Back to the main index
Streamline research with psychopy_ext¶
Part of the introductory series Python for Vision Researchers brought to you by the GestaltReVision group (KU Leuven, Belgium).
In this part we introduce an advanced package, psychopy_ext, that helps you tie together the entire research cycle. It is based on the following paper:
Kubilius, J. (2014). A framework for streamlining research workflow in neuroscience and psychology. Frontiers in Neuroinformatics, 7, 52. doi:10.3389/fninf.2013.00052
Author: Jonas Kubilius
Year: 2014
Copyright: Public Domain as in CC0 (except for figures that, technically speaking, need an attribution as in CC BY because they are part of the publication mentioned above)
Contents¶
Introduction¶
Reproducible research¶
So far we've discussed how to code experiments. But research is more than just making an experiment! You have to analyze data, possibly also compare them to simulated data, present them in conferences and publish in journals. You should also nicely organize and verify your scripts. Ultimately, the goal would be to have your entire project completely reproducible, such that anybody could start from scratch and redo your experiments, regenerate your figures, posters, and papers, and directly build on your work -- this is how knowledge is accumulated and that is the whole Open Science concept that is taking over academia in recent years.
I even made a figure to illustrate that:
So that is what you might want to do. But this is what you and I do instead:
- "My data? Um, here is the folder with the first pilot data, and there are also data from this intern here, but she never finished working on it. I'll have to look more closely which data was used..."
- "I've got this analysis somewhere on an Excel spreadsheet, hold on..."
- "Here is my SPSS analysis file! Oh, you don't have SPSS? Maybe you guys could get it through your department?"
- "I've worked on this presentation for so long, and now my prof told me to run a couple more subjects. I'll have to redraw all my figures again!"
- "Here's the zip file of all scripts for this project. It's a bit messy and without any comments, hope you manage to figure it out:) Oh, the main script doesn't run? Yeah, I see, you have to comment out several bits and also change Line 157 cause there seems to be a bug..."
This is not reproducible at all. We're too often relying on:
- Too many tools that are not tied together in any way and would be hard to tie.
- Commercial software when an equally robust free and open source option is available.
- Tools that are not suited for academic work (yes, I'm looking at you, Excel -- see the Reinhart-Rogoff case).
We need better tools!¶
I do not blame researchers for relying on all these ad-hoc solutions. While in theory it would be nice to code everything from A to Z, in practice we don't have the time to play these silly games. Why we don't have the time is a topic for a separate discussion, but why we have to play these silly games is the problem of software. Simply put, we the lack tools that would seamlessly enact good coding and sharing standards. We need tools that act clever.
Technology rarely has this quality, unfortunately. If you don't agree with this, you obviously have never tried to explain a newbie how to run a Python script: "OK, now run it. I mean, open the command line... it's in... um... OK, click on the Start button, type 'cmd', hit Enter. OK, now navigate to where there script is. OK, open Windows Explorer and get the path...". This is not clever -- this is developers not caring. A smartphone that my grandfather cannot figure out is not clever -- it's pretentious.
There is a reason why people stick to spreadsheets -- they're simple, intuitive and the data is there, as opposed to being only available when you run your analysis script (Bret Victor's point). They're still stupid, of course -- have you ever tried making figures nice in Excel? -- and we rather want tools that:
- Have reasonable defaults, e.g., nice plots by default (hello, matplotlib with the 90's aesthetics)
- Require minimal user intervention, i.e., I don't want to spend hours making Python talk to R or fixing silly LaTeX bugs.
- Have an intuitive interface based on how people -- and not engineers -- think.
- Encourage good habits, just like in Python enforcing indentation -- which makes code more readable! -- is part of the syntax. Guido van Rossum was too experienced to leave it up to users who, frankly, rarely care.
Introducing psychopy_ext¶
Let's build something better, something that would:
- Streamline as many workflow steps as possible ("act clever")
- Seamlessly tie together these workflow steps.
- Facilitate reproducibility of the entire workflow.
Please give a warm welcome to psychopy_ext, a package that has these aims in mind though probably does not live up to them quite yet. Psychopy_ext is nothing but a collection of wrapper scripts to a number useful packages:
- PsychoPy is your bread and butter but you just want to never think again about saving data?
- Love matplotlib but want nice outputs?
- Were impressed by pandas but still don't know how to use it for computing accuracy?
- Would use PyMVPA to analyze your fMRI data but don't know how to even get started?
- Want to compare your data to outputs of computer vision models?
Then psychopy_ext is for you!
Let's go through a simple demo to understand what it gives you.
Step 1: Quick Demo¶
Note that in this demo we import fix so that the code could run from the notebook. In real life you don't do it and the Exp1 inherits from exp.Experiment.
from psychopy import visual
from psychopy_ext import exp
from collections import OrderedDict
import scripts.computer as computer
PATHS = exp.set_paths('trivial', computer)
class Exp1(exp.Experiment):
"""
Instructions (in reST format)
=============================
Press **spacebar** to start.
**Hit 'j'** to advance to the next trial, *Left-Shift + Esc* to exit.
"""
def __init__(self,
name='exp',
info=OrderedDict([('subjid', 'quick_'),
('session', 1),
]),
rp=None,
actions='run'
):
super(Exp1, self).__init__(name=name, info=info,
rp=rp, actions=actions,
paths=PATHS, computer=computer)
# user-defined parameters
self.ntrials = 8
self.stimsize = 2 # in deg
def create_stimuli(self):
"""Define your stimuli here, store them in self.s
"""
self.create_fixation()
self.s = {}
self.s['fix']= self.fixation
self.s['stim'] = visual.GratingStim(self.win, mask='gauss',
size=self.stimsize)
def create_trial(self):
"""Define trial composition
"""
self.trial = [exp.Event(self,
dur=.200, # in seconds
display=[self.s['stim'], self.s['fix']],
func=self.idle_event),
exp.Event(self,
dur=0,
display=self.s['fix'],
func=self.wait_until_response)
]
def create_exp_plan(self):
"""Put together trials
"""
exp_plan = []
for trialno in range(self.ntrials):
exp_plan.append(OrderedDict([
('trialno', trialno),
('onset', ''), # empty ones will be filled up
('dur', ''), # during runtime
('corr_resp', 1),
('subj_resp', ''),
('accuracy', ''),
('rt', ''),
]))
self.exp_plan = exp_plan
if __name__ == "__main__":
Exp1(rp={'no_output':True, 'debug':True}).run()
trial 1
An exception has occurred, use %tb to see the full traceback. SystemExit: 0
To exit: use 'exit', 'quit', or Ctrl-D.
Quick overview¶
Oopsies, that's complex! Let me parse that for you step-by-step:
- Import relevant modules, including computer parameters
- Define the experiment in a class
- Provide some info (instructions)
- Create stimuli
- Define the composition of a trial
- Define trial order and other info
Here's a pic to illustrate that (focus on the class Experiment for now):
More details of what it is doing¶
- Import modules that we need, such as psychopy and psychopy ext. There is also the
computermodule imported where parameters of your computer (screen size etc) are defined. Feel free to edit it. - Define your experiment as a class. Why bother with classes? The major advantage is that you can then inherit methods from a basic template supplied with
psychopy_ext. Then you only have to define or redefine methods that are not in that template. For example, looping through trials is in there, so if yu're happy with it, you don't have to write it again. - Everything within the Experiment is a method for reasons mentioned above. OK, there's an extra advantage: you can divide your script into short, easily readable bits of code and give them meaningful names. I hope you agree that this code is actually better organized than what you do usually.
- First, you give some custom parameters, such as the default subjid, in
__init__(). You can provide instructions how to run the experiment just above this method. If you format them using the reST syntax, as done in the example, it will render nicer-looking instructions. - Next, you define your stimuli in
create_stimuli(). A fancy fixation spot is available frompsychopy_ext. - Then you define the composition of your trial in
create_trial(). Each trial is composed of a series of Events that have a particular duration, stimuli that need to be displayed, and a particular function describing what to do (e.g., how to present stimuli). - Finally, you define all information that you need to run the experiment, such as the order of trials, in
create_exp_plan()as a list ofdictentries. Importantly, all the fields you provide here are written to the output file, and this is the only information that is written out. You can see that some fields, like accuracy, are empty. But they are filled in as the experiment progreses.
And that is all you need to create a full experiment. OK, but where is run()? It's in the Experiment template so you don't have to do anything extra.
Why bother...¶
It may seem that you could have easily written a similar experiment using the same old PsychoPy but don't underestimate how many things are happening behind the scenes:
- A log and data files are created and filled in.
- Everything is adjusted to the particular machine you are using (e.g., in the fMRI scanner you can define a different trigger key than the one you use in the experimental room).
- Looping through trials, trial durations and response collection is automatic too.
And that's only the beginning!
Quick exercises¶
Let's make sure you understand how classes work. What is the output of the following code?
def myfunc():
print 'stuff'
class Output(object):
def __init__(self):
print 'init'
def run(self):
print 'run'
How about this one?
class Output(object):
def __init__(self):
print 'init'
def run(self):
print 'run'
Output()
__main__.Output
And this?
class Output(object):
def __init__(self):
print 'init'
def go(self):
print 'go'
def run(self):
print 'run'
class Child(Output):
def run(self):
print 'child'
Child().run()
init go
Step 2: The Change Blindness Experiment¶
The best way to learn how to use psychopy_ext is to build your own experiment based on the demo above (or on more complex demos that come with the package). So let us reenact the Change Blindness Experiment from Part 2 using the psychopy_ext framework. It may be a good idea to keep both notebooks open as we are going to mostly copy/paste code.
Header: Importing modules¶
The first thing, as usual, is to import all relevant modules. But note that since psychopy_ext extends PsychoPy, we don't have to import most of PsychoPy's modules as in Part 2.
import numpy.random as rnd # for random number generators
from psychopy import visual
from psychopy_ext import exp
from collections import OrderedDict
import computer
PATHS = exp.set_paths('.', computer) # '.' means that the root directory for saving outout is here
PATHS['images'] = 'images'
all modules should be familiar more or less, except the mysterious computer. Well, that's the user-defined module where settings specific to your computers are defined (example settings are here). This is sper handy when you have several machines with different setups (e.g., one in your office, anoter in the testing room, and yet another at home for those of us who have no life).
Also note that we set the paths where all output files are supposed to be saved. This is done to help you organize your project better. Since we set up paths here, it also makes sense to define the path to the images folder here too. (See the example below or check the default paths here.)
Initial user-defined information¶
Next we define the ChangeDet class with its properties. This class is derived from exp.Experiment which, in turn, is nothing but the same old TrialHandler. Thus we ought to pass the relevant parameters here, as we do with method='sequential'. (Other options that the __init__ takes are explained in the documentation.)
The idea of __init__ is to define all (I mean, all) parameters here so that you can easily find and change them later.
There are several kinds of parameters you can define:
info: parameters that you want a user to be able to change on the go, e.g., participant IDrp: parameters conrolling the behavior of the program that you want a user to be bale to change on the go, e.g., whether to save outout or not- Other parameters that an outside user should not change on the go but that define how the program works, e.g., the number of trials. Those are defined as
self.var_namewhereselfmeans these variables are shared within the ChangeDet class -- you can access them from any other function in that class.
info and rp are in fact used in a GUI similar to the dialog box we used before (but more elaborate). You don't have to create the GUI yourself -- it all happens automatically and we'll demonstrate that later.
This is also where we define keys used to respond in the self.computer.valid_responses in the format {'key name': correct or incorrect response}. By default, Shift+Esc is used for escape and spacebar to advance from instructions to testing, so here we only need to define what counts as a correct response to advance to the next trial. Since everything in this experiment is "correct", we set space':1.
Note that we're defining instructions right at the top here. That serve a twofold purpose. On the one hand, it is natural to explain the experiment that the rest of the code enacts. On the other hand, this is also the docstring that is encouraged as a good programming practice, so you're documenting your code at the same time. Trying to act clever here!
We're also omitting writing date string to the output file because psychopy_ext creates a log file (you'll see later) with all this information and more.
Given all this information, psychopy_ext also automatically knows how to create output files and place them in a convenient location. So a large chunk of code is not necessary anymore.
class ChangeDet(exp.Experiment):
"""
Change Detection Experiment
===========================
In this experiment you will see photographs flickering with a tiny detail in them changing.
Your task is to detect where the change is occuring.
To make it harder, there are bubbles randomly covering the part of the photos.
Hit **spacebar to begin**. When you detect a change, hit **spacebar** again.
"""
def __init__(self,
name='exp',
info=OrderedDict([('exp_name', 'Change Detection'),
('subjid', 'cd_'),
('gender', ('male', 'female')),
('age', 18),
('left-handed', False)
]),
rp=None,
actions='run',
order='sequential'
):
super(ChangeDet, self).__init__(name=name, info=info,
rp=rp, actions=actions,
paths=PATHS, computer=computer)
# user-defined parameters
self.imlist = ['1','2','3','4','5','6'] # image names without the suffixes
self.asfx = 'a.jpg' # suffix for the first image
self.bsfx = 'b.jpg' # suffix for the second image
self.scrsize = (900, 600) # screen size in px
self.stimsize = (9, 6) # stimulus size in degrees visual angle
self.timelimit = 30 # sec
self.n_bubbles = 40
self.changetime = .500 #sec
self.computer.valid_responses = {'space': 1}
self.trial_instr = ('Press spacebar to start the trial.\n\n'
'Hit spacebar again when you detect a change.')
Create window¶
The window is usually created automatically for us, but in this particular case we want to be able to define its size so we have to override the particular window creation routine with our custom function. This example is also useful for you to see how to change the default behavior of psychopy_ext.
def create_win(self, *args, **kwargs):
super(ChangeDet, self).create_win(size=self.scrsize, units='deg',
*args, **kwargs)
Create stimuli¶
Should be straightforward by now, except that all stimuli are kept in a dict called self.s. Moreover, the window is defined in terms of degrees visual angle, so stimuli are implicitly using these units too.
def create_stimuli(self):
"""Define your stimuli here, store them in self.s
"""
self.s = {}
self.s['bitmap1'] = visual.ImageStim(self.win, size=self.stimsize)
self.s['bitmap2'] = visual.ImageStim(self.win, size=self.stimsize)
self.s['bubble'] = visual.Circle(self.win, fillColor='black', lineColor='black')
Trial structure¶
Remember, each trial consists of events of a certain duration, and we can pass a custom function of what should be happening during the trial. Here we create structure with a single event that lasts the maximum duration (i.e., 30 sec) and call a custom function show_stim that will control flipping between images, drawing bubbles etc.
def create_trial(self):
"""Define trial composition
"""
self.trial = [exp.Event(self,
dur=self.timelimit, # in seconds
display=[self.s['bitmap1'], self.s['bitmap2'], self.s['bubble']],
func=self.show_stim)
]
Experimental plan¶
Here we put all information about stimuli and so on that will be recorded in the output files.
def create_exp_plan(self):
"""Put together trials
"""
# Check if all images exist
for im in self.imlist:
if (not os.path.exists(os.path.join(self.paths['images'], im+self.asfx)) or
not os.path.exists(os.path.join(self.paths['images'], im+self.bsfx))):
raise Exception('Image files not found in image folder: ' + str(im))
# Randomize the image order
rnd.shuffle(self.imlist)
# Create the orientations list: half upright, half inverted
orilist = [0,1]*(len(self.imlist)/2)
# Randomize the orientation order
rnd.shuffle(orilist)
exp_plan = []
for im, ori in zip(self.imlist, orilist):
exp_plan.append(OrderedDict([
('im', im),
('ori', ori),
('onset', ''), # empty ones will be filled up
('dur', ''), # during runtime
('corr_resp', 1),
('subj_resp', ''),
('accuracy', ''),
('rt', ''),
]))
self.exp_plan = exp_plan
Before trial¶
We need to show instructions before each trial and decide whether stimuli will be upright or inverted. To be more efficient, we first load images (it may take some time) and only when that is ready, show instructions.
def before_trial(self):
"""Set up stimuli prior to a trial
"""
im_fname = os.path.join(self.paths['images'], self.this_trial['im'])
self.s['bitmap1'].setImage(im_fname + self.asfx)
self.s['bitmap1'].setOri(self.this_trial['ori'])
self.s['bitmap2'].setImage(im_fname + self.bsfx)
self.s['bitmap2'].setOri(self.this_trial['ori'])
self.bitmap = self.s['bitmap1']
if self.thisTrialN > 0: # no need for instructions for the first trial
self.show_text(text=self.trial_instr, wait=0)
Show stimuli¶
Finally, we define what happens during each trial. It's mostly copy/paste from our previous implementation with one significant change: we use last_keypress() function to record user responses. This function is aware of the keys that we accept as responses as well as about special keys, such as Shift+Esc for exit. We therefore do not have to then check manually if the participant pressed a spacebar or and exit key. Moreover, the information about responses needs to be passed further (for writing responses to files etc) thus we have to include the return keys statement at the end.
Also notice that since everything is defined in terms of degrees visual angle, we have to adjust bubble size accordingly.
def show_stim(self, *args, **kwargs):
"""Control stimuli during the trial
"""
# Empty the keypresses list
event.clearEvents()
keys = []
change_clock = core.Clock()
# Start the trial
# Stop trial if spacebar or escape has been pressed, or if 30s have passed
while len(keys) == 0 and self.trial_clock.getTime() < self.this_event.dur:
# Switch the image
if self.bitmap == self.s['bitmap1']:
self.bitmap = self.s['bitmap2']
else:
self.bitmap = self.s['bitmap1']
self.bitmap.draw()
# Draw bubbles of increasing radius at random positions
for radius in range(self.n_bubbles):
self.s['bubble'].setRadius(radius/100.)
self.s['bubble'].setPos(((rnd.random()-.5) * self.stimsize[0],
(rnd.random()-.5) * self.stimsize[1] ))
self.s['bubble'].draw()
# Show the new screen we've drawn
self.win.flip()
# For the duration of 'changetime',
# Listen for a spacebar or escape press
change_clock.reset()
while change_clock.getTime() <= self.changetime:
keys = self.last_keypress(keyList=self.computer.valid_responses.keys(),
timeStamped=self.trial_clock)
if len(keys) > 0:
print keys
break
return keys
And that's it!¶
Notice that you did not have to do many things here anymore:
- Define the window and its properties
- Define trial loop
- Define output files and write to them
- Define log files that record what happens during the experiment, including errors
- Catch escapes
- Deal with instructions at the beginning and end
Change Detection Experiment: full code¶
%load scripts/changedet.py
import os
import numpy.random as rnd # for random number generators
from psychopy import visual, core, event
from psychopy_ext import exp
from collections import OrderedDict
import scripts.computer as computer
PATHS = exp.set_paths('change_detection', computer)
PATHS['images'] = '../Part2/images/'
class ChangeDet(exp.Experiment):
"""
Change Detection Experiment
===========================
In this experiment you will see photographs flickering with a tiny detail in them changing.
Your task is to detect where the change is occuring.
To make it harder, there are bubbles randomly covering the part of the photos.
Hit **spacebar to begin**. When you detect a change, hit **spacebar** again.
"""
def __init__(self,
name='exp',
info=OrderedDict([('exp_name', 'Change Detection'),
('subjid', 'cd_'),
('gender', ('male', 'female')),
('age', 18),
('left-handed', False)
]),
rp=None,
actions='run',
order='sequential'
):
super(ChangeDet, self).__init__(name=name, info=info,
rp=rp, actions=actions,
paths=PATHS, computer=computer)
# user-defined parameters
self.imlist = ['1','2','3','4','5','6'] # image names without the suffixes
self.asfx = 'a.jpg' # suffix for the first image
self.bsfx = 'b.jpg' # suffix for the second image
self.scrsize = (900, 600) # screen size in px
self.stimsize = (9, 6) # stimulus size in degrees visual angle
self.timelimit = 30 # sec
self.n_bubbles = 40
self.changetime = .500 #sec
self.computer.valid_responses = {'space': 1}
self.trial_instr = ('Press spacebar to start the trial.\n\n'
'Hit spacebar again when you detect a change.')
def create_win(self, *args, **kwargs):
super(ChangeDet, self).create_win(size=self.scrsize, units='deg',
*args, **kwargs)
def create_stimuli(self):
"""Define your stimuli here, store them in self.s
"""
self.s = {}
self.s['bitmap1'] = visual.ImageStim(self.win, size=self.stimsize)
self.s['bitmap2'] = visual.ImageStim(self.win, size=self.stimsize)
self.s['bubble'] = visual.Circle(self.win, fillColor='black', lineColor='black')
def create_trial(self):
"""Define trial composition
"""
self.trial = [exp.Event(self,
dur=self.timelimit, # in seconds
display=[self.s['bitmap1'], self.s['bitmap2']],
func=self.show_stim)
]
def create_exp_plan(self):
"""Put together trials
"""
# Check if all images exist
for im in self.imlist:
if (not os.path.exists(os.path.join(self.paths['images'], im+self.asfx)) or
not os.path.exists(os.path.join(self.paths['images'], im+self.bsfx))):
raise Exception('Image files not found in image folder: ' + str(im))
# Randomize the image order
rnd.shuffle(self.imlist)
# Create the orientations list: half upright, half inverted
orilist = [0,180]*(len(self.imlist)/2)
# Randomize the orientation order
rnd.shuffle(orilist)
exp_plan = []
for trialno, (im, ori) in enumerate(zip(self.imlist, orilist)):
exp_plan.append(OrderedDict([
('im', im),
('ori', ori),
('onset', ''), # empty ones will be filled up
('dur', ''), # during runtime
('corr_resp', 1),
('subj_resp', ''),
('accuracy', ''),
('rt', ''),
]))
self.exp_plan = exp_plan
def before_trial(self):
"""Set up stimuli prior to a trial
"""
im_fname = os.path.join(self.paths['images'], self.this_trial['im'])
self.s['bitmap1'].setImage(im_fname + self.asfx)
self.s['bitmap1'].setOri(self.this_trial['ori'])
self.s['bitmap2'].setImage(im_fname + self.bsfx)
self.s['bitmap2'].setOri(self.this_trial['ori'])
self.bitmap = self.s['bitmap1']
if self.thisTrialN > 0: # no need for instructions for the first trial
self.show_text(text=self.trial_instr, wait=0)
def show_stim(self, *args, **kwargs):
"""Control stimuli during the trial
"""
# Empty the keypresses list
event.clearEvents()
keys = []
change_clock = core.Clock()
# Start the trial
# Stop trial if spacebar or escape has been pressed, or if 30s have passed
while len(keys) == 0 and self.trial_clock.getTime() < self.this_event.dur:
# Switch the image
if self.bitmap == self.s['bitmap1']:
self.bitmap = self.s['bitmap2']
else:
self.bitmap = self.s['bitmap1']
self.bitmap.draw()
# Draw bubbles of increasing radius at random positions
for radius in range(self.n_bubbles):
self.s['bubble'].setRadius(radius/100.)
self.s['bubble'].setPos(((rnd.random()-.5) * self.stimsize[0],
(rnd.random()-.5) * self.stimsize[1] ))
self.s['bubble'].draw()
# Show the new screen we've drawn
self.win.flip()
# For the duration of 'changetime',
# Listen for a spacebar or escape press
change_clock.reset()
while change_clock.getTime() <= self.changetime:
keys = self.last_keypress(keyList=self.computer.valid_responses.keys(),
timeStamped=self.trial_clock)
if len(keys) > 0:
print keys
break
return keys
if __name__ == "__main__":
ChangeDet(rp={'no_output':True, 'debug':True}).run()
Step 3: Data analysis¶
pscyhopy_ext is not meant only for helping to run experiments. As we discussed above, there are many other tasks that a researcher needs to do. One of them is data analysis. You may be used to doing it in Excel or SPSS, or R, but Python is actually sufficient to carry out many simple and more complex analyses. And it may also be nice to have your experimental and analysis code together in a single file.
There is the pandas package in Python offering great data analysis capabilites. psychopy_ext wraps it with the stats and plot modules to help you do typical analyses efficiently. For more power, you may want to use statsmodels.
So let's look at how to analyze data from your experiment. For this example, we will use data from a paper by de-Wit, Kubilius et al. (2013).
Reading in data¶
Reading in data is done by a clever read_csv method that can get datga both from local sources (your computer) and the internet. In this example, we fecth data for 12 control participants (so that is twelve files) and concatenate them together into a single large structure, called a DataFrame, as seen in the output.
import pandas
# get data from de-Wit, Kubilius et al. (2013); will take some time
path = 'https://bitbucket.org/qbilius/df/raw/aed0ac3eba09d1d688e87816069f5b05e127519e/data/controls2_%02d.csv'
data = [pandas.read_csv(path % i) for i in range(1,13)]
df = pandas.concat(data, ignore_index=True)
df
| expName | subjID | runNo | runType | paraType | paraName | blockNo | startBlock | trialNo | cond | onset | actualOnset | dur | context | pos | corrResp | subjResp | accuracy | RT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | run | controls2_01 | 1 | main | event | para06 | 0 | 1 | 0 | 7 | 0.0 | 0.004020 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1691.731086 |
| 1 | run | controls2_01 | 1 | main | event | para06 | 1 | 1 | 1 | 1 | 0.3 | 1.693590 | 0.3 | Parts | Top Left | 1 | 4 | Incorrect | 1847.628840 |
| 2 | run | controls2_01 | 1 | main | event | para06 | 2 | 1 | 2 | 1 | 0.6 | 3.541586 | 0.3 | Parts | Top Left | 1 | 1 | Correct | 1663.642791 |
| 3 | run | controls2_01 | 1 | main | event | para06 | 3 | 1 | 3 | 6 | 0.9 | 5.205333 | 0.3 | Whole | Top Right | 2 | 2 | Correct | 1511.654446 |
| 4 | run | controls2_01 | 1 | main | event | para06 | 4 | 1 | 4 | 1 | 1.2 | 6.717345 | 0.3 | Parts | Top Left | 1 | 1 | Correct | 1959.769867 |
| 5 | run | controls2_01 | 1 | main | event | para06 | 5 | 1 | 5 | 1 | 1.5 | 8.677413 | 0.3 | Parts | Top Left | 1 | 1 | Correct | 1335.666005 |
| 6 | run | controls2_01 | 1 | main | event | para06 | 6 | 1 | 6 | 8 | 1.8 | 10.013345 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1495.693448 |
| 7 | run | controls2_01 | 1 | main | event | para06 | 7 | 1 | 7 | 3 | 2.1 | 11.509252 | 0.3 | Parts | Bottom Left | 3 | 3 | Correct | 2319.651968 |
| 8 | run | controls2_01 | 1 | main | event | para06 | 8 | 1 | 8 | 7 | 2.4 | 13.829205 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1919.638110 |
| 9 | run | controls2_01 | 1 | main | event | para06 | 9 | 1 | 9 | 5 | 2.7 | 15.749133 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1463.658959 |
| 10 | run | controls2_01 | 1 | main | event | para06 | 10 | 1 | 10 | 7 | 3.0 | 17.213078 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1335.654028 |
| 11 | run | controls2_01 | 1 | main | event | para06 | 11 | 1 | 11 | 8 | 3.3 | 18.549068 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1495.674200 |
| 12 | run | controls2_01 | 1 | main | event | para06 | 12 | 1 | 12 | 4 | 3.6 | 20.044989 | 0.3 | Parts | Bottom Right | 4 | 4 | Correct | 2079.699771 |
| 13 | run | controls2_01 | 1 | main | event | para06 | 13 | 1 | 13 | 5 | 3.9 | 22.125191 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1319.653252 |
| 14 | run | controls2_01 | 1 | main | event | para06 | 14 | 1 | 14 | 3 | 4.2 | 23.444928 | 0.3 | Parts | Bottom Left | 3 | 4 | Incorrect | 1967.667815 |
| 15 | run | controls2_01 | 1 | main | event | para06 | 15 | 1 | 15 | 3 | 4.5 | 25.412903 | 0.3 | Parts | Bottom Left | 3 | 3 | Correct | 1455.684448 |
| 16 | run | controls2_01 | 1 | main | event | para06 | 16 | 1 | 16 | 1 | 4.8 | 26.868817 | 0.3 | Parts | Top Left | 1 | 4 | Incorrect | 2111.706457 |
| 17 | run | controls2_01 | 1 | main | event | para06 | 17 | 1 | 17 | 4 | 5.1 | 28.980800 | 0.3 | Parts | Bottom Right | 4 | 1 | Incorrect | 1575.592539 |
| 18 | run | controls2_01 | 1 | main | event | para06 | 18 | 1 | 18 | 1 | 5.4 | 30.556723 | 0.3 | Parts | Top Left | 1 | 1 | Correct | 2583.716753 |
| 19 | run | controls2_01 | 1 | main | event | para06 | 19 | 1 | 19 | 5 | 5.7 | 33.140642 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1383.702553 |
| 20 | run | controls2_01 | 1 | main | event | para06 | 20 | 1 | 20 | 4 | 6.0 | 34.524658 | 0.3 | Parts | Bottom Right | 4 | 4 | Correct | 1615.689649 |
| 21 | run | controls2_01 | 1 | main | event | para06 | 21 | 1 | 21 | 6 | 6.3 | 36.140685 | 0.3 | Whole | Top Right | 2 | 2 | Correct | 1399.671678 |
| 22 | run | controls2_01 | 1 | main | event | para06 | 22 | 1 | 22 | 8 | 6.6 | 37.540624 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1631.644232 |
| 23 | run | controls2_01 | 1 | main | event | para06 | 23 | 1 | 23 | 5 | 6.9 | 39.172514 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1423.674768 |
| 24 | run | controls2_01 | 1 | main | event | para06 | 24 | 1 | 24 | 2 | 7.2 | 40.596501 | 0.3 | Parts | Top Right | 2 | 3 | Incorrect | 2207.632844 |
| 25 | run | controls2_01 | 1 | main | event | para06 | 25 | 1 | 25 | 5 | 7.5 | 42.804405 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1447.695395 |
| 26 | run | controls2_01 | 1 | main | event | para06 | 26 | 1 | 26 | 6 | 7.8 | 44.252363 | 0.3 | Whole | Top Right | 2 | 2 | Correct | 1447.696678 |
| 27 | run | controls2_01 | 1 | main | event | para06 | 27 | 1 | 27 | 8 | 8.1 | 45.700631 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1303.450589 |
| 28 | run | controls2_01 | 1 | main | event | para06 | 28 | 1 | 28 | 0 | 8.4 | 47.004304 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 29 | run | controls2_01 | 1 | main | event | para06 | 29 | 1 | 29 | 0 | 8.7 | 47.304275 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 30 | run | controls2_01 | 1 | main | event | para06 | 30 | 1 | 30 | 0 | 9.0 | 47.604785 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 31 | run | controls2_01 | 1 | main | event | para06 | 31 | 1 | 31 | 6 | 9.3 | 47.904776 | 0.3 | Whole | Top Right | 2 | 2 | Correct | 1427.156452 |
| 32 | run | controls2_01 | 1 | main | event | para06 | 32 | 1 | 32 | 5 | 9.6 | 49.332309 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1287.643999 |
| 33 | run | controls2_01 | 1 | main | event | para06 | 33 | 1 | 33 | 8 | 9.9 | 50.620247 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1287.665385 |
| 34 | run | controls2_01 | 1 | main | event | para06 | 34 | 1 | 34 | 2 | 10.2 | 51.908175 | 0.3 | Parts | Top Right | 2 | 2 | Correct | 1991.666627 |
| 35 | run | controls2_01 | 1 | main | event | para06 | 35 | 1 | 35 | 3 | 10.5 | 53.900168 | 0.3 | Parts | Bottom Left | 3 | 3 | Correct | 1487.696267 |
| 36 | run | controls2_01 | 1 | main | event | para06 | 36 | 1 | 36 | 1 | 10.8 | 55.388153 | 0.3 | Parts | Top Left | 1 | 4 | Incorrect | 1607.679209 |
| 37 | run | controls2_01 | 1 | main | event | para06 | 37 | 1 | 37 | 5 | 11.1 | 56.996092 | 0.3 | Whole | Top Left | 1 | 1 | Correct | 1495.672061 |
| 38 | run | controls2_01 | 1 | main | event | para06 | 38 | 1 | 38 | 3 | 11.4 | 58.492151 | 0.3 | Parts | Bottom Left | 3 | 3 | Correct | 2031.672205 |
| 39 | run | controls2_01 | 1 | main | event | para06 | 39 | 1 | 39 | 4 | 11.7 | 60.523999 | 0.3 | Parts | Bottom Right | 4 | 3 | Incorrect | 3711.614753 |
| 40 | run | controls2_01 | 1 | main | event | para06 | 40 | 1 | 40 | 2 | 12.0 | 64.235910 | 0.3 | Parts | Top Right | 2 | 2 | Correct | 1815.614455 |
| 41 | run | controls2_01 | 1 | main | event | para06 | 41 | 1 | 41 | 1 | 12.3 | 66.051824 | 0.3 | Parts | Top Left | 1 | 4 | Incorrect | 1759.731286 |
| 42 | run | controls2_01 | 1 | main | event | para06 | 42 | 1 | 42 | 4 | 12.6 | 67.811819 | 0.3 | Parts | Bottom Right | 4 | 1 | Incorrect | 1607.676643 |
| 43 | run | controls2_01 | 1 | main | event | para06 | 43 | 1 | 43 | 7 | 12.9 | 69.419797 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1303.664879 |
| 44 | run | controls2_01 | 1 | main | event | para06 | 44 | 1 | 44 | 7 | 13.2 | 70.723707 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1359.686204 |
| 45 | run | controls2_01 | 1 | main | event | para06 | 45 | 1 | 45 | 0 | 13.5 | 72.083681 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 46 | run | controls2_01 | 1 | main | event | para06 | 46 | 1 | 46 | 0 | 13.8 | 72.383803 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 47 | run | controls2_01 | 1 | main | event | para06 | 47 | 1 | 47 | 3 | 14.1 | 72.684001 | 0.3 | Parts | Bottom Left | 3 | 3 | Correct | 2583.301432 |
| 48 | run | controls2_01 | 1 | main | event | para06 | 48 | 1 | 48 | 0 | 14.4 | 75.267621 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 49 | run | controls2_01 | 1 | main | event | para06 | 49 | 1 | 49 | 4 | 14.7 | 75.567601 | 0.3 | Parts | Bottom Right | 4 | 3 | Incorrect | 3435.445341 |
| 50 | run | controls2_01 | 1 | main | event | para06 | 50 | 1 | 50 | 8 | 15.0 | 79.003531 | 0.3 | Whole | Bottom Right | 4 | 4 | Correct | 1655.706348 |
| 51 | run | controls2_01 | 1 | main | event | para06 | 51 | 1 | 51 | 7 | 15.3 | 80.659547 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1415.669888 |
| 52 | run | controls2_01 | 1 | main | event | para06 | 52 | 1 | 52 | 6 | 15.6 | 82.075451 | 0.3 | Whole | Top Right | 2 | 2 | Correct | 1455.673755 |
| 53 | run | controls2_01 | 1 | main | event | para06 | 53 | 1 | 53 | 7 | 15.9 | 83.531444 | 0.3 | Whole | Bottom Left | 3 | 3 | Correct | 1311.654360 |
| 54 | run | controls2_01 | 1 | main | event | para06 | 54 | 1 | 54 | 2 | 16.2 | 84.843354 | 0.3 | Parts | Top Right | 2 | 2 | Correct | 1751.674652 |
| 55 | run | controls2_01 | 1 | main | event | para06 | 55 | 1 | 55 | 1 | 16.5 | 86.595483 | 0.3 | Parts | Top Left | 1 | 4 | Incorrect | 1639.529776 |
| 56 | run | controls2_01 | 1 | main | event | para06 | 56 | 1 | 56 | 1 | 16.8 | 88.235367 | 0.3 | Parts | Top Left | 1 | 1 | Correct | 2151.664557 |
| 57 | run | controls2_01 | 1 | main | event | para06 | 57 | 1 | 57 | 2 | 17.1 | 90.387212 | 0.3 | Parts | Top Right | 2 | 2 | Correct | 2087.715344 |
| 58 | run | controls2_01 | 1 | main | event | para06 | 58 | 1 | 58 | 0 | 17.4 | 92.475212 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| 59 | run | controls2_01 | 1 | main | event | para06 | 59 | 1 | 59 | 0 | 17.7 | 92.775342 | 0.3 | Fixation | NaN | NaN | NaN | Correct | NaN |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
1920 rows × 19 columns
If you did your experiment using psychopy_ext, then there is a helper function in the exp module, called get_behav_data(), that will find and import the relevant data from you experiment.
Aggegating data¶
Typically we want to average data across participants and plot it comparing several conditions. Aggregating data in pandas is not too bad but still might take some effort, and plotting it in a nice way is definitely a not trivial. Let's see how that can be done in psychopy_ext. Let's first compute reaction times using the stats.aggregate() function:
from psychopy_ext import stats
rt = stats.aggregate(df, values='RT', cols='context')
rt
| cols.context | Whole | Parts | Fixation |
|---|---|---|---|
| RT | 1414.651203 | 2073.345894 | NaN |
1 rows × 3 columns
If you are used to Excel PivotCharts, this should look familiar. We simply specify the data source (df), which column we want to aggregate (values) and how it should be structured (cols). Here we say that we want to split data by the context column. If you look at that column, you'll see there are three unique values in it: 'Whole', 'Parts' and 'Fixation', thus in the output you see an average for each of these values. There were no responses during fixation, so the average is coded as 'NaN' ('not a number').
Don't want this fixation? Let's filter it out:
df = df[df.context != 'Fixation']
rt = stats.aggregate(df, values='RT', cols='context')
rt
| cols.context | Whole | Parts |
|---|---|---|
| RT | 1414.651203 | 2073.345894 |
1 rows × 2 columns
The way it works is by first evaluating which elements in the 'context' column are not 'Fixation' (df.context != 'Fixation'). The output of this is a boolean vector, whcih we then use to filter the entire DataFrame.
Now let's compute these averages for each participant separately (this will be used to compute error bars in plotting later):
rt = stats.aggregate(df, values='RT', cols='context', yerr='subjID')
rt
| cols.context | Whole | Parts |
|---|---|---|
| yerr.subjID | ||
| controls2_01 | 1436.928122 | 1907.336854 |
| controls2_02 | 2008.916406 | 2456.431393 |
| controls2_03 | 1369.047977 | 1907.168522 |
| controls2_04 | 1261.510856 | 1992.553333 |
| controls2_05 | 1376.929082 | 1966.202955 |
| controls2_06 | 1413.390391 | 1912.792788 |
| controls2_07 | 1202.785780 | 2412.898507 |
| controls2_08 | 1420.174382 | 2025.540335 |
| controls2_09 | 1530.765389 | 2571.260439 |
| controls2_10 | 1244.360752 | 1788.609975 |
| controls2_11 | 1304.750426 | 1938.166938 |
| controls2_12 | 1406.254871 | 2001.188694 |
12 rows × 2 columns
Also for more conditions:
rt = stats.aggregate(df, values='RT', cols=['pos', 'context'])
rt
| cols.pos | Bottom Left | Top Left | Top Right | Bottom Right | ||||
|---|---|---|---|---|---|---|---|---|
| cols.context | Whole | Parts | Whole | Parts | Whole | Parts | Whole | Parts |
| RT | 1399.365869 | 1954.816429 | 1406.562769 | 2183.723032 | 1425.831487 | 2005.071479 | 1427.743824 | 2149.539713 |
1 rows × 8 columns
But what it you want to compute accuracy? There's a function for that too, called accuracy(). For it to work, we need to specify which values are considered "correct" and which are considered "incorrect":
acc = stats.accuracy(df, values='accuracy', cols='context', yerr='subjID', correct='Correct', incorrect='Incorrect')
acc
| cols.context | Whole | Parts |
|---|---|---|
| yerr.subjID | ||
| controls2_01 | 0.985507 | 0.797101 |
| controls2_02 | 1.000000 | 0.956522 |
| controls2_03 | 0.985507 | 0.898551 |
| controls2_04 | 0.971014 | 0.840580 |
| controls2_05 | 0.971014 | 0.985507 |
| controls2_06 | 0.971014 | 0.985507 |
| controls2_07 | 0.985507 | 0.913043 |
| controls2_08 | 1.000000 | 0.884058 |
| controls2_09 | 1.000000 | 0.811594 |
| controls2_10 | 0.971014 | 0.927536 |
| controls2_11 | 1.000000 | 0.913043 |
| controls2_12 | 1.000000 | 0.971014 |
12 rows × 2 columns
Plotting¶
Because we aggregated data using psychopy_ext, plotting it is super quick now with the plot() function:
%matplotlib inline
from psychopy_ext import plot
plt = plot.Plot()
plt.plot(acc, kind='bar')
plt.show()
Notice how you get error bars for free and even if the two conditions are significantly different from each other!
It can also produce other kinds of plots (see the Gallery). One of the nicer ones is called a bean plot. It cleverly combines all data points (as these horizontal bars; if several data points coincide, the line is longer) and the estimated density of the measurements, so that you can quickly see the distribution of your data and spot any outliers or non-normality.
plt = plot.Plot()
plt.plot(acc, kind='bean')
plt.show()
You can also easily plot several subplots:
rt = stats.aggregate(df, values='RT', cols='context', subplots='pos', yerr='subjID')
plt = plot.Plot()
plt.plot(rt, kind='bean')
plt.show()
There are many more option available in this module, so check out its documentation.
Also, I hope you have noticed by now that the plots in this tutorial are beautiful. They are so pretty by default thanks to a great design by the Seaborn package, so you may want to check out that library too.
Step 4: Integrated development¶
More tasks¶
So far we've looked at examples where a single experiment is implemented. But often we have several experiments in the same study -- how could we accomodate for this? psychopy_ext has a concept of a Task: Every experiment of composed of several Tasks that we ask participants to perform. Thus, if you have two tasks, it would looks something like the following (borrowing code from the twotasks.py demo):
class TwoTasks(exp.Experiment):
def __init__(self, ...):
self.tasks = [Train, Test]
class Train(exp.Task):
def __init__(self, parent):
...
class Test(exp.Task):
def __init__(self, parent):
...
Here the TwoTasks class knows about Train and Test because we put them in the self.tasks variable. Train and Test know about TwoTasks through the parent argument that is passed when these classes are initiated during runtime.
More experiments¶
We can also have several separate experiments, like Study 1 and Study 2. You simply make two files in the scripts folder, study1.py and study2.py. The data for these experiments by default are saved in separate locations (that are called, guess what, study1 and study2).
GUI¶
As secretly mentioned before, psychopy_ext can automatically produce rather complex GUIs such that you can fully customize your experiment before running it. These GUIs are constructed using information you provide at the top of Experiment class from the info and rp parameters. And it looks like this:
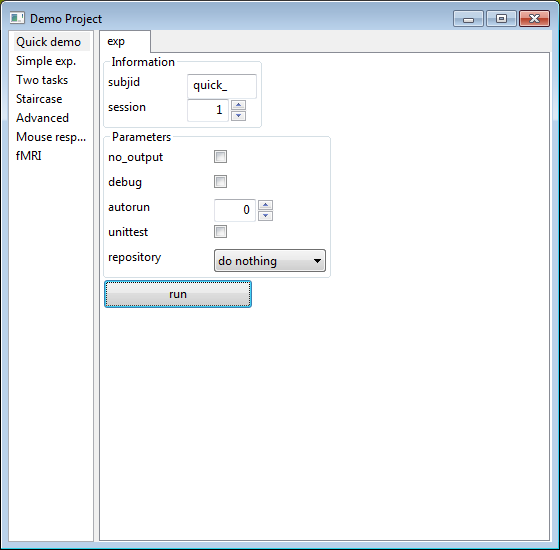
Because this GUI is so convenient, it is actually the default mode of running code in psychopy_ext. Calling any command is unified within the run.py file, providing a very easy, replicable way to run code and analyses. It looks something like this:
%load run.py
#! /usr/bin/env python
from psychopy_ext import ui
__author__ = "Jonas Kubilius"
__version__ = "0.1"
exp_choices = [
ui.Choices('scripts.trivial', name='Quick demo'),
ui.Choices('scripts.changedet', name='Change Detection Experiment')
]
# bring up the graphic user interface or interpret command line inputs
# usually you can skip the size parameter
ui.Control(exp_choices, title='Demo Project', size=(560,550))
Here we have two important bits: defining Choices (tabs on the left side of GUI) that correspond to diffferent experiments (not tasks), and Control that creates the GUI itself.
It is not possible to demonstrate this functionality from a notebook directly so we will use IPython magic commands to execute a shell command. Try this:
%run run.py
Command line interface¶
But not everybody is keen to use these GUIs. Thus, psychopy_ext also offers command-line interface in the following manner (running it from the Terminal, Powershell, cmd or a similar program):
python run.py myproject exp run --subjid subj_01 --n
Here we provide the name of the project (in case there are several), task name in it (experiment, analysis, simulation etc.), function we want to call (run) and parameters for info and rp. Look at this figure above for a graphical illustration. Notice how you can abbreviate parameters: --n instead of no_output.
Try it in practice:
%run run.py changedet exp run --subjid subj_01 --debug --n
Extra: Other features that will blow your mind¶
Autorun¶
You don't want your experiment to fail with your first participant or after a small tweak in the middle of a pilot run, do you? Imagine you run an experiment for an hour only to learn later that no data was recorded! But then the only way to know if it is really ready is to run it yourself -- which is reasonable to do several times but definitely not after every little tweak that "shouldn't change anything". People with a long enough history in development know that these small innocent-looking tweaks can sometimes lead to small accidental issues such as output files not being saved or the script breaking in the middle of running...
To prevent from such unforseen problems occuring, the best strategy is to have automated tests, called unit tests, that would quickly check if everything is in order. For experiments, this means being able to run the experiment automatically to make sure it works and produces meaningful output. psychopy_ext comes with this functionality out of the box. Simply choose the "unittest" option in the gui or --unittest in the command line.
Let's try that for the Change Detection experiment:
%run run.py changedet exp run --d --n --unittest
initializing...FreeType import Failed: expected string or Unicode object, NoneType found
Change Detection Experiment
===========================
In this experiment you will see photographs flickering with a tiny detail in them changing.
Your task is to detect where the change is occuring.
To make it harder, there are bubbles randomly covering the part of the photos.
Hit **spacebar to begin**. When you detect a change, hit **spacebar** again.
trial 1Press spacebar to start the trial.
Hit spacebar again when you detect a change.
trial 2Press spacebar to start the trial.
Hit spacebar again when you detect a change.
trial 3Press spacebar to start the trial.
Hit spacebar again when you detect a change.
trial 4Press spacebar to start the trial.
Hit spacebar again when you detect a change.
trial 5Press spacebar to start the trial.
Hit spacebar again when you detect a change.
End of Experiment. Thank you!
You'll notice that the whole experiment runs on its own at a very high pace -- or you may not even see anything really because it's so short. But you see that it prints out what it can see on the screen and thus you can easily verify it went through the entire experiment without any errors.
So that's cool and good for a quick reassurance that all is in order! But sometimes, especially for longer experiments composed of multiple tasks, you actually want to run the experiment half manually, such that you can read instructions and advance to testing, then quickly go through trials, then read the instruction again etc. For this, there is an autorun option that also allows you to choose how quickly we run through trials ('1' means the actualy speed, '100' would be 100x faster).
Notice that the program is actually performing the experiment just like a participant would, so in the end we get an output file that we can use to meaningfully test our analysis scripts. In fact, it is greatly encouraged to write your data analysis scripts at the same time as your experimental scripts. You will often see that by doing the analysis on such simulated data you will learn that perhaps a particular information about a stimulus or condition is missing and would be useful for the analysis.
Analyzer¶
psychopy_ext also has a prototype for a quick data analysis, drawing ideas from Excel's PivotChart, and, consistent with PsychoPy's Builder and Coder modules, named the Analyzer. It is really an early prototype and not even documented yet but here's a quick preview:
# first get the data from de-Wit, Kubilius et al., (2013) again
import pandas
path = 'https://bitbucket.org/qbilius/df/raw/aed0ac3eba09d1d688e87816069f5b05e127519e/data/controls2_%02d.csv'
data = [pandas.read_csv(path % i) for i in range(1,13)]
df = pandas.concat(data, ignore_index=True)
df.to_csv('data.csv') # save to a file
# now open the Analyzer GUI
from psychopy_ext import analyzer
analyzer.run()
Computer vision models included¶
Suppose you run an experiment and find that people can tell if there is an animal in an image based on a very brief presentation. Obviously, you may want to make claims that people process this high level object and scene information very quickly, perhaps even in a feedforward manner. But you have to be careful here. Maybe people are able to do this task based on some low level information, such as a particular power spectrum difference between animal and non-animal stimuli.
A good strategy to address these concerns, to a certain extent at least, is to process your stimuli with a model of an early visual cortex, and use some sort of categorization algorithm (such as computing a distance betweet the two categories, applying a support vector machine, or using a number of other machine learning techniques). Typically it is a tedious procedure but psychopy_ext comes with several simple models included in the models module, such as Pixelwise (for pixelwise differences), GaborJet (a very simplistic V1 model) from Biederman lab, and HMAX'99, the early implementation of the HMAX model.
In the example below, we use the images from the Change Detection experiment to see how different they appear to the HMAX'99 model. You shoudl see that some stimuli are much more different from the others (dark red spots) but on the diagonal images are quite similar to each other, as it should be since version a and version b are only slightly different.
import glob
from scipy import misc
from psychopy_ext import models
import matplotlib.pyplot as plt
# read images from the Change Detection experiment
ims = glob.glob('../Part2/images/*.jpg')
ims = [misc.imread(im) for im in ims]
# crop and resize them to (128, 128)
ims = [misc.imresize(im[:,im.shape[0]], (128, 128)) for im in ims]
hmax = models.HMAX()
hmax.compare(ims)
Export your stimuli to vector graphics¶
Often you want to be able to export stimuli that you used in the experiment for using in a paper. One possibility is to capture display using PsychoPy's getMovieFrame and saveMovieFrames fucntionality that captures what is presented on the screen. However, the resolution of this export is going to be low and you will often be unable to use these images on a poster or for a paper.
A better approach is to export stimuli in the SVG (scalable vector graphics) format that exports objects rather than pixels, and, as the name implies, you can scale them as much as you like without losing quality in programs like Inkscape, Scribus, and Adobe Illustrator. To help you with that, psychopy_ext provides an undocumented (read: not fully functional) feature: a whole SVG module in the exp class that will try to export your stimuli in the svg format as much as possible. Note that currently it only works with shape and text stimuli (lines, circles, etc) and images (that are, of course, not really scalable). This is how it works:
from IPython.display import SVG, display
from psychopy import visual, event
from psychopy_ext import exp
win = visual.Window(size=(400,400))
stim1 = visual.Circle(win)
stim2 = visual.TextStim(win, "Ceci n'est pas un cercle!", height=.1)
# write to svg
svg = exp.SVG(win, filename='stimuli.svg')
svg.write(stim1)
svg.write(stim2)
svg.svgfile.save()
display(SVG('stimuli.svg'))
# optional: show stimuli on the screen too
stim1.draw()
stim2.draw()
win.flip()
event.waitKeys()
win.close()
FreeType import Failed: expected string or Unicode object, NoneType found
1.2722 WARNING Creating new monitor... 1.2724 WARNING Creating new monitor...