
Stacking Guide¶
import numpy as np
np.random.seed(0)
import kts
from kts import *
train = kts.load('train')
test = kts.load('test')
stl.stack¶
To stack models, stl.stack is used:
stl.stack
stack(experiment_id, noise_level, random_state)
For indices used for fitting the experiment returns OOF predictions.
For unseen indices returns predictions obtained via experiment.predict().
If specified, then uniformly distributed value from range [-noise_level/2, noise_level/2]
is added to each prediction.
>>> stl.stack('ABCDEF') >>> stl.stack('ABCDEF', noise_level=0.3, random_state=42) >>> stl.concat([stl.stack('ABCDEF'), stl.stack('GHIJKL')])
In case if we pass a train set slice, it just returns OOF predictions:
Note that it cannot be used in parallel features.
@preview(train, 10, parallel=False)
def preview_stack(df):
return stl.stack('KPBVAI')(df)
| KPBVAI | |
|---|---|
| PassengerId | |
| 1 | 0.109578 |
| 2 | 0.985013 |
| 3 | 0.635773 |
| 4 | 0.906962 |
| 5 | 0.132054 |
| 6 | 0.124416 |
| 7 | 0.322831 |
| 8 | 0.599516 |
| 9 | 0.520683 |
| 10 | 0.922890 |
But for test set, inference is run:
@preview(test, 10, parallel=False)
def preview_stack(df):
return stl.stack('KPBVAI')(df)
| KPBVAI | |
|---|---|
| PassengerId | |
| 892 | 0.150248 |
| 893 | 0.589290 |
| 894 | 0.107736 |
| 895 | 0.229384 |
| 896 | 0.756447 |
| 897 | 0.191558 |
| 898 | 0.758887 |
| 899 | 0.271479 |
| 900 | 0.785848 |
| 901 | 0.282044 |
Anti-overfitting¶
KTS provides two basic ways to prevent overfitting during stacking.
Noise¶
First of them is adding random uniform noise to first-level model predictions during training stage:
@preview(train, 4, 4, parallel=False)
def preview_stack(df):
return stl.stack('KPBVAI', noise_level=0.1, random_state=None)(df)
| KPBVAI | |
|---|---|
| PassengerId | |
| 1 | 0.112647 |
| 2 | 0.977061 |
| 3 | 0.683239 |
| 4 | 0.921687 |
| KPBVAI | |
|---|---|
| PassengerId | |
| 1 | 0.077990 |
| 2 | 1.005335 |
| 3 | 0.609979 |
| 4 | 0.933971 |
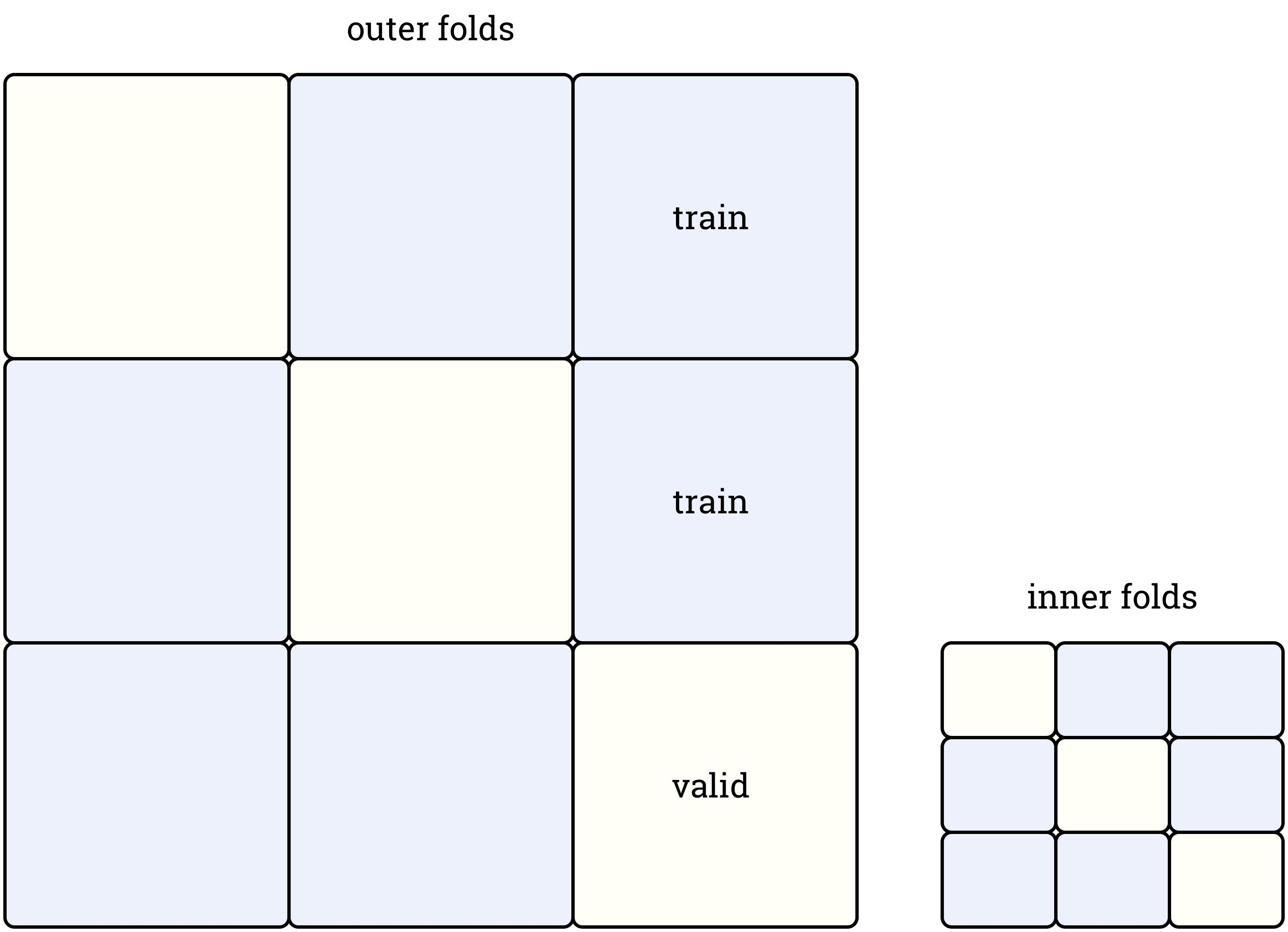
Refiner¶
The second available option is a special splitter called Refiner, which splits each fold of an outer splitter using inner splitter. It allows to train a second-level model without even indirect leaks, as in this case each second-level model is trained using validation set of the corresponding first-level model.
from kts.validation.split import Refiner
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import roc_auc_score
outer_skf = StratifiedKFold(5, True, 42) # splitter used to train the first-level model
inner_skf = StratifiedKFold(3, True, 42) # splitter to be used to split its folds
refiner = Refiner(outer_skf, inner_skf)
val_stack = Validator(refiner, roc_auc_score)
fs = FeatureSet([stl.stack('KPBVAI'), stl.stack('FYCMDA'), tfidf()],
train_frame=train,
targets='Survived')
from kts.models.binary import *
model = LogisticRegression(solver='lbfgs', C=11)
val_stack.score(model, fs)
{'score': 0.8348066445892532, 'id': 'CBUQLG'}
Stacked experiments behave exactly as usual experiments:
lb.CBUQLG.predict(test)[:10]
array([0.09516588, 0.46953291, 0.09342506, 0.15601523, 0.7111576 ,
0.11536574, 0.56980286, 0.16907218, 0.66821078, 0.13896083])
lb.CBUQLG.feature_importances(estimator=Permutation(train, n_iters=10))
lb.CBUQLG.feature_importances(estimator=PermutationBlind(test, n_iters=20))
Deep Stacking¶
As stl.stack is no more than a usual feature constructor, you can build as complex stackings as you want just by adding it to feature sets.
Let's write a five-level stacking with resudual connections. In this demo we don't care about overfitting and model performance and just show that:
- Stacking is as easy as adding
stl.stack(id)to feature set - Stacking inference is no different from ordinary experiments
- In case if two or more next-level models need predictions from model A, model A will still be run only once
skf = StratifiedKFold(5, True, 42)
val = Validator(skf, roc_auc_score)
current_features = [tfidf(), num_aggs('SibSp'), num_aggs('Parch')]
for i in range(5):
model = RandomForestClassifier(n_estimators=50)
fs = FeatureSet(current_features, train_frame=train, targets='Survived')
summary = val.score(model, fs, leaderboard='deepstack')
current_features.append(stl.stack(summary['id']))
lbs.deepstack
So, BBFXPT is a fifth-level model.
lb.BBFXPT.predict(test.head(21))
array([0.07853333, 0.848 , 0.065 , 0.11357619, 0.508 ,
0.008 , 0.68731746, 0.37766667, 0.42224762, 0.24766667,
0.14748889, 0.14460952, 0.58071429, 0.33666667, 0.632 ,
0.768 , 0.542 , 0.0744 , 0.50438095, 0.46020952,
0.27 ])
lb.BBFXPT.feature_importances()