual Mapping: Visualization of High Dimensional Spatial Patterns in a Single Geo-Map, (Computers, Environment and Urban Systems 61 (2017) 1–12).**
In [392]:
x = np.linspace(-5,5,100)[:,np.newaxis]
y = x**2 + 1*x**3
# y = np.sqrt(7-x**2)
plt.plot(x,y,'.-')
# y = -1*np.sqrt(7-x**2)
# plt.plot(x,y)
Data5 = np.concatenate((x,y),axis=1)
In [393]:
msz1 =1
msz0 = 50
data = Data5
somCN= SOM.SOM('som1', data, mapsize = [msz0, msz1],norm_method = 'var',initmethod='pca')
# som1 = SOM.SOM('som1', D, mapsize = [1, 100],norm_method = 'var',initmethod='pca')
somCN.train(n_job = 1, shared_memory = 'no',verbose='final')
codebook_CN= denormalize_by(somCN.data_raw, somCN.codebook[:], n_method = 'var')
Total time elapsed: 0.238000 secodns final quantization error: 0.036120
In [395]:
fig = plt.figure()
K = codebook_CN.shape[0]
plt.plot(data[:,0],data[:,1],'.',alpha=.4)
c = range(codebook_CN.shape[0])
plt.scatter(codebook_CN[:,0],codebook_CN[:,1],s=60,marker='o',alpha=1.,c=plt.cm.RdYlBu_r(np.asarray(c)/float(K)));
fig.set_size_inches(7,7);
Further Discusssions¶
- Using Contextual numbers in the context of dynamical systems
- Looking at SOM not as a deterministic machine, but as a probabilistic machine!
- Resampling application
- Using online randomized SOM for several runs to find invariances