Tools¶
The below line is a magic command that allows plots to appear in the notebook.
%matplotlib inline
The first thing is always to import the packages we'll use.
import pandas as pd
import numpy as np
import networkx as nx
Tutorials on pandas can be found at:
- https://pandas.pydata.org/pandas-docs/stable/10min.html
- https://pandas.pydata.org/pandas-docs/stable/tutorials.html
Tutorials on numpy can be found at:
- https://docs.scipy.org/doc/numpy/user/quickstart.html
- http://www.scipy-lectures.org/intro/numpy/index.html
- http://www.scipy-lectures.org/advanced/advanced_numpy/index.html
A tutorial on networkx can be found at:
Import the data¶
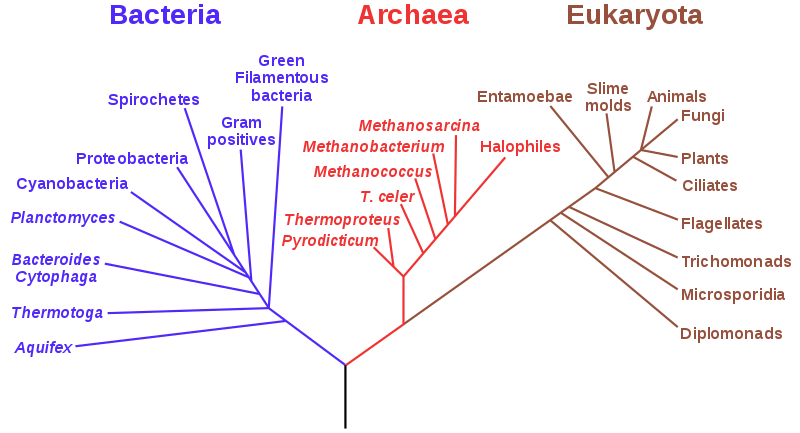
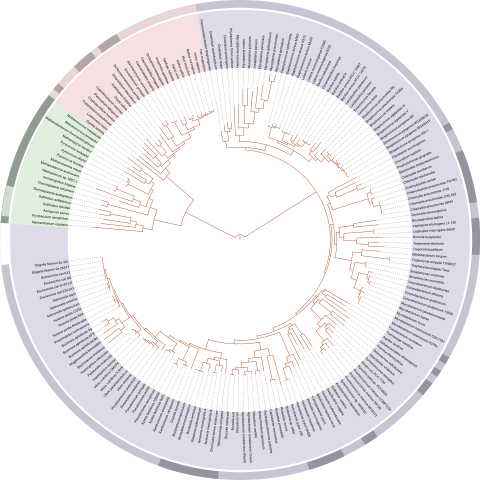
We will play with a excerpt of the Tree of Life, that can be found together with this notebook. This dataset is reduced to the first 1000 taxons (starting from the root node). The full version is available here: Open Tree of Life.
tree_of_life = pd.read_csv('data/taxonomy_small.tsv', sep='\t\|\t?', encoding='utf-8', engine='python')
If you do not remember the details of a function:
pd.read_csv?
For more info on the separator, see regex.
Now, what is the object tree_of_life? It is a Pandas DataFrame.
tree_of_life
| uid | parent_uid | name | rank | sourceinfo | uniqname | flags | Unnamed: 7 | |
|---|---|---|---|---|---|---|---|---|
| 0 | 805080 | NaN | life | no rank | silva:0,ncbi:1,worms:1,gbif:0,irmng:0 | NaN | NaN | NaN |
| 1 | 93302 | 805080.0 | cellular organisms | no rank | ncbi:131567 | NaN | NaN | NaN |
| 2 | 996421 | 93302.0 | Archaea | domain | silva:D37982/#1,ncbi:2157,worms:8,gbif:2,irmng:12 | Archaea (domain silva:D37982/#1) | NaN | NaN |
| 3 | 5246114 | 996421.0 | Marine Hydrothermal Vent Group 1(MHVG-1) | no rank - terminal | silva:AB302039/#2 | NaN | NaN | NaN |
| 4 | 102415 | 996421.0 | Thaumarchaeota | phylum | silva:D87348/#2,ncbi:651137,worms:559429,irmng... | NaN | NaN | NaN |
| 5 | 5246628 | 102415.0 | terrestrial group | no rank - terminal | silva:AB600373/#3 | NaN | NaN | NaN |
| 6 | 4795965 | 102415.0 | Marine Group I | no rank | silva:D87348/#3,ncbi:905826 | NaN | NaN | NaN |
| 7 | 5205649 | 4795965.0 | uncultured marine crenarchaeote 'Gulf of Maine' | species | silva:AGBE01001967,ncbi:1089683 | NaN | sibling_higher | NaN |
| 8 | 5208050 | 4795965.0 | uncultured marine archaeon DCM858 | species | silva:AF121992,ncbi:105567 | NaN | sibling_higher | NaN |
| 9 | 5205092 | 4795965.0 | uncultured marine group I thaumarchaeote | species | silva:JF715361,ncbi:360837 | NaN | sibling_higher | NaN |
| 10 | 5205072 | 4795965.0 | uncultured Nitrosopumilaceae archaeon | species | silva:JN591993,ncbi:1118069 | NaN | sibling_higher | NaN |
| 11 | 5208765 | 4795965.0 | uncultured marine archaeon DCM874 | species | silva:AF122001,ncbi:105576 | NaN | sibling_higher | NaN |
| 12 | 179705 | 4795965.0 | Cenarchaeales | order | silva:AY192631/#4,ncbi:205948,worms:573555,irm... | NaN | NaN | NaN |
| 13 | 189165 | 179705.0 | Cenarchaeaceae | family | silva:AY192631/#5,ncbi:205957,worms:573556,gbi... | NaN | NaN | NaN |
| 14 | 888219 | 189165.0 | Cenarchaeum | genus | silva:AY192631/#6,ncbi:46769,worms:573557,gbif... | NaN | NaN | NaN |
| 15 | 5207306 | 888219.0 | Thermoplasmatales archaeon Gpl | species | silva:JN881616,ncbi:261391 | NaN | NaN | NaN |
| 16 | 376618 | 888219.0 | Cenarchaeum symbiosum A | no rank - terminal | silva:DQ397549,ncbi:414004 | NaN | NaN | NaN |
| 17 | 4796244 | 888219.0 | crenarchaeote symbiont of Axinella sp. | species | silva:AF421159,ncbi:173517 | NaN | NaN | NaN |
| 18 | 4796252 | 888219.0 | crenarchaeote symbiont of Axinella verrucosa | species | silva:AF420237,ncbi:171716 | NaN | NaN | NaN |
| 19 | 5204995 | 888219.0 | uncultured Cenarchaeaceae thaumarchaeote | species | silva:DQ299278,ncbi:375545 | NaN | NaN | NaN |
| 20 | 363497 | 888219.0 | Cenarchaeum symbiosum | species | silva:AF083072,ncbi:46770,worms:573558,gbif:59... | NaN | NaN | NaN |
| 21 | 376617 | 363497.0 | Cenarchaeum symbiosum B | no rank - terminal | ncbi:414005 | NaN | infraspecific | NaN |
| 22 | 5204996 | 888219.0 | Cenarchaeum environmental samples | no rank - terminal | ncbi:355925 | NaN | was_container | NaN |
| 23 | 5204994 | 189165.0 | Cenarchaeaceae environmental samples | no rank - terminal | ncbi:375544 | NaN | was_container | NaN |
| 24 | 5204998 | 179705.0 | Cenarchaeales environmental samples | no rank - terminal | ncbi:260466 | NaN | was_container | NaN |
| 25 | 5204999 | 179705.0 | uncultured Cenarchaeales thaumarchaeote | species | ncbi:260467 | NaN | sibling_higher,not_otu | NaN |
| 26 | 5205398 | 4795965.0 | uncultured crenarchaeote ODPB-A18 | species | silva:AF121098,ncbi:95930 | NaN | sibling_higher | NaN |
| 27 | 5205625 | 4795965.0 | uncultured crenarchaeote ODPB-A3 | species | silva:AF121093,ncbi:95925 | NaN | sibling_higher | NaN |
| 28 | 5205019 | 4795965.0 | uncultured marine crenarchaeote KM3-86-C1 | species | silva:EU686625,ncbi:526685 | NaN | sibling_higher | NaN |
| 29 | 5205058 | 4795965.0 | uncultured Nitrosopumilales archaeon | species | silva:EF069380,ncbi:171534 | NaN | sibling_higher | NaN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 969 | 5205123 | 102415.0 | thaumarchaeote enrichment culture clone Ec.FBa... | species | ncbi:1238031 | NaN | environmental | NaN |
| 970 | 5205150 | 102415.0 | thaumarchaeote enrichment culture clone Ec.FBa... | species | ncbi:1237975 | NaN | environmental | NaN |
| 971 | 5571747 | 102415.0 | uncultured marine thaumarchaeote KM3_158_B05 | species | ncbi:1456024 | NaN | environmental,not_otu | NaN |
| 972 | 5571649 | 102415.0 | uncultured marine thaumarchaeote KM3_84_D12 | species | ncbi:1456311 | NaN | environmental,not_otu | NaN |
| 973 | 5572006 | 102415.0 | uncultured marine thaumarchaeote AD1000_60_A11 | species | ncbi:1455927 | NaN | environmental,not_otu | NaN |
| 974 | 5205251 | 102415.0 | thaumarchaeote enrichment culture clone Ec.FBa... | species | ncbi:1237979 | NaN | environmental | NaN |
| 975 | 5571794 | 102415.0 | uncultured marine thaumarchaeote KM3_201_H03 | species | ncbi:1456096 | NaN | environmental,not_otu | NaN |
| 976 | 5571869 | 102415.0 | uncultured marine thaumarchaeote KM3_82_D05 | species | ncbi:1456304 | NaN | environmental,not_otu | NaN |
| 977 | 5571712 | 102415.0 | uncultured marine thaumarchaeote AD1000_24_H07 | species | ncbi:1455902 | NaN | environmental,not_otu | NaN |
| 978 | 5571961 | 102415.0 | uncultured marine thaumarchaeote KM3_75_F06 | species | ncbi:1456280 | NaN | environmental,not_otu | NaN |
| 979 | 5571926 | 102415.0 | uncultured marine thaumarchaeote KM3_71_E12 | species | ncbi:1456258 | NaN | environmental,not_otu | NaN |
| 980 | 5205222 | 102415.0 | thaumarchaeote enrichment culture clone Ec.MTa... | species | ncbi:1238100 | NaN | environmental | NaN |
| 981 | 5571900 | 102415.0 | uncultured marine thaumarchaeote KM3_06_B06 | species | ncbi:1455974 | NaN | environmental,not_otu | NaN |
| 982 | 5205166 | 102415.0 | thaumarchaeote enrichment culture clone Ec.FBb... | species | ncbi:1238069 | NaN | environmental | NaN |
| 983 | 5571740 | 102415.0 | uncultured marine thaumarchaeote KM3_193_A03 | species | ncbi:1456081 | NaN | environmental,not_otu | NaN |
| 984 | 5205235 | 102415.0 | thaumarchaeote enrichment culture clone Ec.MTa... | species | ncbi:1238126 | NaN | environmental | NaN |
| 985 | 5571679 | 102415.0 | uncultured marine thaumarchaeote KM3_23_E01 | species | ncbi:1456099 | NaN | environmental,not_otu | NaN |
| 986 | 5571865 | 102415.0 | uncultured marine thaumarchaeote SAT1000_48_A08 | species | ncbi:1456414 | NaN | environmental,not_otu | NaN |
| 987 | 5571850 | 102415.0 | uncultured marine thaumarchaeote SAT1000_10_G06 | species | ncbi:1456374 | NaN | environmental,not_otu | NaN |
| 988 | 5571611 | 102415.0 | uncultured marine thaumarchaeote AD1000_26_G12 | species | ncbi:1455904 | NaN | environmental,not_otu | NaN |
| 989 | 5572016 | 102415.0 | uncultured marine thaumarchaeote KM3_65_D04 | species | ncbi:1456224 | NaN | environmental,not_otu | NaN |
| 990 | 5571969 | 102415.0 | uncultured marine thaumarchaeote KM3_186_C08 | species | ncbi:1456070 | NaN | environmental,not_otu | NaN |
| 991 | 5571667 | 102415.0 | uncultured marine thaumarchaeote KM3_41_H02 | species | ncbi:1456146 | NaN | environmental,not_otu | NaN |
| 992 | 5571890 | 102415.0 | uncultured marine thaumarchaeote KM3_52_F05 | species | ncbi:1456177 | NaN | environmental,not_otu | NaN |
| 993 | 5571807 | 102415.0 | uncultured marine thaumarchaeote AD1000_54_F09 | species | ncbi:1455926 | NaN | environmental,not_otu | NaN |
| 994 | 5571591 | 102415.0 | uncultured marine thaumarchaeote KM3_175_A05 | species | ncbi:1456051 | NaN | environmental,not_otu | NaN |
| 995 | 5571756 | 102415.0 | uncultured marine thaumarchaeote KM3_46_E07 | species | ncbi:1456159 | NaN | environmental,not_otu | NaN |
| 996 | 5571888 | 102415.0 | uncultured marine thaumarchaeote KM3_02_A10 | species | ncbi:1455955 | NaN | environmental,not_otu | NaN |
| 997 | 5205131 | 102415.0 | thaumarchaeote enrichment culture clone Ec.FBa... | species | ncbi:1238015 | NaN | environmental | NaN |
| 998 | 5572032 | 102415.0 | uncultured marine thaumarchaeote KM3_53_B02 | species | ncbi:1456180 | NaN | environmental,not_otu | NaN |
999 rows × 8 columns
The description of the entries is given here: https://github.com/OpenTreeOfLife/reference-taxonomy/wiki/Interim-taxonomy-file-format
Explore the table¶
tree_of_life.columns
Index(['uid', 'parent_uid', 'name', 'rank', 'sourceinfo', 'uniqname', 'flags',
'Unnamed: 7'],
dtype='object')
Let us drop some columns.
tree_of_life = tree_of_life.drop(columns=['sourceinfo', 'uniqname', 'flags','Unnamed: 7'])
tree_of_life.head()
| uid | parent_uid | name | rank | |
|---|---|---|---|---|
| 0 | 805080 | NaN | life | no rank |
| 1 | 93302 | 805080.0 | cellular organisms | no rank |
| 2 | 996421 | 93302.0 | Archaea | domain |
| 3 | 5246114 | 996421.0 | Marine Hydrothermal Vent Group 1(MHVG-1) | no rank - terminal |
| 4 | 102415 | 996421.0 | Thaumarchaeota | phylum |
Pandas infered the type of values inside each column (int, float, string and string). The parent_uid column has float values because there was a missing value, converted to NaN
print(tree_of_life['uid'].dtype, tree_of_life.parent_uid.dtype)
int64 float64
How to access individual values.
tree_of_life.iloc[0, 2]
'life'
tree_of_life.loc[0, 'name']
'life'
Exercise: Guess the output of the below line.
# tree_of_life.uid[0] == tree_of_life.parent_uid[1]
Ordering the data.
tree_of_life.sort_values(by='name').head()
| uid | parent_uid | name | rank | |
|---|---|---|---|---|
| 297 | 5246638 | 102415.0 | AB64A-17 | no rank - terminal |
| 293 | 5246632 | 102415.0 | AK31 | no rank - terminal |
| 298 | 5246637 | 102415.0 | AK56 | no rank - terminal |
| 202 | 5246635 | 102415.0 | AK59 | no rank - terminal |
| 204 | 5246636 | 102415.0 | AK8 | no rank - terminal |
Operation on the columns¶
Unique values, useful for categories:
tree_of_life['rank'].unique()
array(['no rank', 'domain', 'no rank - terminal', 'phylum', 'species',
'order', 'family', 'genus', 'class'], dtype=object)
Selecting only one category.
tree_of_life[tree_of_life['rank'] == 'species'].head()
| uid | parent_uid | name | rank | |
|---|---|---|---|---|
| 7 | 5205649 | 4795965.0 | uncultured marine crenarchaeote 'Gulf of Maine' | species |
| 8 | 5208050 | 4795965.0 | uncultured marine archaeon DCM858 | species |
| 9 | 5205092 | 4795965.0 | uncultured marine group I thaumarchaeote | species |
| 10 | 5205072 | 4795965.0 | uncultured Nitrosopumilaceae archaeon | species |
| 11 | 5208765 | 4795965.0 | uncultured marine archaeon DCM874 | species |
How many species do we have?
len(tree_of_life[tree_of_life['rank'] == 'species'])
912
tree_of_life['rank'].value_counts()
species 912 no rank - terminal 58 no rank 12 genus 8 order 3 family 3 domain 1 phylum 1 class 1 Name: rank, dtype: int64
Building the graph¶
Let us build the adjacency matrix of the graph. For that we need to reorganize the data. First we separate the nodes and their properties from the edges.
nodes = tree_of_life[['uid', 'name','rank']]
edges = tree_of_life[['uid', 'parent_uid']]
When using an adjacency matrix, nodes are indexed by their row or column number and not by a uid. Let us create a new index for the nodes.
# Create a column for node index.
nodes.reset_index(level=0, inplace=True)
nodes = nodes.rename(columns={'index':'node_idx'})
nodes.head()
| node_idx | uid | name | rank | |
|---|---|---|---|---|
| 0 | 0 | 805080 | life | no rank |
| 1 | 1 | 93302 | cellular organisms | no rank |
| 2 | 2 | 996421 | Archaea | domain |
| 3 | 3 | 5246114 | Marine Hydrothermal Vent Group 1(MHVG-1) | no rank - terminal |
| 4 | 4 | 102415 | Thaumarchaeota | phylum |
# Create a conversion table from uid to node index.
uid2idx = nodes[['node_idx', 'uid']]
uid2idx = uid2idx.set_index('uid')
uid2idx.head()
| node_idx | |
|---|---|
| uid | |
| 805080 | 0 |
| 93302 | 1 |
| 996421 | 2 |
| 5246114 | 3 |
| 102415 | 4 |
edges.head()
| uid | parent_uid | |
|---|---|---|
| 0 | 805080 | NaN |
| 1 | 93302 | 805080.0 |
| 2 | 996421 | 93302.0 |
| 3 | 5246114 | 996421.0 |
| 4 | 102415 | 996421.0 |
Now we are ready to use yet another powerful function of Pandas. Those familiar with SQL will recognize it: the join function.
# Add a new column, matching the uid with the node_idx.
edges = edges.join(uid2idx, on='uid')
# Do the same with the parent_uid.
edges = edges.join(uid2idx, on='parent_uid', rsuffix='_parent')
# Drop the uids.
edges = edges.drop(columns=['uid','parent_uid'])
edges.head()
| node_idx | node_idx_parent | |
|---|---|---|
| 0 | 0 | NaN |
| 1 | 1 | 0.0 |
| 2 | 2 | 1.0 |
| 3 | 3 | 2.0 |
| 4 | 4 | 2.0 |
The above table is a list of edges connecting nodes and their parents.
Building the (weighted) adjacency matrix¶
We will use numpy to build this matrix. Note that we don't have edge weights here, so our graph is going to be unweighted.
n_nodes = len(nodes)
adjacency = np.zeros((n_nodes, n_nodes), dtype=int)
for idx, row in edges.iterrows():
if np.isnan(row.node_idx_parent):
continue
i, j = int(row.node_idx), int(row.node_idx_parent)
adjacency[i, j] = 1
adjacency[j, i] = 1
adjacency[:15, :15]
array([[0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
[1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 1, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 1, 0, 0, 1, 1, 1, 1, 1, 1, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0],
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1],
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0]])
Congratulations, you have built the adjacency matrix!
Graph visualization¶
To conclude, let us visualize the graph. We will use the python module networkx.
# A simple command to create the graph from the adjacency matrix.
graph = nx.from_numpy_array(adjacency)
In addition, let us add some attributes to the nodes:
node_props = nodes.to_dict()
for key in node_props:
# print(key, node_props[key])
nx.set_node_attributes(graph, node_props[key], key)
Let us check if it is correctly recorded:
graph.node[1]
{'node_idx': 1, 'uid': 93302, 'name': 'cellular organisms', 'rank': 'no rank'}
Draw the graph with two different layout algorithms.
nx.draw_spectral(graph)
/home/michael/.conda/envs/ntds_2018/lib/python3.6/site-packages/networkx/drawing/nx_pylab.py:611: MatplotlibDeprecationWarning: isinstance(..., numbers.Number) if cb.is_numlike(alpha):
nx.draw_spring(graph)
/home/michael/.conda/envs/ntds_2018/lib/python3.6/site-packages/networkx/drawing/nx_pylab.py:611: MatplotlibDeprecationWarning: isinstance(..., numbers.Number) if cb.is_numlike(alpha):
Save the graph to disk in the gexf format, readable by gephi and other tools that manipulate graphs. You may now explore the graph using gephi and compare the visualizations.
nx.write_gexf(graph, 'tree_of_life.gexf')