The NumPy array - and a few other things¶
A structure for efficient numerical computation¶
Most material here is from: Stéfan van der Walt (@stefanvdwalt)Some from Nicolas Rougier's book ("From Python to Numpy")
Some additions by JB Poline Nipype Workshop, Boston
Accompanying problem sets are on the Wiki at: https://python.g-node.org/wiki/numpy

Num-what?¶
This talk discusses some of the more advanced NumPy features. If you’ve never seen NumPy before, you may have more fun doing the tutorial at
http://mentat.za.net/numpy/intro/intro.html
or studying the course notes at
http://scipy-lectures.github.io
You can always catch up by reading:
The NumPy array: a structure for efficient numerical computation. Stéfan van der Walt, S. Chris Colbert and Gaël Varoquaux. In IEEE Computing in Science Engineering, March/April 2011.
Setup¶
import numpy as np # We always use this convention,
# also in the documentation
print(np.__version__) # version 1.7 or greater
# Version 1.9 released 7th of September!
1.11.2
ATLAS is a fast implementation of BLAS (Basic Linear Algebra Routines). On
OSX you have Accelerate; students can get Intel's MKL for free. On Ubuntu,
install libatlas3-base.
Make use of IPython's powerful features! TAB-completion, documentation, source inspection, timing, cpaste, etc.
print(np.show_config ()) # got ATLAS / Accelerate / MKL?
lapack_opt_info:
define_macros = [('HAVE_CBLAS', None)]
language = c
library_dirs = ['/usr/local/lib']
libraries = ['openblas', 'openblas']
openblas_info:
define_macros = [('HAVE_CBLAS', None)]
language = c
library_dirs = ['/usr/local/lib']
libraries = ['openblas', 'openblas']
blas_opt_info:
define_macros = [('HAVE_CBLAS', None)]
language = c
library_dirs = ['/usr/local/lib']
libraries = ['openblas', 'openblas']
blas_mkl_info:
NOT AVAILABLE
openblas_lapack_info:
define_macros = [('HAVE_CBLAS', None)]
language = c
library_dirs = ['/usr/local/lib']
libraries = ['openblas', 'openblas']
None
Ultra fast primer¶
# Array are not list
l = [1,2]
a = np.asarray(l)
print('l*2: ', l*2, '; a*2: ', a*2)
#
print('l+l+l: ', l+l+l, '; a+a+a: ',a+a+a)
l*2: [1, 2, 1, 2] ; a*2: [2 4] l+l+l: [1, 2, 1, 2, 1, 2] ; a+a+a: [3 6]
Arrays are mutable¶
So, if you pass an array as an argument and change it in a function...
a = np.arange(4)
def change_my_array(arr):
arr[0] = -10
return None
print(a)
change_my_array(a)
print(a)
[0 1 2 3] [-10 1 2 3]
# READABILITY
def function_1(seq, sub):
return [i for i in range(len(seq) - len(sub)) if seq[i:i+len(sub)] == sub]
def function_2(seq, sub):
target = np.dot(sub, sub)
candidates = np.where(np.correlate(seq, sub, mode='valid') == target)[0]
check = candidates[:, np.newaxis] + np.arange(len(sub))
mask = np.all((np.take(seq, check) == sub), axis=-1)
return candidates[mask]
from timeit import timeit
n, m = 5000, 200
s = np.kron(np.arange(n),np.eye(m)).reshape(n*m*m)
print(s.shape)
%timeit("function_1(s, [6,7,8,9])")
%timeit("function_2(s, [6,7,8,9])")
(200000000,) 100000000 loops, best of 3: 7.2 ns per loop 100000000 loops, best of 3: 7.15 ns per loop


Taking a look at numpy/core/include/numpy/ndarraytypes.h:
ndarray typedef struct PyArrayObject {
PyObject_HEAD
char *data; /* pointer to data buffer */
int nd; /* number of dimensions */
npy_intp *dimensions; /* size in each dimension */
npy_intp *strides; /* bytes to jump to get
* to the next element in
* each dimension
*/
PyObject *base; /* Pointer to original array
/* Decref this object */
/* upon deletion. */
PyArray_Descr *descr; /* Pointer to type struct */
int flags; /* Flags */
PyObject *weakreflist; /* For weakreferences */
} PyArrayObject ;
a = np.ones((3,4))
print(a)
print(dir(a))
[[ 1. 1. 1. 1.] [ 1. 1. 1. 1.] [ 1. 1. 1. 1.]] ['T', '__abs__', '__add__', '__and__', '__array__', '__array_finalize__', '__array_interface__', '__array_prepare__', '__array_priority__', '__array_struct__', '__array_wrap__', '__bool__', '__class__', '__contains__', '__copy__', '__deepcopy__', '__delattr__', '__delitem__', '__dir__', '__divmod__', '__doc__', '__eq__', '__float__', '__floordiv__', '__format__', '__ge__', '__getattribute__', '__getitem__', '__gt__', '__hash__', '__iadd__', '__iand__', '__ifloordiv__', '__ilshift__', '__imatmul__', '__imod__', '__imul__', '__index__', '__init__', '__int__', '__invert__', '__ior__', '__ipow__', '__irshift__', '__isub__', '__iter__', '__itruediv__', '__ixor__', '__le__', '__len__', '__lshift__', '__lt__', '__matmul__', '__mod__', '__mul__', '__ne__', '__neg__', '__new__', '__or__', '__pos__', '__pow__', '__radd__', '__rand__', '__rdivmod__', '__reduce__', '__reduce_ex__', '__repr__', '__rfloordiv__', '__rlshift__', '__rmatmul__', '__rmod__', '__rmul__', '__ror__', '__rpow__', '__rrshift__', '__rshift__', '__rsub__', '__rtruediv__', '__rxor__', '__setattr__', '__setitem__', '__setstate__', '__sizeof__', '__str__', '__sub__', '__subclasshook__', '__truediv__', '__xor__', 'all', 'any', 'argmax', 'argmin', 'argpartition', 'argsort', 'astype', 'base', 'byteswap', 'choose', 'clip', 'compress', 'conj', 'conjugate', 'copy', 'ctypes', 'cumprod', 'cumsum', 'data', 'diagonal', 'dot', 'dtype', 'dump', 'dumps', 'fill', 'flags', 'flat', 'flatten', 'getfield', 'imag', 'item', 'itemset', 'itemsize', 'max', 'mean', 'min', 'nbytes', 'ndim', 'newbyteorder', 'nonzero', 'partition', 'prod', 'ptp', 'put', 'ravel', 'real', 'repeat', 'reshape', 'resize', 'round', 'searchsorted', 'setfield', 'setflags', 'shape', 'size', 'sort', 'squeeze', 'std', 'strides', 'sum', 'swapaxes', 'take', 'tobytes', 'tofile', 'tolist', 'tostring', 'trace', 'transpose', 'var', 'view']
A homogeneous container¶
char *data; /* pointer to data buffer */
Data is just a pointer to bytes in memory:
<img src="images/array_dtype.png"/ width="60%">
x = np.array([1, 2, 3])
x.dtype
dtype('int64')
x.__array_interface__['data']
(32558752, False)
print(x.data)
<memory at 0x7f5f8c0fd588>
Dimensions¶
int nd;
npy_intp *dimensions; /* number of dimensions */
/* size in each dimension */



x = np.array([])
np.array(0).shape
()
x = np.random.random((3, 2, 3, 3))
x.shape
(3, 2, 3, 3)
Data type descriptors¶
PyArray_Descr * descr ; /* Pointer to type struct */
Common types in include int, float, bool:
np.array ([-1, 0, 1], dtype=int)
array([-1, 0, 1])
np.array([-1, 0, 1], dtype=float)
array([-1., 0., 1.])
np.array([-1 , 0, 1], dtype=bool)
array([ True, False, True], dtype=bool)
Each item in the array has to have the same type (occupy a fixed nr of bytes in memory), but that does not mean a type has to consist of a single item:
dt = np.dtype([('value', np.int), ('status', np.bool)])
np.array([(0, True), (1, False)], dtype=dt)
array([(0, True), (1, False)],
dtype=[('value', '<i8'), ('status', '?')])
This is called a structured array.
Strides¶
npy_intp *strides; /* bytes to jump to get */
/* to the next element */
x = np.arange(12).reshape((3, 4))
x
array([[ 0, 1, 2, 3],
[ 4, 5, 6, 7],
[ 8, 9, 10, 11]])
x.dtype
dtype('int64')
x.dtype.itemsize
8
x.strides
(32, 8)
We see (4 * itemsize, itemsize) or (skip_bytes_row, skip_bytes_col).

Example: Taking the transpose¶
We can take the transpose of the array without copying any data by simply manipulating shape and strides. What should they be?

View versus copy¶
A copy will allocate new memory, a view will not¶
A change in the copy will not affect the original array, a change in the view will¶
Some numpy operation return copies, others return views¶
- flatten return a copy
- slicing will return a view
- ravel will attempt to return a view
- reshape returns a view
Check if array is a copy or a view with flags¶
Z = np.zeros((5,5))
Z.ravel().base is Z == True
#
Z[::2,::2].ravel().base is Z == True
#
assert(Z.flatten().base is not Z)
#
assert(Z.reshape((25,1)).base is Z)
Z1 = np.arange(10)
Z2 = Z1[1:-1:2]
print(Z2)
[1 3 5 7]
Flags¶
int flags; /* Flags */
x = np.array([1, 2, 3])
z = x[::2]
print(z)
print(x.flags)
[1 3] C_CONTIGUOUS : True F_CONTIGUOUS : True OWNDATA : True WRITEABLE : True ALIGNED : True UPDATEIFCOPY : False
z.flags
C_CONTIGUOUS : False F_CONTIGUOUS : False OWNDATA : False WRITEABLE : True ALIGNED : True UPDATEIFCOPY : False
Temporary copy¶
>>> X = np.ones(10, dtype=np.int)
>>> Y = np.ones(10, dtype=np.int)
>>> A = 2*X + 2*Y
Versus:
>>> X = np.ones(10, dtype=np.int)
>>> Y = np.ones(10, dtype=np.int)
>>> np.multiply(X, 2, out=X)
>>> np.multiply(Y, 2, out=Y)
>>> np.add(X, Y, out=X)
Structured arrays¶
Like we saw earlier, each item in an array has the same type, but that does not mean a type has to consist of a single item:
dt = np.dtype([('value', np.int), ('status', np.bool)])
np.array([(0, True), (1,False)], dtype=dt)
array([(0, True), (1, False)],
dtype=[('value', '<i8'), ('status', '?')])
This is called a structured array, and is accessed like a dictionary:
x = np.array([(0, True), (1, False)], dtype=dt)
x['value']
array([0, 1])
x['status']
array([ True, False], dtype=bool)
Reading structured data from file¶
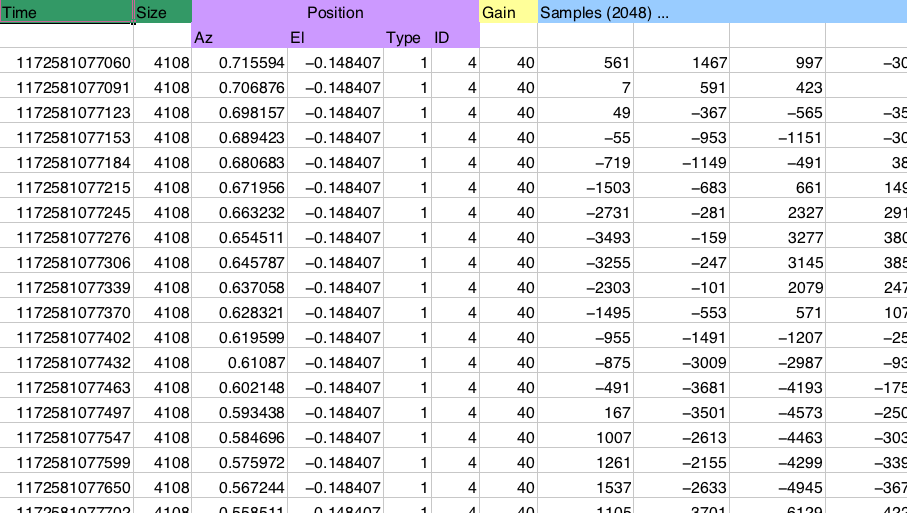
Reading this kind of data can be troublesome:
while ((count > 0) && (n <= NumPoints))
% get time - I8 [ ms ]
[lw, count] = fread(fid, 1, 'uint32');
if (count > 0) % then carry on
uw = fread(fid, 1, 'int32');
t(1, n) = (lw + uw * 2^32) / 1000;
% get number of bytes of data
numbytes = fread(fid, 1, 'uint32');
% read sMEASUREMENTPOSITIONINFO (11 bytes)
m(1, n) = fread(fid, 1, 'float32'); % az [ rad ]
m(2, n) = fread(fid, 1, 'float32'); % el [ rad ]
m(3, n) = fread(fid, 1, 'uint8'); % region type
m(4, n) = fread(fid, 1, 'uint16'); % region ID
g(1, n) = fread(fid, 1, 'uint8');
numsamples = ( numbytes -12)/2; % 2 byte int
...
Using structured arrays:
dt = np.dtype([('time', np.uint64),
('size', np.uint32),
('position', [('az', np.float32),
('el', np.float32),
('region_type', np.uint8),
('region_ID', np.uint16)]),
('gain', np.uint8),
('samples', (np.int16, 2048))])
data = np.fromfile(f, dtype=dt)
We can then access this structured array as before:
data['position']['az']
Array functions¶
a = np.random.normal((3,30))
print(a)
a = np.random.normal(0,1, size=(3,10))
print(a)
a = np.random.randint(0,10, size=(3,10))
print(a)
[ 0.64488785 29.78139454] [[ 1.69439088 0.72536413 -2.02163215 -1.00667092 1.8816507 -0.03026824 -0.93375708 0.14278375 -0.09448514 0.41436576] [ 1.715446 1.08301563 1.59472585 0.50601605 -0.5222702 -0.53497445 -2.05725302 -1.86172097 0.56197185 0.49018729] [-0.06686053 -0.1104309 -1.20906256 -0.70505399 1.41420279 1.49779434 -0.22250591 -0.09190751 0.67891195 0.01016829]] [[6 5 0 4 7 9 1 9 6 7] [2 5 3 4 4 4 1 1 2 3] [5 6 7 8 6 2 4 5 6 6]]
print("a.min(), a.max(), a.mean(), a.std(), a.sum()")
print(a.min(), a.max(), a.mean(), a.std(), a.sum())
a = np.arange(7)
print(a.cumsum())
a = np.arange(1,7)
print(a.cumprod())
a.min(), a.max(), a.mean(), a.std(), a.sum() 1 6 3.5 1.70782512766 21 [ 0 1 3 6 10 15 21] [ 1 2 6 24 120 720]
Broadcasting¶
Combining of differently shaped arrays without creating large intermediate arrays:
x = np.arange(4)
print(x)
print(x + 3)
[0 1 2 3] [3 4 5 6]
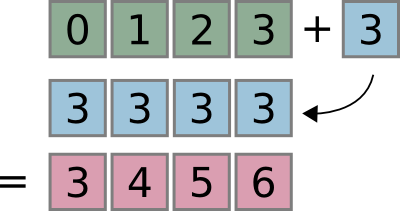
See docstring of np.doc.broadcasting for more detail.
Broadcasting (2D)¶
a = np.arange(12).reshape((3, 4))
b = np.array([1, 2, 3])[:, np.newaxis]
print(a)
print()
print(b)
[[ 0 1 2 3] [ 4 5 6 7] [ 8 9 10 11]] [[1] [2] [3]]
a + b
array([[ 1, 2, 3, 4],
[ 6, 7, 8, 9],
[11, 12, 13, 14]])

Broadcasting (3D)¶
x = np.zeros((3, 5))
print(x.shape)
y = np.zeros(8)
z = x[..., np.newaxis]
print(z.shape)
w = z + y
print(w.shape)
(3, 5) (3, 5, 1) (3, 5, 8)
x = np.zeros((3, 5))
y = np.zeros(8)
z = x[..., np.newaxis]
X shape = (3, 5, 1)
Y shape = ( 8)
Z shape = (3, 5, 8)

Broadcasting rules¶
The broadcasting rules are straightforward: compare dimensions, starting from
the last. Match when either dimension is one or None, or if dimensions are
equal:
Scalar 2D 3D Bad
( ,) (3, 4) (3, 5, 1) (3, 5, 2)
(3,) (3, 1) ( 8) ( 8)
---- ------ --------- ---------
(3,) (3, 4) (3, 5, 8) XXX
a = np.random.normal(size=(3,1,5,1))
b = np.random.normal(size=(7,1,4))
c=a+b
c.shape
(3, 7, 5, 4)
Explicit broadcasting¶
x = np.zeros((3, 5, 1))
y = np.zeros((3, 5, 8))
xx, yy = np.broadcast_arrays(x, y)
xx.shape
(3, 5, 8)
np.broadcast_arrays([1, 2, 3],
[[1], [2], [3]])
[[1 2 3] [[1 1 1]
[1 2 3] [2 2 2]
[1 2 3]] [3 3 3]]
Some useful array manipulations¶
# meshgrid ?
# np.where
# boolean operation
a = np.arange(12).reshape(3,4)
b = (a == 6)
print(b)
b = np.logical_or(a == 6, a == 7)
print(b)
a[b]
[[False False False False] [False False True False] [False False False False]] [[False False False False] [False False True True] [False False False False]]
array([6, 7])
print(a)
print(np.where(a>3))
print(a[np.where(a>3)])
print(a > 3)
[[ 0 1 2 3] [ 4 5 6 7] [ 8 9 10 11]] (array([1, 1, 1, 1, 2, 2, 2, 2]), array([0, 1, 2, 3, 0, 1, 2, 3])) [ 4 5 6 7 8 9 10 11] [[False False False False] [ True True True True] [ True True True True]]
Axis operation¶
# reshape
# flatten / ravel
# transpose
# axis rotation
a = np.arange(24).reshape(1,2,3,4)
a.T.shape
b = np.rollaxis(a, 2, 1)
print(b.shape)
b = np.rollaxis(a, 2, 0)
print(b.shape)
b = np.rollaxis(a, 2, 3)
print(b.shape)
(1, 3, 2, 4) (3, 1, 2, 4) (1, 2, 3, 4)
hstack - vstack¶
a1 = np.ones((1,10))
a3 = np.ones((3,10))
# vstack
a4 = np.vstack((a1,a3))
print(a4.shape)
# hstack
a10 = np.hstack((a1.T, a3.T))
print(a10.shape)
print(np.concatenate((a1, a3), axis=0).shape)
print(np.concatenate((a1.T, a3.T), axis=1).shape)
(4, 10) (10, 4) (4, 10) (10, 4)
dot product: at the basis of so many things :)¶
I = np.eye(4)
a = np.random.lognormal(size=(4,4))
print(a)
b = a.dot(I)
print(np.allclose(b, a))
print(((a-b)**2).sum())
[[ 1.38102561 0.27409858 1.26081074 0.2600873 ] [ 0.86771584 0.85718828 0.21770531 0.94686422] [ 5.09592854 0.25703762 2.07068062 0.36530305] [ 1.4798213 0.37511234 7.01844686 2.6445728 ]] True 0.0
b@I
array([[ 1.38102561, 0.27409858, 1.26081074, 0.2600873 ],
[ 0.86771584, 0.85718828, 0.21770531, 0.94686422],
[ 5.09592854, 0.25703762, 2.07068062, 0.36530305],
[ 1.4798213 , 0.37511234, 7.01844686, 2.6445728 ]])
Fancy indexing¶
An ndarray can be indexed in two ways:
- Using slices and scalars
- Using ndarrays ("fancy indexing")
For example:
x = np.arange(9).reshape((3, 3))
x
array([[0, 1, 2],
[3, 4, 5],
[6, 7, 8]])
x[:, [1, 1, 2]]
array([[1, 1, 2],
[4, 4, 5],
[7, 7, 8]])
Which is equivalent to:
np.array((x[:, 1], x[:, 1], x[:, 2])).T
array([[1, 1, 2],
[4, 4, 5],
[7, 7, 8]])
Output shape of an indexing op¶
- Broadcast all index arrays against one another.
- Use the dimensions of slices as-is.
x = np.array([[0, 1, 2], [3, 4, 5], [6, 7, 8]])
print(x)
[[0 1 2] [3 4 5] [6 7 8]]
x.shape
(3, 3)
But what would happen if we do
idx0 = np.array([[0, 1],
[1, 2]]) # row indices
idx1 = np.array([[0, 1]]) # column indices
array([[0, 4],
[3, 7]])
x[idx0, idx1]
x[idx0, idx1]
array([[0, 4],
[3, 7]])
Output shape of indexing op (cont'd)¶
idx0.shape, idx1.shape
((2, 2), (1, 2))
a, b = np.broadcast_arrays(idx0, idx1)
a
array([[0, 1],
[1, 2]])
b
array([[0, 1],
[0, 1]])
x
array([[0, 1, 2],
[3, 4, 5],
[6, 7, 8]])
x[idx0, idx1]
array([[0, 4],
[3, 7]])
Output shape of an indexing op (cont'd)¶
x = np.random.random((15, 12, 16, 3))
index_one = np.array([[0, 1], [2, 3], [4, 5]])
index_two = np.array([[0, 1]])
print(index_one.shape)
print(index_two.shape)
(3, 2) (1, 2)
Predict the output shape of
x[5:10, index_one, :, index_two]
Warning! When mixing slicing and fancy indexing, the order of the output dimensions is less easy to predict. Play it safe and don't mix the two!
Output shape of an indexing op (cont'd)¶
x = np.random.random((15, 12, 16, 3))
index_one = np.array([[0, 1], [2, 3], [4, 5]])
index_two = np.array([[0, 1]])
print(index_one.shape)
print(index_two.shape)
(3, 2) (1, 2)
Broadcast index1 against index2:
(3, 2) # shape of index1
(1, 2) # shape of index2
------
(3, 2)
The shape of x[5:10, index_one, :, index_two] is
(3, 2, 5, 16)
Jack's dilemma¶
Indexing and broadcasting are intertwined, as we’ll see in the following example. One of my favourites from the NumPy mailing list:
Date: Wed, 16 Jul 2008 16:45:37 -0500
From: Jack Cook
To: <numpy-discussion@scipy.org>
Subject: Numpy Advanced Indexing Question
Greetings,
I have an I,J,K 3D volume of amplitude values at regularly sampled time intervals. I have an I,J 2D slice which contains a time (K) value at each I, J location. What I would like to do is extract a subvolume at a constant +/- K window around the slice. Is there an easy way to do this using advanced indexing or some other method? Thanks in advanced for your help.
-- Jack
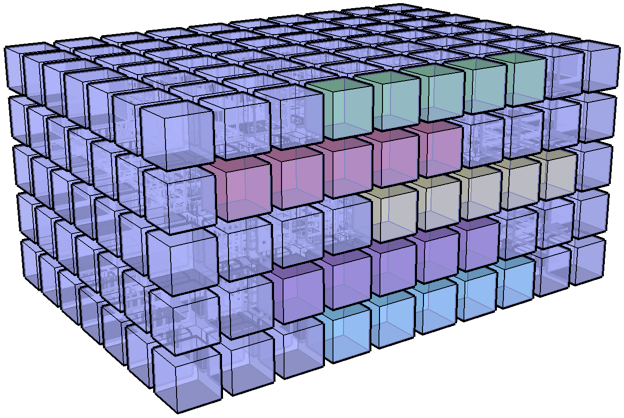
Test setup¶
ni, nj, nk = (10, 15, 20) # dimensions
Make a fake data block such that block[i, j, k] == k for all i, j, k.
block = np.empty((ni, nj, nk) , dtype=int)
block[:] = np.arange(nk)[np.newaxis, np.newaxis, :]
Pick out a random fake horizon in k:
k = np.random.randint(5, 15, size=(ni, nj))
k
array([[ 5, 14, 11, 13, 8, 12, 11, 8, 13, 8, 7, 10, 12, 8, 10],
[ 7, 5, 6, 8, 9, 10, 10, 14, 5, 14, 14, 12, 5, 8, 9],
[ 7, 14, 8, 14, 6, 6, 13, 9, 6, 5, 6, 13, 13, 11, 7],
[14, 9, 6, 12, 14, 14, 8, 10, 14, 10, 13, 12, 12, 11, 9],
[12, 11, 14, 14, 13, 5, 7, 8, 8, 14, 14, 5, 7, 8, 12],
[ 5, 7, 10, 11, 12, 12, 8, 9, 14, 13, 8, 6, 13, 8, 6],
[ 8, 6, 12, 14, 5, 12, 5, 13, 14, 13, 10, 14, 9, 7, 6],
[10, 6, 14, 14, 11, 9, 11, 8, 11, 5, 13, 13, 12, 14, 13],
[ 5, 11, 9, 5, 6, 8, 7, 11, 10, 9, 14, 5, 13, 9, 6],
[13, 12, 6, 12, 8, 13, 5, 11, 14, 9, 13, 13, 12, 10, 12]])
half_width = 3
Solution¶
These two indices assure that we take a slice at each (i, j) position
idx_i = np.arange(ni)[:, np.newaxis, np.newaxis]
idx_j = np.arange(nj)[np.newaxis, :, np.newaxis]
This is the substantitive part that picks out the window:
idx_k = k[:, :, np.newaxis] + np.arange(-half_width, half_width + 1)
slices = block[idx_i, idx_j, idx_k] # Slice
Apply the broadcasting rules:
(ni, 1, 1 ) # idx_i
(1, nj, 1 ) # idx_j
(ni, nj, 2 * half_width + 1) # idx_k
# ---
(ni, nj, 7) # <-- this is what we wanted!
(10, 15, 7)
Solution verification¶
slices = block[idx_i, idx_j, idx_k]
slices.shape
(10, 15, 7)
(10, 15, 7)
Now verify that our window is centered on k everywhere:
slices[:, :, 3]
array([[ 5, 14, 11, 13, 8, 12, 11, 8, 13, 8, 7, 10, 12, 8, 10],
[ 7, 5, 6, 8, 9, 10, 10, 14, 5, 14, 14, 12, 5, 8, 9],
[ 7, 14, 8, 14, 6, 6, 13, 9, 6, 5, 6, 13, 13, 11, 7],
[14, 9, 6, 12, 14, 14, 8, 10, 14, 10, 13, 12, 12, 11, 9],
[12, 11, 14, 14, 13, 5, 7, 8, 8, 14, 14, 5, 7, 8, 12],
[ 5, 7, 10, 11, 12, 12, 8, 9, 14, 13, 8, 6, 13, 8, 6],
[ 8, 6, 12, 14, 5, 12, 5, 13, 14, 13, 10, 14, 9, 7, 6],
[10, 6, 14, 14, 11, 9, 11, 8, 11, 5, 13, 13, 12, 14, 13],
[ 5, 11, 9, 5, 6, 8, 7, 11, 10, 9, 14, 5, 13, 9, 6],
[13, 12, 6, 12, 8, 13, 5, 11, 14, 9, 13, 13, 12, 10, 12]])
np.all(slices[:, :, 3] == k)
True
The __array_interface__¶
Any object that exposes a suitable dictionary named
__array_interface__ may be converted to a NumPy array. This is
handy for exchanging data with external libraries. The array interface
has the following important keys (see
http://docs.scipy.org/doc/numpy/reference/arrays.interface):
- shape
- typestr
- data: (20495857, True); 2-tuple—pointer to data and boolean to
indicate whether memory is read-only
- strides
- version: 3
Example: Repeating sequence¶
data = np.array([1, 2, 3, 4])
print(data.shape, data.strides)
(4,) (8,)
N = 400000
repeat = np.lib.stride_tricks.as_strided(data, shape=(N, 4), strides=(0, 8))
repeat
array([[1, 2, 3, 4],
[1, 2, 3, 4],
[1, 2, 3, 4],
...,
[1, 2, 3, 4],
[1, 2, 3, 4],
[1, 2, 3, 4]])
Questions, discussion and exercizes¶
%reload_ext load_style
%load_style css/notebook.css