Overview¶
[ Back to top ]
What is Dask?¶
Dask is a flexible, open source library for parallel computing in Python
GitHub: https://github.com/dask/dask
Documentation: https://docs.dask.org
Scales the existing Python ecosystem
Why Dask?¶
Enables parallel and larger-than-memory computations
Uses familiar APIs you're used to from projects like NumPy, Pandas, and scikit-learn
Allows you to scale existing workflows with minimal code changes
Dask works on your laptop, but also scales out to large clusters
Offers great built-in diagnosic tools
Components of Dask¶
Task Graphs¶
def inc(i):
return i + 1
def add(a, b):
return a + b
a, b = 1, 12
c = inc(a)
d = inc(b)
output = add(c, d)
print(f'output = {output}')
This computation can be encoded in the following task graph:

Graph of inter-related tasks with dependencies between them
Circular nodes in the graph are Python function calls
Square nodes are Python objects that are created by one task as output and can be used as inputs in another task
High Level Collections¶
[ Back to top ]
Dask Arrays¶
Dask arrays are chunked, n-dimensional arrays
Can think of a Dask array as a collection of NumPy
ndarrayarraysDask arrays implement a large subset of the NumPy API using blocked algorithms
For many purposes Dask arrays can serve as drop-in replacements for NumPy arrays
import numpy as np
import dask.array as da
arr_np = np.arange(1, 50, 3)
arr_np
We can create a Dask array in a similar manner, but need to specify a chunks argument to tell Dask how to break up the underlying array into chunks.
arr_da = da.arange(1, 50, 3, chunks=5) # Each chunk is 5 items long
type(arr_da)
arr_da # Dask arrays have nice HTML output in Jupyter notebooks
Dask arrays look and feel like NumPy arrays. For example, they have dtype and shape attributes
print(arr_da.dtype)
print(arr_da.shape)
Dask arrays are lazily evaluated. The result from a computation isn't computed until you ask for it. Instead, a Dask task graph for the computation is produced. You can visualize the task graph using the visualize() method.
arr_da.visualize()
To compute a task graph call the compute() method
arr_da.compute() # We'll go into more detail about .compute() later on
The result of this computation is a fimilar NumPy ndarray
type(arr_da.compute())
Dask arrays support a large portion of the NumPy interface:
Arithmetic and scalar mathematics:
+,*,exp,log, ...Reductions along axes:
sum(),mean(),std(),sum(axis=0), ...Tensor contractions / dot products / matrix multiply:
tensordotAxis reordering / transpose:
transposeSlicing:
x[:100, 500:100:-2]Fancy indexing along single axes with lists or numpy arrays:
x[:, [10, 1, 5]]Array protocols like
__array__and__array_ufunc__Some linear algebra:
svd,qr,solve,solve_triangular,lstsq, ......
See the Dask array API docs for full details about what portion of the NumPy API is implemented for Dask arrays.
Blocked Algorithms¶
Dask arrays are implemented using blocked algorithms. These algorithms break up a computation on a large array into many computations on smaller peices of the array. This minimizes the memory load (amount of RAM) of computations and allows for working with larger-than-memory datasets in parallel.
x = da.random.random(20, chunks=5)
x
result = x.sum()
result.visualize()
result.compute()
We can build more complex computations using the familiar NumPy operations we're used to.
x = da.random.random(size=(1_000, 1_000), chunks=(250, 500))
x
result = (x + x.T).sum(axis=0).mean()
result.visualize()
result.compute()
We can even perform computations on larger-than-memory arrays!
x = da.random.random(size=(40_000, 40_000), chunks=(2_000, 2_000))
x.nbytes / 1e9 # Size of array in gigabytes
result = (x + x.T).sum(axis=0).mean()
from dask.diagnostics import ProgressBar
with ProgressBar():
result.compute()
Note: Dask can be used to scale other array-like libraries that support the NumPy ndarray interface. For example, pydata/sparse for sparse arrays or CuPy for GPU-accelerated arrays.
Dask DataFrames¶
Pandas is great for dealing with tabular datasets that can fit in memory on a single machine. Dask becomes useful when dealing with larger-than-memory datasets. We saw that a Dask arrays are composed of many NumPy arrays, chunked along one or more dimensions. It's similar for Dask DataFrames: a Dask DataFrame is composed of many Pandas DataFrames and the partitioning happens only along the index.
Dask DataFrames are a collection of Pandas DataFrames
Dask DataFrames implement a large subset of the Pandas API
Backed by blocked algorithms that allow for parallel and out of core computation
For many purposes Dask DataFrames can serve as drop-in replacements for Pandas DataFrames
import pandas as pd
import dask.dataframe as dd
Dask DataFrames support many of the same data I/O methods as Pandas. For example,
read_csv\to_csvread_hdf\to_hdfread_json\to_jsonread_parquet\to_parquet
Can specify a chunksize argument to set the number of rows per partition
ddf = dd.read_hdf('example_data.hdf', key='dataframe', chunksize=25)
ddf
The npartitions attribute tells us how many Pandas DataFrames make up our Dask DataFrame
ddf.npartitions
Like Dask arrays, Dask DataFrames are lazily evaluated. Here, the dd.read_hdf function wraps several calls to pd.read_hdf, once for each partition of the Dask DataFrame.
ddf.visualize()
Dask DataFrames cover a well-used portion of the Pandas API:
Elementwise operations:
df.x+df.y,df * dfRow-wise selections:
df[df.x > 0]Loc:
df.loc[4.0:10.5]Common aggregations:
df.x.max(),df.max()Is in:
df[df.x.isin([1, 2, 3])]Datetime/string accessors:
df.timestamp.monthFroupby-aggregate (with common aggregations):
df.groupby(df.x).y.max(),df.groupby('x').max()...
See the Dask DataFrame API docs for full details about what portion of the Pandas API is implemented for Dask DataFrames.
col_mean = ddf['col_1'].mean()
col_mean
col_mean.visualize()
col_mean.compute()
Dask Delayed¶
[ Back to top ]
Sometimes problems don’t fit nicely into one of the high-level collections like Dask arrays or Dask DataFrames. In these cases, you can parallelize custom algorithms using the lower-level Dask delayed interface. This allows one to manually create task graphs with a light annotation of normal Python code.
import time
import random
def inc(x):
time.sleep(random.random())
return x + 1
def double(x):
time.sleep(random.random())
return 2 * x
def add(x, y):
time.sleep(random.random())
return x + y
%%time
data = [1, 2, 3, 4]
output = []
for x in data:
a = inc(x)
b = double(x)
c = add(a, b)
output.append(c)
total = sum(output)
Dask delayed wraps function calls and delays their execution. delayed functions record what we want to compute (a function and input parameters) as a task in a graph that we’ll run later on parallel hardware by calling compute.
from dask import delayed
@delayed
def lazy_inc(x):
time.sleep(random.random())
return x + 1
lazy_inc
inc_output = lazy_inc(3) # lazily evaluate inc(3)
inc_output
inc_output.compute()
Using delayed functions, we can build up a task graph for the particular computation we want to perform
double_inc_output = lazy_inc(inc_output)
double_inc_output
double_inc_output.visualize()
double_inc_output.compute()
We can use delayed to make our previous example computation lazy by wrapping all the function calls with delayed
import time
import random
@delayed
def inc(x):
time.sleep(random.random())
return x + 1
@delayed
def double(x):
time.sleep(random.random())
return 2 * x
@delayed
def add(x, y):
time.sleep(random.random())
return x + y
%%time
data = [1, 2, 3, 4]
output = []
for x in data:
a = inc(x)
b = double(x)
c = add(a, b)
output.append(c)
total = delayed(sum)(output)
total
total.visualize()
%%time
total.compute()
I highly recommend checking out the Dask delayed best practices page to avoid some common pitfalls when using delayed.
Schedulers¶
[ Back to top ]
High-level collections like Dask arrays and Dask DataFrames, as well as the low-level dask.delayed interface build up task graphs for a computation. After these graphs are generated, they need to be executed (potentially in parallel). This is the job of a task scheduler. Different task schedulers exist within Dask. Each will consume a task graph and compute the same result, but with different performance characteristics.
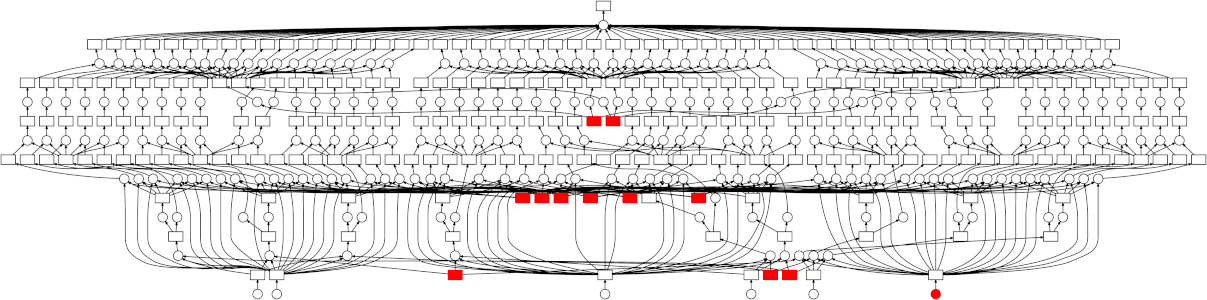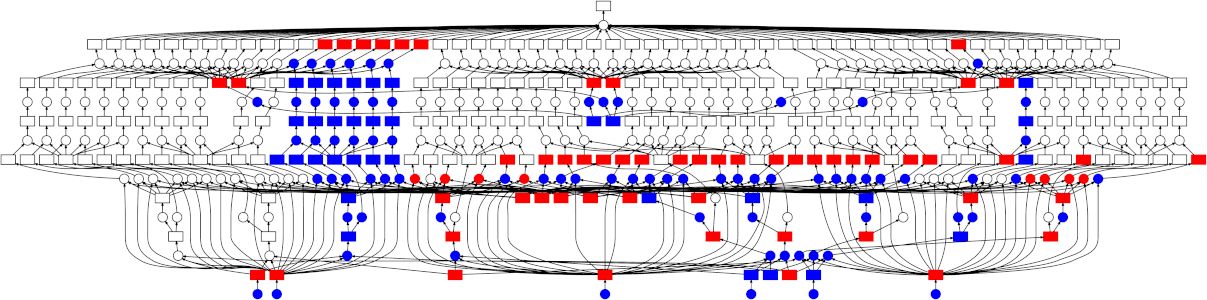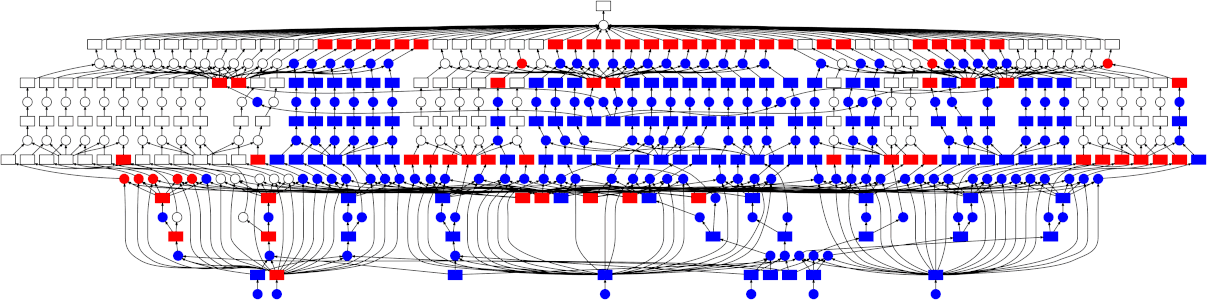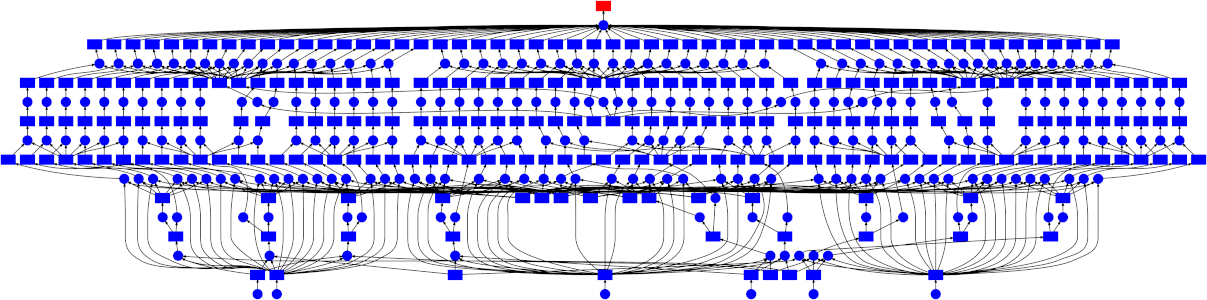
Dask has two different classes of schedulers: single-machine schedulers and a distributed scheduler.
Single Machine Schedulers¶
[ Back to top ]
Single machine schedulers provide basic features on a local process or thread pool and require no setup (only use the Python standard library). The different single machine schedulers Dask provides are:
'threads': The threaded scheduler executes computations with a localmultiprocessing.pool.ThreadPool. The threaded scheduler is the default choice for Dask arrays, Dask DataFrames, and Dask delayed.'processes': The multiprocessing scheduler executes computations with a localmultiprocessing.Pool.'single-threaded': The single-threaded synchronous scheduler executes all computations in the local thread, with no parallelism at all. This is particularly valuable for debugging and profiling, which are more difficult when using threads or processes.
You can configure which scheduler is used in a few different ways. You can set the scheduler globally by using the dask.config.set(scheduler=) command
import dask
dask.config.set(scheduler='threads')
total.compute(); # Will use the multi-threading scheduler
or use it as a context manager to set the scheduler for a block of code
with dask.config.set(scheduler='processes'):
total.compute() # Will use the multi-processing scheduler
or even within a single compute call
total.compute(scheduler='threads'); # Will use the multi-threading scheduler
The num_workers argument is used to specify the number of threads or processes to use
total.compute(scheduler='threads', num_workers=4);
Distributed Scheduler¶
[ Back to top ]
Despite having "distributed" in it's name, the distributed scheduler works well on both single and multiple machines. Think of it as the "advanced scheduler".
The Dask distributed cluster is composed of a single centralized scheduler and one or more worker processes. A Client object is used as the user-facing entry point to interact with the cluster.
Deploying a remote Dask cluster involves some additional setup. There are several projects for easily deploying a Dask cluster on commonly used computing resources:
- Dask-Kubernetes for deploying Dask using native Kubernetes APIs
- Dask-Yarn for deploying Dask on YARN clusters
- Dask-MPI for deploying Dask on existing MPI environments
- Dask-Jobqueue for deploying Dask on job queuing systems (e.g. PBS, Slurm, etc.)
Setting up the distributed scheduler locally just involves creating a Client object, which lets you interact with the "cluster" (local threads or processes on your machine).
from dask.distributed import Client
client = Client(threads_per_worker=4, n_workers=1)
client
Note: when we create a distributed scheduler Client, by default it registers itself as the default Dask scheduler. All .compute() calls will automatically start using the distributed scheduler unless otherwise told to use a different scheduler.
The distributed scheduler has many features:
See the Dask distributed documentation for full details about all the distributed scheduler features.
For this talk, I'd like to highlight two of the distributed scheduler features: real time diagnostics and the futures interface.
x = da.ones((20_000, 20_000), chunks=(400, 400))
result = (x + x.T).sum(axis=0).mean()
result.compute()
Futures interface¶
[ Back to top ]
The Dask distributed scheduler implements a superset of Python's concurrent.futures interface that allows for finer control and asynchronous computation.
import time
import random
def inc(x):
time.sleep(random.random())
return x + 1
def double(x):
time.sleep(random.random())
return 2 * x
def add(x, y):
time.sleep(random.random())
return x + y
We can run these functions locally
inc(1)
Or we can submit them to run remotely on a Dask worker node
future = client.submit(inc, 1)
future
The submit function sends a function and arguments to the distributed scheduler for processing. It returns a Future object that refer to remote data on the cluster. The Future returns immediately while the computations run remotely in the background. There is no blocking of the local Python session.
If you wait a moment, and then check on the future again, you'll see that it has finished.
future
Once the computation for a Future is complete, you can retrieve the result using the .result() method
future.result()
Specifying task dependencies¶
Much like the delayed interface, we can submit tasks based on other futures. This will create a dependency between the inputs and outputs. Dask will track the execution of all tasks and ensure that downstream tasks are run at the proper time and place and with the proper data.
x = client.submit(inc, 1)
y = client.submit(double, 2)
z = client.submit(add, x, y)
z
z.result()
As an example, we can submit many tasks that depend on each other in a for-loop
%%time
zs = []
for i in range(64):
x = client.submit(inc, i) # x = inc(i)
y = client.submit(double, x) # y = inc(x)
z = client.submit(add, x, y) # z = inc(y)
zs.append(z)
total = client.submit(sum, zs)
total
total.result()
Custom computation: Tree summation¶
As an example of a non-trivial algorithm, consider the classic tree reduction. We accomplish this with a nested for-loop and a bit of normal Python logic.
finish total single output
^ / \
| c1 c2 neighbors merge
| / \ / \
| b1 b2 b3 b4 neighbors merge
^ / \ / \ / \ / \
start a1 a2 a3 a4 a5 a6 a7 a8 many inputs
We can easily scale our distributed cluster up or down depending on our needs
client.cluster.scale(3) # ask for 3 4-thread workers
L = zs
while len(L) > 1:
new_L = []
for i in range(0, len(L), 2):
future = client.submit(add, L[i], L[i + 1]) # add neighbors
new_L.append(future)
L = new_L # swap old list for new
del future, L, new_L, total
Building a computation dynamically¶
We can even dynamically submit tasks based on the output of other tasks. This gives more flexibility in situations where the computations may evolve over time.
For this, we can use operations like as_completed(), which returns futures in the order in which they complete.
from dask.distributed import as_completed
zs = client.map(inc, zs)
seq = as_completed(zs)
while seq.count() > 2: # at least two futures left
a = next(seq)
b = next(seq)
new = client.submit(add, a, b) # add them together
seq.add(new) # add new future back into loop
This was a brief demo of the distributed scheduler. It's has lots of other cool features not touched on here. For more information, check out the Distributed documentation.
Scalable Machine Learning with Dask-ML¶
[ Back to top ]
Dask-ML is a Python library for scalable machine learning in Python. Build on top of Dask collections and supports the scikit-learn API.
Three different approaches are taken to scaling modern machine learning algorithms:
Parallelize scikit-learn directly
Reimplement scalable algorithms with Dask arrays
Partner with other distributed libraries (like XGBoost and TensorFlow)
from dask_ml.datasets import make_classification
from dask_ml.model_selection import train_test_split
from dask_ml.metrics import accuracy_score
from dask_ml.linear_model import LogisticRegression
X, y = make_classification(n_samples=1000,
n_features=2,
n_classes=2,
random_state=2,
chunks=10)
X
X_train, X_test, y_train, y_test = train_test_split(X, y,
test_size=0.3,
random_state=2)
clf = LogisticRegression(max_iter=2)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
accuracy_score(y_test, y_pred)
Additional Resources¶
[ Back to top ]
Dask links:
GitHub repository: https://github.com/dask/dask
Documentation: https://docs.dask.org
Dask examples repository: https://github.com/dask/dask-examples
There are lots of great Dask tutorial from various conference on YouTube. For example:
If you have a Dask usage questions, please ask it on Stack Overflow with the #dask tag. Dask developers monitor this tag and will answer questions.
If you run into a bug, feel free to file a report on the Dask GitHub issue tracker.