Exploring high dimensional data¶
You'll be introduced to the concept of dimensionality reduction and will learn when an why this is important. You'll learn the difference between feature selection and feature extraction and will apply both techniques for data exploration. The chapter ends with a lesson on t-SNE, a powerful feature extraction technique that will allow you to visualize a high-dimensional dataset. This is the Summary of lecture "Dimensionality Reduction in Python", via datacamp.
- toc: true
- badges: true
- comments: true
- author: Chanseok Kang
- categories: [Python, Datacamp, Machine_Learning]
- image: images/tsne_gender.png
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (10, 5)
Introduction¶
Removing features without variance¶
A sample of the Pokemon dataset has been loaded as pokemon_df. To get an idea of which features have little variance you should use the IPython Shell to calculate summary statistics on this sample. Then adjust the code to create a smaller, easier to understand, dataset.
pokemon_df = pd.read_csv('./dataset/pokemon_gen1.csv')
pokemon_df.head()
| HP | Attack | Defense | Generation | Name | Type | Legendary | |
|---|---|---|---|---|---|---|---|
| 0 | 45 | 49 | 49 | 1 | Bulbasaur | Grass | False |
| 1 | 60 | 62 | 63 | 1 | Ivysaur | Grass | False |
| 2 | 80 | 82 | 83 | 1 | Venusaur | Grass | False |
| 3 | 80 | 100 | 123 | 1 | VenusaurMega Venusaur | Grass | False |
| 4 | 39 | 52 | 43 | 1 | Charmander | Fire | False |
pokemon_df.describe()
| HP | Attack | Defense | Generation | |
|---|---|---|---|---|
| count | 160.00000 | 160.00000 | 160.000000 | 160.0 |
| mean | 64.61250 | 74.98125 | 70.175000 | 1.0 |
| std | 27.92127 | 29.18009 | 28.883533 | 0.0 |
| min | 10.00000 | 5.00000 | 5.000000 | 1.0 |
| 25% | 45.00000 | 52.00000 | 50.000000 | 1.0 |
| 50% | 60.00000 | 71.00000 | 65.000000 | 1.0 |
| 75% | 80.00000 | 95.00000 | 85.000000 | 1.0 |
| max | 250.00000 | 155.00000 | 180.000000 | 1.0 |
# Remove the feature without variance from this list
number_cols = ['HP', 'Attack', 'Defense']
# Leave this list as is for now
non_number_cols = ['Name', 'Type', 'Legendary']
# Sub-select by combining the lists with chosen features
df_selected = pokemon_df[number_cols + non_number_cols]
# Prints the first 5 lines of the new dataframe
print(df_selected.head())
HP Attack Defense Name Type Legendary 0 45 49 49 Bulbasaur Grass False 1 60 62 63 Ivysaur Grass False 2 80 82 83 Venusaur Grass False 3 80 100 123 VenusaurMega Venusaur Grass False 4 39 52 43 Charmander Fire False
# Leave this list as is
number_cols = ['HP', 'Attack', 'Defense']
# Remove the feature without variance from this list
non_number_cols = ['Name', 'Type', ]
# Create a new dataframe by subselecting the chosen features
df_selected = pokemon_df[number_cols + non_number_cols]
# Prints the first 5 lines of the new dataframe
print(df_selected.head())
HP Attack Defense Name Type 0 45 49 49 Bulbasaur Grass 1 60 62 63 Ivysaur Grass 2 80 82 83 Venusaur Grass 3 80 100 123 VenusaurMega Venusaur Grass 4 39 52 43 Charmander Fire
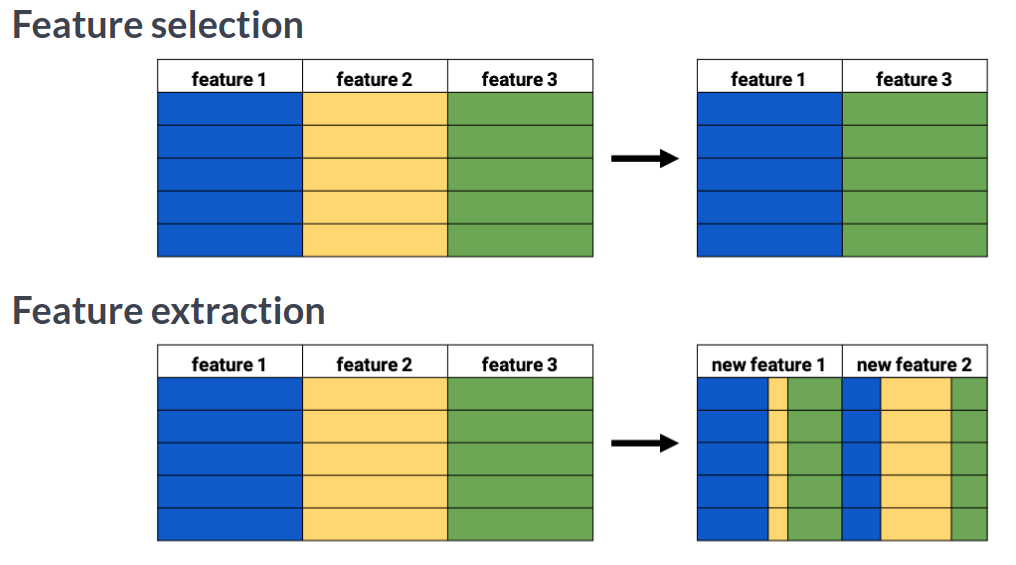
Feature selection vs feature extraction¶
- Why reduce dimensionality?
- Your dataset will:
- be less complex
- require less disk space
- require less computation time
- have lower chance of model overfitting
- Your dataset will:
Visually detecting redundant features¶
Data visualization is a crucial step in any data exploration. Let's use Seaborn to explore some samples of the US Army ANSUR body measurement dataset.
ansur_df_1 = pd.read_csv('./dataset/ansur_df_1.csv')
ansur_df_2 = pd.read_csv('./dataset/ansur_df_2.csv')
# Create a pairplot and color the points using the 'Gender' feature
sns.pairplot(ansur_df_1, hue='Gender', diag_kind='hist');
# Remove one of the redundant features
reduced_df = ansur_df_1.drop('body_height', axis=1)
# Creat a pairplot and color the points using the 'Gender' feature
sns.pairplot(reduced_df, hue='Gender');
# Create a pairplot and color the points using the 'Gender' feature
sns.pairplot(ansur_df_2, hue='Gender', diag_kind='hist');
# Remove the redundant feature
reduced_df = ansur_df_2.drop(['n_legs'], axis=1)
# Create a pairplot and color the points using the 'Gender' feature
sns.pairplot(reduced_df, hue='Gender', diag_kind='hist');
the body height (inches) and stature (meters) hold the same information in a different unit + all the individuals in the second sample have two legs.
t-SNE visualization of high-dimensional data¶
Fitting t-SNE to the ANSUR data¶
t-SNE is a great technique for visual exploration of high dimensional datasets. In this exercise, you'll apply it to the ANSUR dataset. You'll remove non-numeric columns from the pre-loaded dataset df and fit TSNE to his numeric dataset.
ansur_male = pd.read_csv('./dataset/ANSUR_II_MALE.csv')
ansur_female = pd.read_csv('./dataset/ANSUR_II_FEMALE.csv')
df = pd.concat([ansur_male, ansur_female])
from sklearn.manifold import TSNE
# Non-numeric columns in the dataset
non_numeric = ['Branch', 'Gender', 'Component', 'BMI_class', 'Height_class']
# Drop the non-numeric columns from df
df_numeric = df.drop(non_numeric, axis=1)
# Create a t-SNE model with learning rate 50
m = TSNE(learning_rate=50)
# fit and transform the t-SNE model on the numeric dataset
tsne_features = m.fit_transform(df_numeric)
print(tsne_features.shape)
(6068, 2)
t-SNE reduced the more than 90 features in the dataset to just 2 which you can now plot.
t-SNE visualisation of dimensionality¶
Time to look at the results of your hard work. In this exercise, you will visualize the output of t-SNE dimensionality reduction on the combined male and female Ansur dataset. You'll create 3 scatterplots of the 2 t-SNE features ('x' and 'y') which were added to the dataset df. In each scatterplot you'll color the points according to a different categorical variable.
df['x'] = tsne_features[:, 0]
df['y'] = tsne_features[:, 1]
# Color the points according to Army Component
sns.scatterplot(x='x', y='y', hue='Component', data=df);
# Color the points by Army Branch
sns.scatterplot(x='x', y='y', hue='Branch', data=df);
# Color the points by Gender
sns.scatterplot(x='x', y='y', hue='Gender', data=df);
There is a Male and a Female cluster. t-SNE found these gender differences in body shape without being told about them explicitly! From the second plot you learned there are more males in the Combat Arms Branch.