2D datasets with Mesh Neural Networks¶
0. Choose dataset¶
In [1]:
dataset = "spirals"
1. Import Libraries¶
In [2]:
from sklearn import datasets
import numpy as np
import matplotlib as mpl
mpl.use('Agg')
import matplotlib.pyplot as plt
from matplotlib import animation, rc
import IPython.display
from PIL import Image
rc('animation', html='html5')
2. Set MNN hyperparameters¶
In [3]:
inputs=2 + 1 # bias input
hidden=7
outputs=2 if dataset != "blobs" and dataset != "blobs2" else 3
batch_size=10
ticks = 3
epochs = 1500
lr = 0.001
samples=1000
test_perc = 0.3
noise = 0.1
3. Load dataset¶
In [4]:
if dataset == "moons":
X, Y = datasets.make_moons(n_samples=samples, noise=noise)
elif dataset == "circles":
X, Y = datasets.make_circles(n_samples=samples, noise=noise, factor=0.5)
elif dataset == "blobs":
X, Y = datasets.make_blobs(n_samples=samples, cluster_std=[1.0, 2.5, 0.5])
elif dataset == "blobs2":
X1, Y1 = datasets.make_blobs(n_samples=samples//2)
X2, Y2 = datasets.make_blobs(n_samples=samples//2)
X = np.concatenate((X1, X2))
Y = np.concatenate((Y1, Y2))
elif dataset == "spirals":
n = np.sqrt(np.random.rand(samples,1)) * 780 * (2*np.pi)/360
d1x = -np.cos(n)*n + np.random.rand(samples,1) * noise
d1y = np.sin(n)*n + np.random.rand(samples,1) * noise
X, Y = (np.vstack((np.hstack((d1x,d1y)),np.hstack((-d1x,-d1y)))),
np.hstack((np.zeros(samples),np.ones(samples))))
else:
raise Exception("Unknown dataset")
pivot = int(X.shape[0]*(1-test_perc))
train_X = X[:pivot]
train_Y = Y[:pivot]
test_X = X[pivot:]
test_Y = Y[pivot:]
In [5]:
def f(x, derivative=False):
gt = (x > 0)
if derivative:
return 1 * gt
else:
return x * gt
4.2 CrossEntropy loss function and its derivative¶
In [6]:
def softmax(x):
exps = np.exp(x - np.max(x))
return exps / np.expand_dims(np.sum(exps, axis=1), axis=1)
def ce_loss(out, y, grad):
y = y.astype(int)
m = y.shape[0]
p = softmax(out)
log_likelihood = -np.log(p[range(m),y])
loss = np.mean(log_likelihood, axis=0)
de = p
de[range(m),y] -= 1
de = de/m
de = np.expand_dims(np.expand_dims(de, axis=2), axis=2)
grad = de*grad
return (loss, grad.sum(axis=1).sum(axis=0))
4.3 Adam Optimizer¶
In [7]:
class Adam:
def __init__(self, lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-8, decay=0., **kwargs):
allowed_kwargs = {'clipnorm', 'clipvalue'}
for k in kwargs:
if k not in allowed_kwargs:
raise TypeError('Unexpected keyword argument '
'passed to optimizer: ' + str(k))
self.__dict__.update(kwargs)
self.iterations = 0
self.lr = lr
self.beta_1 = beta_1
self.beta_2 = beta_2
self.decay = decay
self.epsilon = epsilon
self.initial_decay = decay
def step(self, params, grads):
original_shapes = [x.shape for x in params]
params = [x.flatten() for x in params]
grads = [x.flatten() for x in grads]
lr = self.lr
if self.initial_decay > 0:
lr *= (1. / (1. + self.decay * self.iterations))
t = self.iterations + 1
lr_t = lr * (np.sqrt(1. - np.power(self.beta_2, t)) /
(1. - np.power(self.beta_1, t)))
if not hasattr(self, 'ms'):
self.ms = [np.zeros(p.shape) for p in params]
self.vs = [np.zeros(p.shape) for p in params]
ret = [None] * len(params)
for i, p, g, m, v in zip(range(len(params)), params, grads, self.ms, self.vs):
m_t = (self.beta_1 * m) + (1. - self.beta_1) * g
v_t = (self.beta_2 * v) + (1. - self.beta_2) * np.square(g)
p_t = p - lr_t * m_t / (np.sqrt(v_t) + self.epsilon)
self.ms[i] = m_t
self.vs[i] = v_t
ret[i] = p_t
self.iterations += 1
for i in range(len(ret)):
ret[i] = ret[i].reshape(original_shapes[i])
return ret
4.4 Plotting¶
In [8]:
def make_meshgrid(x, y, h=.02):
x_min, x_max = x.min() - 1, x.max() + 1
y_min, y_max = y.min() - 1, y.max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
return xx, yy
def plot_contours(net, xx, yy, **params):
input = np.c_[xx.ravel(), yy.ravel()]
bs = input.shape[0]
state = np.zeros(shape=(bs, neurons)) # Init state
grad = np.zeros(shape=(bs, neurons, neurons, neurons)) # Init gradients
for t in range(0, ticks):
state, _ = net(state, grad, input, bs=bs, compute_grad=False)
outs = state[:,inputs+hidden:neurons]
Z = np.argmax(outs, axis=1).astype(int)
Z = Z.reshape(xx.shape)
out = plt.contourf(xx, yy, Z, **params)
return out
def fig2data(fig):
fig.canvas.draw()
w,h = fig.canvas.get_width_height()
buf = np.frombuffer(fig.canvas.tostring_argb(), dtype=np.uint8)
buf.shape = (w, h, 4)
buf = np.roll(buf, 3, axis = 2)
return buf
def fig2img(fig):
buf = fig2data(fig)
w, h, d = buf.shape
return Image.frombytes("RGBA",(w ,h), buf.tostring())
def get_figure():
fig = plt.figure()
X0, X1 = train_X[:, 0], train_X[:, 1]
xx, yy = make_meshgrid(X0, X1)
plot_contours(net, xx, yy, cmap=plt.cm.coolwarm, alpha=0.8)
plt.scatter(X0, X1, c=train_Y, cmap=plt.cm.coolwarm, s=20, edgecolors='k')
ret = fig2img(fig)
plt.close(fig)
return ret
In [9]:
def derivate(grad, t):
sn = np.zeros(shape=grad.shape)
sn[:,np.eye(neurons).astype(bool)] = state[:,np.newaxis]
sn = np.transpose(sn, (0,3,2,1))
return f(t, derivative=True)[:, np.newaxis, np.newaxis] * (np.matmul(grad, A) + sn)
5.2 State update function as described in Section 2.1 of the paper¶
In [10]:
def net(state, grad, x, bs=batch_size, compute_grad=True):
# set inputs neurons state with input vector
state[:,0:inputs] = np.concatenate((x, np.ones((bs,1))), axis=1)
# compute ti
t = np.matmul(state, A)
# Forward propagate the gradients
grad = derivate(grad, t) if compute_grad else None
# Compute new state
state = f(t)
return state, grad
5.3 Learning algorithm as described in Section 2.2.1 of the paper¶
In [11]:
neurons = inputs + hidden + outputs
A = np.random.rand(neurons, neurons) # Adjacency Matrix
optimizer = Adam(lr=lr)
frames = []
for epoch in range(0, epochs):
losses = 0
for i in range(0, len(train_X)//batch_size):
# Training batches
batch_X = train_X[i*batch_size:(i+1)*batch_size]
batch_Y = train_Y[i*batch_size:(i+1)*batch_size]
state = np.zeros(shape=(batch_size, neurons)) # Init state
grad = np.zeros(shape=(batch_size, neurons, neurons, neurons)) # Init gradients
# Update MNN in time
for t in range(0, ticks):
state, grad = net(state, grad, batch_X)
# Permute gradients for error function
grad = np.transpose(grad, (0,3,1,2))
# Slice output values and gradients
outs = state[:,inputs+hidden:neurons]
outs_grad = grad[:,inputs+hidden:neurons]
# Compute loss and error gradients
loss, err_grad = ce_loss(outs, batch_Y, outs_grad)
losses += loss
# Update weights
A = optimizer.step(A, err_grad)
if epoch % 50 == 0:
print("epoch: %d loss: %f" % (epoch, losses/(len(train_X)//(batch_size))))
frames.append(get_figure())
frames[0].save(dataset+'_training.gif', format='GIF', append_images=frames[1:],
save_all=True, duration=100, loop=0)
IPython.display.Image(dataset+"_training.gif")
epoch: 0 loss: 5.085141 epoch: 50 loss: 0.616959 epoch: 100 loss: 0.583152 epoch: 150 loss: 0.548342 epoch: 200 loss: 0.343124 epoch: 250 loss: 0.197952 epoch: 300 loss: 0.161359 epoch: 350 loss: 0.121376 epoch: 400 loss: 0.095786 epoch: 450 loss: 0.064127 epoch: 500 loss: 0.043824 epoch: 550 loss: 0.034114 epoch: 600 loss: 0.026477 epoch: 650 loss: 0.023128 epoch: 700 loss: 0.022933 epoch: 750 loss: 0.017043 epoch: 800 loss: 0.017234 epoch: 850 loss: 0.012081 epoch: 900 loss: 0.281040 epoch: 950 loss: 0.009276 epoch: 1000 loss: 0.006655 epoch: 1050 loss: 0.490068 epoch: 1100 loss: 0.002726 epoch: 1150 loss: 0.002117 epoch: 1200 loss: 0.001836 epoch: 1250 loss: 0.003380 epoch: 1300 loss: 0.002132 epoch: 1350 loss: 0.055572 epoch: 1400 loss: 0.000964 epoch: 1450 loss: 0.000897
Out[11]:
<IPython.core.display.Image object>
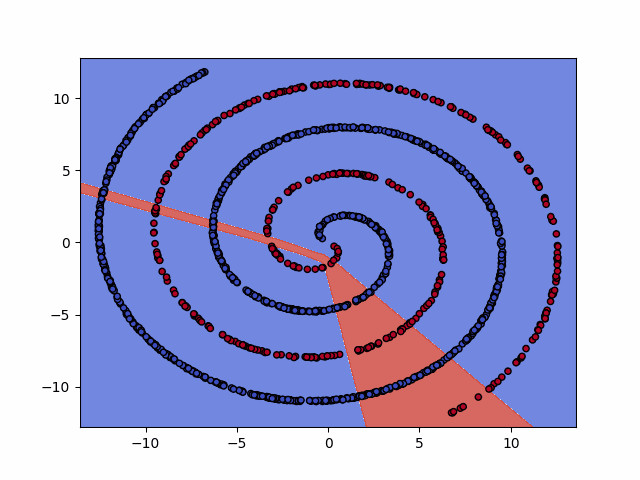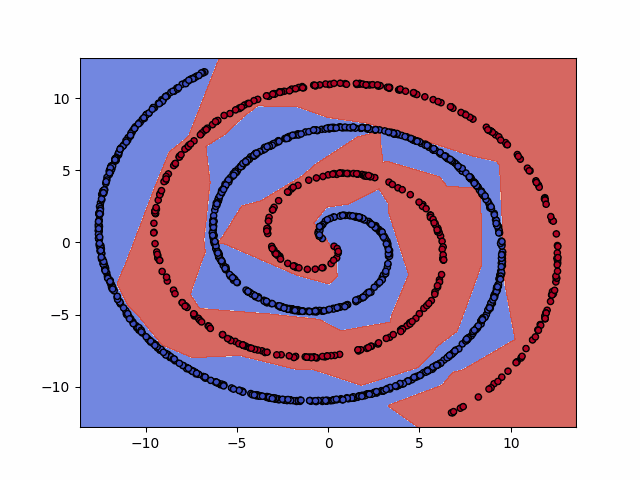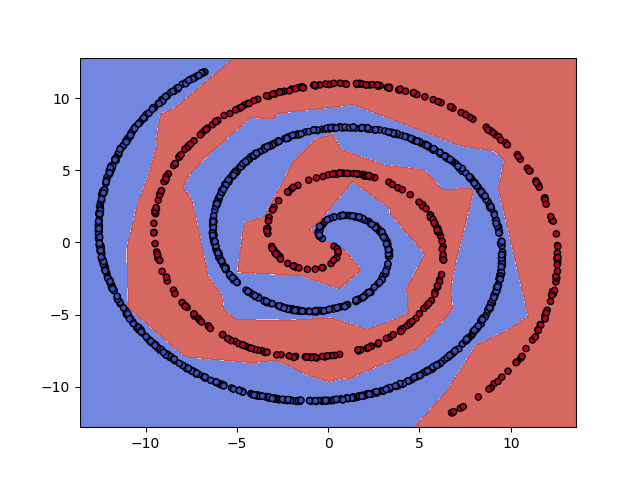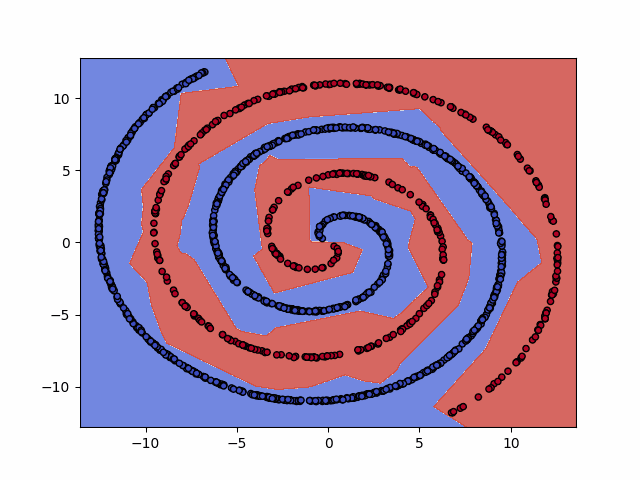
6. Test the model¶
In [12]:
rights = 0
tots = 0
for i in range(0, len(test_X)//batch_size):
# Testing batches
batch_X = test_X[i*batch_size:(i+1)*batch_size]
batch_Y = test_Y[i*batch_size:(i+1)*batch_size]
# Init MNN
state = np.zeros(shape=(batch_size, neurons))
grad = np.zeros(shape=(batch_size, neurons, neurons, neurons))
for t in range(0, ticks):
state, grad = net(state, grad, batch_X)
outs = state[:,inputs+hidden:neurons]
rights += (np.argmax(outs, axis=1) == batch_Y).astype(int).sum()
tots += batch_size
print("Accuracy: %f" % (rights/tots) )
Accuracy: 1.000000