


Trees - for BHSA data (Hebrew)¶
About¶
This notebook composes syntax trees out of the BHSA dataset of the Hebrew Bible, its text and it linguistic annotations.
The source data is the text-fabric representation of this dataset.
The result is a set of roughly 65,000 tree structures, one for each sentence, in Penn Treebank notation, like this
(S (NP (NNP John))
(VP (VPZ loves)
(NP (NNP Mary)))
(. .))
but not necessarily adhering to any known tag set. Note that the tags generated by this notebook can be can be customized.
Trees in this format can be searched by e.g. tgrep.
We create a new feature, tree, that holds these the tree structure for each sentence.
In this way the trees are readily available when you need them in your Text-Fabric workflow.
See the example notebook.
This notebook can also be used to generate text files with all trees in it. For that you have to run the notebook on your own computer.
BHSA data and syntax trees¶
The process of tree construction is not straightforward,
since the BHSA data have not been coded as syntax trees.
Rather they take the shape of a collection of features that describe
observable characteristics of the words, phrases, clauses and sentences.
Moreover, if a phrase, clause or sentence is discontinuous,
it is divided in phrase_atoms, clause_atoms,
or sentence_atoms, respectively, which are by definition continuous.
There are no explicit hierarchical relationships between these objects, or rather nodes. But there is an implicit hierarchy: embedding. Every node is linked to the set of word nodes, these are the words that are "contained" in the node. This induces a containment relationship on the set of nodes.
This notebook makes use of a Python module tree.py (in the same directory).
This module works on top of Text-Fabric and knows the general structure of an ancient text.
It constructs a hierarchy of words, subphrases, phrases, clauses and sentences
based on the embedding relationship.
But this is not all.
The BHSA data contains a mother relationship,
which denotes linguistic dependency.
The module trees.py reconstructs the tree obtained from the embedding relationship
by using the mother relationship as a set of instructions to move certain nodes below others.
In some cases extra nodes will be constructed as well.
The embedding relationship¶
Nodes:¶
The BHSA data is coded in such a way that every node is associated with a type and a slot set.
The type of a node, $T(O)$, determines which features a node has.
BHSA types are sentence, sentence_atom,
clause, clause_atom, phrase, phrase_atom, subphrase, word, and there are also
the non-linguistic types book, chapter, verse and half_verse.
There is an implicit ordering of node types, given by the sequence above, where word comes first and
sentence comes last. We denote this ordering by $<$.
The slot set of a node, $m(O)$, is the set of word occurrences linked to that node. Every word occurrence in the source occupies a unique slot, which is a number, so slot sets are sets of numbers. Think of the slots as the textual positions of individual words throughout the whole text.
Note that when a sentence contains a clause which contains a phrase, the sentence, clause, and phrase are linked to slot sets that contain each other. The fact that a sentence "contains" a clause is not marked directly, it is a consequence of how the slot sets they are linked to are embedded.
Definition (slot set order):¶
There is a natural order on slot sets, which we will use.
We will not base our trees on all node types,
since in the BHSA data they do not constitute a single hierarchy.
We will restrict ourselves to the set $\cal O = \{$ sentence, clause, phrase, word $\}$.
Definition (directly below):¶
Node type $T_1$ is directly below $T_2$ ( $T_1 <_1 T_2 $ ) in $\cal O$ if $T_1 < T_2$ and there is no $T$ in $\cal O$ with $T_1 < T < T_2$.
Now we can introduce the notion of (tree) parent with respect to a set of node types $\cal O$ (e.g. ):
Definition (parent)¶
Node $A$ is a parent of node $B$ if the following are true:
- $m(A) \subseteq\ m(B)$
- $T(A) <_1 T(B)$ in $\cal O$.
The mother relationship¶
While using the embedding got us trees,
using the mother relationship will give us more interesting trees.
In general, the mother in the BHSA dataset points to a node
on which the node in question is, in some sense, dependent.
The nature of this dependency is coded in a specific feature on clauses,
the clause_constituent_relation in version 3,
or rela in later versions.
Later in this notebook we'll show a frequency list of the values.
Here is a description of what we do with the mother relationship.
If a clause has a mother, there are three cases for the rela value of this clause:
- its value is in $\{$
Adju,Objc,Subj,PrAd,PreC,Cmpl,Attr,RgRc,Spec$\}$ - its value is
Coor - its value is in $\{$
Resu,ReVo,none$\}$
In case 3 we do nothing.
In case 1 we remove the link of the clause to its parent and add the clause as a child to either the node that the mother points to, or to the parent of the mother. We do the latter only if the mother is a word. We will not add children to words.
In the diagrams, the red arrows represent the mother relationship, and the black arrows the embedding relationships, and the fat black arrows the new parent relationships. The gray arrows indicated severed parent links.
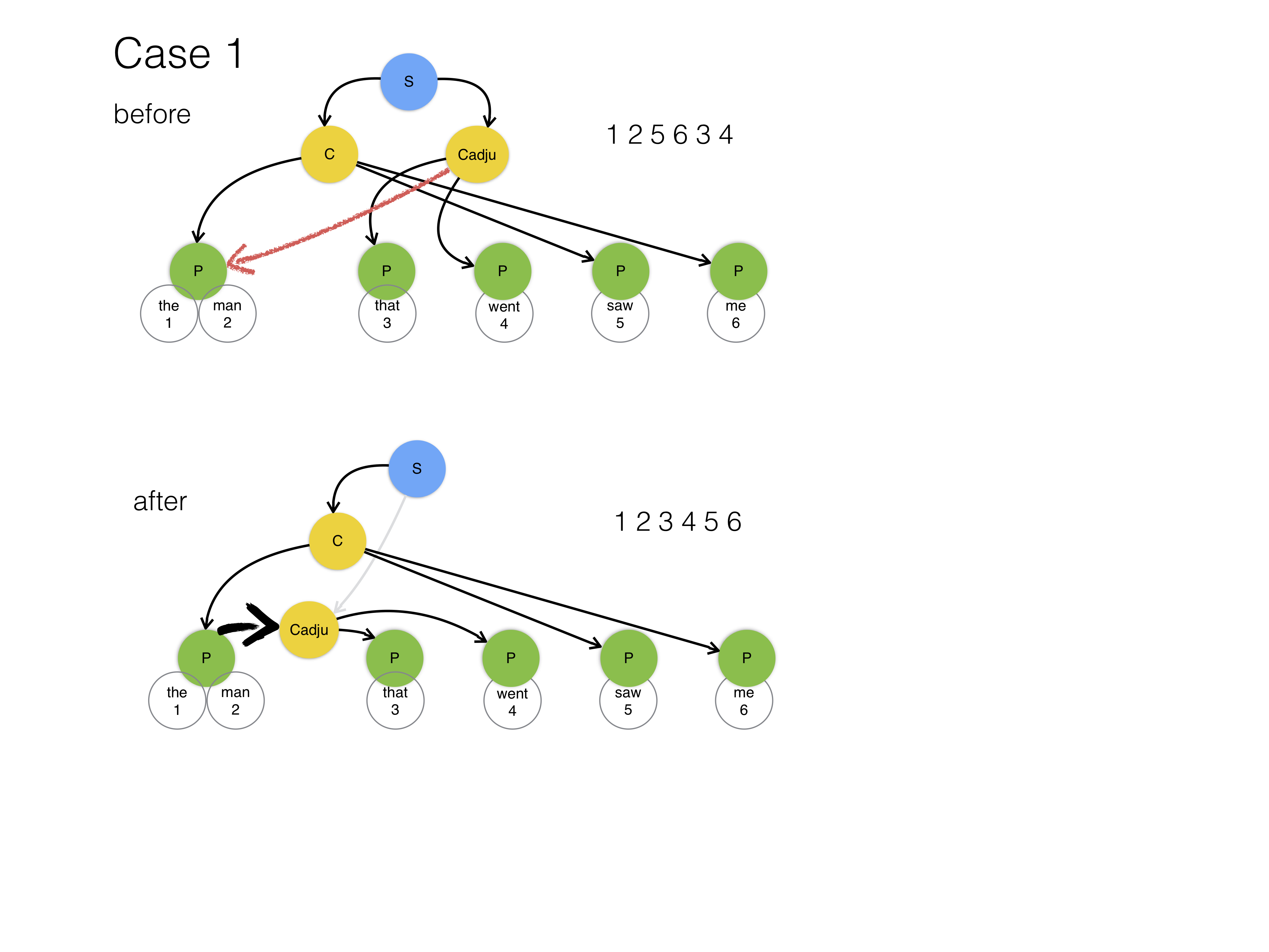
In case 2 we create a node between the mother and its parent.
This node takes the name of the mother, and the mother will be added as child,
but with name Ccoor, and the clause which points to the mother is added as a sister.
This is a rather complicated case, but the intuition is not that difficult. Consider the sentence:
John thinks that Mary said it and did it
We have a compound object sentence, with Mary said it and did it as coordinated components.
The way this has been marked up in the BHSA database is as follows:
Mary said it, clause with clause_constituent_relation=Objc, mother=John thinks(clause)
and did it, clause with clause_constituent_relation=Coor, mother=Mary said it(clause)
So the second coordinated clause is simply linked to the first coordinated clause. Restructuring means to create a parent for both coordinated clauses and treat both as sisters at the same hierarchical level. See the diagram.

Note on order¶
When we add nodes to new parents, we let them occupy the sequential position among its new sisters that corresponds with the slot set ordering.
Note on discontinuity¶
Sentences, clauses and phrases are not always continuous. Before restructuring it will not always be the case that if you walk the tree in pre-order, you will end up with the leaves (the words) in the same order as the original sentence. Restructuring generally improves that, because it often puts a node under a non-continuous parent object precisely at the location that corresponds with the a gap in the parent.
However, there is no guarantee that every discontinuity will be resolved in this graceful manner. When we create the trees, we also output the list of slot numbers that you get when you walk the tree in pre-order. Whenever this list is not monotonic, there is an issue with the ordering.
Note on cycles¶
If a mother points to itself or a descendant of itself, we have a cycle in the mother relationship. In these cases, the restructuring algorithm will disconnect a parent link without introducing a new link to the tree above it: a whole fragment of the tree becomes disconnected and will get lost.
Sanity check 6 below reveals that this occurs in fact 4 times in the BHSA version 4 (it occurred 13 times in the BHSA 3 version). We will exclude these trees from further processing.
Note on stretch¶
If a mother points outside the sentence of the clause on which it is specified we have a case of stretch. This should not happen. Mothers may point outside their sentences, but not in the cases that trigger restructuring. Yet, the sanity checks below reveal that this does occur in some versions. We will exclude these cases from further processing.
Customization¶
There are several levels of customization.
If you want to apply this notebook to other resources than the BHSA, you can specify the names of the relevant node types and features that the tree algorithm may uses for structuring the trees and restructuring them (if needed).
If you want to change the tags of the nodes in the output, you can write a new function getTag()
and pass that to the tree algorithm. See below.
%load_ext autoreload
%autoreload 2
import sys
import os
import collections
import yaml
from tf.fabric import Fabric
from tf.core.helpers import formatMeta
from tree import Tree
import utils
if "SCRIPT" not in locals():
SCRIPT = False
FORCE = True
CORE_NAME = "bhsa"
NAME = "trees"
VERSION = "2021"
def stop(good=False):
if SCRIPT:
sys.exit(0 if good else 1)
This notebook can run a lot of tests and create a lot of examples.
However, when run in the pipeline, we only want to create the two tree features.
So, further on, there will be quite a bit of code under the condition not SCRIPT.
Setting up the context: source file and target directories¶
The conversion is executed in an environment of directories, so that sources, temp files and results are in convenient places and do not have to be shifted around.
repoBase = os.path.expanduser("~/github/etcbc")
coreRepo = "{}/{}".format(repoBase, CORE_NAME)
thisRepo = "{}/{}".format(repoBase, NAME)
coreTf = "{}/tf/{}".format(coreRepo, VERSION)
thisTemp = "{}/_temp/{}".format(thisRepo, VERSION)
thisTempTf = "{}/tf".format(thisTemp)
thisTf = "{}/tf/{}".format(thisRepo, VERSION)
Test¶
Check whether this conversion is needed in the first place. Only when run as a script.
if SCRIPT:
(good, work) = utils.mustRun(
None, "{}/.tf/{}.tfx".format(thisTf, "tree"), force=FORCE
)
if not good:
stop(good=False)
if not work:
stop(good=True)
Load the TF data¶
utils.caption(4, "Load the existing TF dataset")
TF = Fabric(locations=coreTf, modules=[""])
.............................................................................................. . 0.00s Load the existing TF dataset . .............................................................................................. This is Text-Fabric 9.1.7 Api reference : https://annotation.github.io/text-fabric/tf/cheatsheet.html 114 features found and 0 ignored
Load data¶
We load the some features of the BHSA data. See the feature documentation for more info.
sp = "part_of_speech" if VERSION == "3" else "sp"
rela = "clause_constituent_relation" if VERSION == "3" else "rela"
ptyp = "phrase_type" if VERSION == "3" else "typ"
ctyp = "clause_atom_type" if VERSION == "3" else "typ"
g_word_utf8 = "text" if VERSION == "3" else "g_word_utf8"
# -
api = TF.load(
f"""
{sp} {rela} {ptyp} {ctyp}
{g_word_utf8}
mother
"""
)
api.makeAvailableIn(globals())
0.00s loading features ...
| 0.00s Dataset without structure sections in otext:no structure functions in the T-API
11s All features loaded/computed - for details use TF.isLoaded()
[('Computed',
'computed-data',
('C Computed', 'Call AllComputeds', 'Cs ComputedString')),
('Features', 'edge-features', ('E Edge', 'Eall AllEdges', 'Es EdgeString')),
('Fabric', 'loading', ('TF',)),
('Locality', 'locality', ('L Locality',)),
('Nodes', 'navigating-nodes', ('N Nodes',)),
('Features',
'node-features',
('F Feature', 'Fall AllFeatures', 'Fs FeatureString')),
('Search', 'search', ('S Search',)),
('Text', 'text', ('T Text',))]
We are going to make convenient labels for constituents, words and clauses, based on the
the types of textual objects and the features
sp and rela.
Node types¶
typeInfo = (
("word", ""),
("subphrase", "U"),
("phrase", "P"),
("clause", "C"),
("sentence", "S"),
)
typeTable = dict(t for t in typeInfo)
typeOrder = [t[0] for t in typeInfo]
Part of speech¶
sorted(Fs(sp).freqList(), key=lambda x: x[0])
[('adjv', 10141),
('advb', 4603),
('art', 30387),
('conj', 62737),
('inrg', 1303),
('intj', 1912),
('nega', 6059),
('nmpr', 35607),
('prde', 2678),
('prep', 73298),
('prin', 1026),
('prps', 5035),
('subs', 125583),
('verb', 75451)]
posTable = {
"adjv": "aj",
"adjective": "aj",
"advb": "av",
"adverb": "av",
"art": "dt",
"article": "dt",
"conj": "cj",
"conjunction": "cj",
"inrg": "ir",
"interrogative": "ir",
"intj": "ij",
"interjection": "ij",
"nega": "ng",
"negative": "ng",
"nmpr": "n-pr",
"pronoun": "pr",
"prde": "pr-dem",
"prep": "pp",
"preposition": "pp",
"prin": "pr-int",
"prps": "pr-ps",
"subs": "n",
"noun": "n",
"verb": "vb",
}
rela¶
sorted(Fs(rela).freqList(), key=lambda x: x[0])
[('Adju', 6426),
('Appo', 5884),
('Attr', 5811),
('Cmpl', 298),
('Coor', 3660),
('Link', 1317),
('NA', 630059),
('Objc', 1327),
('Para', 1559),
('PrAd', 308),
('PreC', 162),
('ReVo', 312),
('Resu', 1666),
('RgRc', 856),
('Sfxs', 75),
('Spec', 5565),
('Subj', 506),
('adj', 4138),
('atr', 3064),
('dem', 1847),
('mod', 941),
('par', 11946),
('rec', 34989)]
ccrInfo = {
"Adju": ("r", "Cadju"),
"Appo": ("r", "Cappo"),
"Attr": ("r", "Cattr"),
"Cmpl": ("r", "Ccmpl"),
"Coor": ("x", "Ccoor"),
"CoVo": ("n", "Ccovo"),
"Link": ("r", "Clink"),
"Objc": ("r", "Cobjc"),
"Para": ("r", "Cpara"),
"PrAd": ("r", "Cprad"),
"PreC": ("r", "Cprec"),
"Pred": ("r", "Cpred"),
"ReVo": ("n", "Crevo"),
"Resu": ("n", "Cresu"),
"RgRc": ("r", "Crgrc"),
"Sfxs": ("r", "Csfxs"),
"Spec": ("r", "Cspec"),
"Subj": ("r", "Csubj"),
"NA": ("n", "C"),
"none": ("n", "C"),
}
treeTypes = ("sentence", "clause", "phrase", "subphrase", "word")
(rootType, leafType, clauseType, phraseType) = (
treeTypes[0],
treeTypes[-1],
treeTypes[1],
treeTypes[2],
)
ccrTable = dict((c[0], c[1][1]) for c in ccrInfo.items())
ccrClass = dict((c[0], c[1][0]) for c in ccrInfo.items())
Now we can actually construct the tree by initializing a tree object.
After that we call its restructureClauses() method.
Then we have two tree structures for each sentence:
- the
etree, i.e. the tree obtained by working out the embedding relationships and nothing else - the
rtree, i.e. the tree obtained by restructuring theetree
We have several tree relationships at our disposal:
eparentand its inverseechildrenrparentand its inverserchildreneldest_sistergoing from a mother clause of kindCoor(case 2) to its daughter clauses- sisters being the inverse of
eldest_sister
where eldest_sister and sisters only occur in the rtree.
This will take a while (25 seconds approx on a MacBook Air 2012, 6 on a MacBook Pro in 2017, and now just 4 seconds).
tree = Tree(
TF,
otypes=treeTypes,
phraseType=phraseType,
clauseType=clauseType,
ccrFeature=rela,
ptFeature=ptyp,
posFeature=sp,
motherFeature="mother",
)
0.00s loading features ... 0.11s All additional features loaded - for details use TF.isLoaded() 0.11s Start computing parent and children relations for objects of type sentence, clause, phrase, subphrase, word 0.41s 100000 nodes 0.69s 200000 nodes 2.37s 300000 nodes 2.66s 400000 nodes 2.93s 500000 nodes 3.21s 600000 nodes 3.50s 700000 nodes 3.79s 800000 nodes 4.07s 900000 nodes 4.19s 945491 nodes: 881774 have parents and 518901 have children
tree.restructureClauses(ccrClass)
results = tree.relations()
parent = results["rparent"]
sisters = results["sisters"]
children = results["rchildren"]
elderSister = results["elderSister"]
utils.caption(4, "Ready for processing")
4.20s Restructuring clauses: deep copying tree relations
5.91s Pass 0: Storing mother relationship
5.96s 20791 clauses have a mother
5.96s All clauses have mothers of types in {'word', 'subphrase', 'clause', 'phrase', 'sentence'}
5.96s Pass 1: all clauses except those of type Coor
6.01s Pass 2: clauses of type Coor only
6.04s Mothers applied. Found 0 motherless clauses.
6.04s 3233 nodes have 1 sisters
6.04s 200 nodes have 2 sisters
6.04s 9 nodes have 3 sisters
6.04s There are 3660 sisters, 3442 nodes have sisters.
..............................................................................................
. 17s Ready for processing .
..............................................................................................
# # Sanity check
# 6
# If there are blocking errors we collect the nodes of those sentences in the set
# `skip`, so that later on we can skip them easily.
utils.caption(4, "Verifying whether all slots are preserved under restructuring")
expectedMismatches = {
"3": 13,
"4": 3,
"4b": 0,
"2016": 0,
"2017": 0,
}
utils.caption(4, "Expected mismatches: {}".format(expectedMismatches.get(VERSION, "??")))
.............................................................................................. . 17s Verifying whether all slots are preserved under restructuring . .............................................................................................. .............................................................................................. . 17s Expected mismatches: ?? . ..............................................................................................
skip = set()
errors = []
for snode in F.otype.s(rootType):
declaredSlots = set(E.oslots.s(snode))
results = {}
thisgood = {}
for kind in ("e", "r"):
results[kind] = set(
lf for lf in tree.getLeaves(snode, kind) if F.otype.v(lf) == leafType
)
thisgood[kind] = declaredSlots == results[kind]
# if not thisgood[kind]:
# print('{} D={}\n L={}'.format(kind, declaredSlots, results[kind]))
# i -= 1
# if i == 0: break
if False in thisgood.values():
errors.append((snode, thisgood["e"], thisgood["r"]))
nErrors = len(errors)
if nErrors:
utils.caption(4, "{} mismatches:".format(len(errors)), good=False)
mine = min(20, len(errors))
skip |= {e[0] for e in errors}
for (s, e, r) in errors[0:mine]:
utils.caption(
4,
"{} embedding: {}; restructd: {}".format(
s, "OK" if e else "XX", "OK" if r else "XX"
),
good=False,
)
else:
utils.caption(4, "0 mismatches")
.............................................................................................. . 19s 0 mismatches . ..............................................................................................
Delivering Trees¶
We are going to deliver the trees as a feature of sentence nodes.
There will be two features in fact:
tree: minimalistictreen: with node information of non-slot nodes
In order to produce this, we need to write appropriate functions to pass to writeTree: getTag() and getTagN().
getTag(node)¶
This function produces for each node
- a tag string,
- a part-of-speech representation,
- a textual position (slot number),
- a boolean which tells if this node is a leaf or not.
This function will be passed to the writeTree() function in the tree module.
By supplying a different function, you can control a lot of the characteristics of the
written tree.
def getTag(node):
otype = F.otype.v(node)
tag = typeTable[otype]
if tag == "P":
tag = Fs(ptyp).v(node)
elif tag == "C":
tag = ccrTable[Fs(rela).v(node)]
isWord = tag == ""
pos = posTable[Fs(sp).v(node)] if isWord else None
slot = node if isWord else None
text = '"{}"'.format(Fs(g_word_utf8).v(node)) if isWord else None
return (tag, pos, slot, text, isWord)
This is a variant on getTag() where we put the node number into the tag, between { }.
def getTagN(node):
otype = F.otype.v(node)
tag = typeTable[otype]
if tag == "P":
tag = Fs(ptyp).v(node)
elif tag == "C":
tag = ccrTable[Fs(rela).v(node)]
isWord = tag == ""
if not isWord:
tag += "{" + str(node) + "}"
pos = posTable[Fs(sp).v(node)] if isWord else None
slot = node if isWord else None
text = '"{}"'.format(Fs(g_word_utf8).v(node)) if isWord else None
return (tag, pos, slot, text, isWord)
Finally, here is the production of the whole set of trees.
Now we generate the data for two TF features tree and treen:
utils.caption(4, "Exporting {} trees to TF".format(rootType))
s = 0
chunk = 10000
sc = 0
treeData = {}
treeDataN = {}
for node in F.otype.s(rootType):
if node in skip:
continue
(treeRep, wordsRep, bSlot) = tree.writeTree(
node, "r", getTag, rev=False, leafNumbers=True
)
(treeNRep, wordsNRep, bSlotN) = tree.writeTree(
node, "r", getTagN, rev=False, leafNumbers=True
)
treeData[node] = treeRep
treeDataN[node] = treeNRep
s += 1
sc += 1
if sc == chunk:
utils.caption(0, "{} trees composed".format(s))
sc = 0
utils.caption(4, "{} trees composed".format(s))
.............................................................................................. . 19s Exporting sentence trees to TF . .............................................................................................. | 20s 10000 trees composed | 22s 20000 trees composed | 23s 30000 trees composed | 24s 40000 trees composed | 25s 50000 trees composed | 27s 60000 trees composed .............................................................................................. . 27s 63717 trees composed . ..............................................................................................
genericMetaPath = f"{thisRepo}/yaml/generic.yaml"
treesMetaPath = f"{thisRepo}/yaml/trees.yaml"
with open(genericMetaPath) as fh:
genericMeta = yaml.load(fh, Loader=yaml.FullLoader)
genericMeta["version"] = VERSION
with open(treesMetaPath) as fh:
treesMeta = formatMeta(yaml.load(fh, Loader=yaml.FullLoader))
metaData = {"": genericMeta, **treesMeta}
nodeFeatures = dict(tree=treeData, treen=treeDataN)
for f in nodeFeatures:
metaData[f]["valueType"] = "str"
utils.caption(4, "Writing tree feature to TF")
TFw = Fabric(locations=thisTempTf, silent=True)
TFw.save(nodeFeatures=nodeFeatures, edgeFeatures={}, metaData=metaData)
.............................................................................................. . 5m 01s Writing tree feature to TF . ..............................................................................................
True
Diffs¶
Check differences with previous versions.
utils.checkDiffs(thisTempTf, thisTf, only=set(nodeFeatures))
Deliver¶
Copy the new TF features from the temporary location where they have been created to their final destination.
In[38]:
utils.deliverDataset(thisTempTf, thisTf)
Compile TF¶
In[39]:
utils.caption(4, "Load and compile the new TF features")
TF = Fabric(locations=[coreTf, thisTf], modules=[""])
api = TF.load(" ".join(nodeFeatures))
api.makeAvailableIn(globals())
In[40]:
utils.caption(4, "Basic tests")
utils.caption(4, "Sample sentences in tree form")
sentences = F.otype.s("sentence")
examples = (sentences[0], sentences[len(sentences) // 2], sentences[-1])
for s in examples:
utils.caption(0, F.tree.v(s))
for s in examples:
utils.caption(0, F.treen.v(s))
End of pipeline¶
If this notebook is run with the purpose of generating data, this is the end then.
After this tests and examples are run.
In[41]:
if SCRIPT:
stop(good=True)
See the tutorial trees for how to make use of this feature.
Checking for sanity¶
Let us see whether the trees we have constructed satisfy some sanity constraints. After all, the algorithm is based on certain assumptions about the data, but are those assumptions valid? And restructuring is a tricky operation, do we have confidence that nothing went wrong?
- How many sentence nodes? From earlier queries we know what to expect.
- Does any sentence have a parent? If so, there is something wrong with our assumptions or algorithm.
- Is every top node a sentence? If not, we have material outside a sentence, which contradicts the assumptions.
- Do you reach all sentences if you go up from words? If not, some sentences do not contain words.
- Do you reach all words if you go down from sentences? If not, some words have become disconnected from their sentences.
- Do you reach the same words in reconstructed trees as in embedded trees? If not, some sentence material has got lost during the restructuring process.
- From what object types to what object types does the parent relationship link? Here we check that parents do not link object types that are too distant in the object type ranking.
- How many nodes have mothers and how many mothers can a node have? We expect at most one.
- From what object types to what object types does the mother relationship link?
- Is the mother of a clause always in the same sentence?
If not, foreign sentences will be drawn in, leading to (very) big chunks.
This may occur when we use mother relationships in cases where
relahas different values than the ones that should trigger restructuring. - Has the max/average tree depth increased after restructuring? By how much? This is meant as an indication by how much our tree structures improve in significant hierarchy when we take the mother relationship into account.
# 1
expectedSentences = {
"3": 71354,
"4": 66045,
"4b": 63586,
"2016": 63570,
"2017": 63711,
}
TF.info(
"Counting {}s ... (expecting {})".format(
rootType, expectedSentences.get(VERSION, "??")
)
)
TF.info("There are {} {}s".format(len(list(F.otype.s(rootType))), rootType))
14s Counting sentences ... (expecting 63711)
14s There are 63711 sentences
# 2
TF.info("Checking parents of {}s ... (expecting none)".format(rootType))
exceptions = set()
for node in F.otype.s(rootType):
if node in parent:
exceptions.add(node)
if len(exceptions) == 0:
TF.info("No {} has a parent".format(rootType))
else:
TF.error("{} {}s have a parent:".format(len(exceptions), rootType))
for n in sorted(exceptions):
p = parent[n]
TF.error(
"{} {} [{}] has {} parent {} [{}]".format(
rootType, n, tree.slotss(n), F.otype.v(p), p, tree.slotss(p)
)
)
15s Checking parents of sentences ... (expecting none)
15s No sentence has a parent
# 3 (again a check on #1)
TF.info("Checking the types of root nodes ... (should all be {}s)".format(rootType))
expectedTops = {
"3": 0,
"4": "3 subphrases",
"4b": 0,
"2016": 0,
"2017": 0,
}
TF.info(
"Expected roots which are non-{}s: {}".format(
rootType, expectedTops.get(VERSION, "??")
)
)
exceptions = collections.defaultdict(lambda: [])
sn = 0
for node in N.walk():
otype = F.otype.v(node)
if otype not in typeTable:
continue
if otype == rootType:
sn += 1
if node not in parent and node not in elderSister and otype != rootType:
exceptions[otype].append(node)
TF.info("{} {}s seen".format(sn, rootType))
if len(exceptions) == 0:
TF.info("All top nodes are {}s".format(rootType))
else:
TF.error("Top nodes which are not {}s:".format(rootType))
for t in sorted(exceptions):
TF.error("{}: {}x".format(t, len(exceptions[t])), tm=False)
for c in exceptions[clauseType]:
(s, st) = tree.getRoot(c, "e")
v = T.sectionFromNode(s)
TF.error(
"{}={}, {}={}={}, verse={}".format(clauseType, c, rootType, st, s, v), tm=False
)
15s Checking the types of root nodes ... (should all be sentences)
15s Expected roots which are non-sentences: 0
16s 63711 sentences seen
16s All top nodes are sentences
# 4, 5
def getTop(kind, rel, rela, multi):
seen = set()
topNodes = set()
startNodes = set(F.otype.s(kind))
nextNodes = startNodes
TF.info("Starting from {} nodes ...".format(kind))
while len(nextNodes):
newNextNodes = set()
for node in nextNodes:
if node in seen:
continue
seen.add(node)
isTop = True
if node in rel:
isTop = False
if multi:
for c in rel[node]:
newNextNodes.add(c)
else:
newNextNodes.add(rel[node])
if node in rela:
isTop = False
if multi:
for c in rela[node]:
newNextNodes.add(c)
else:
newNextNodes.add(rela[node])
if isTop:
topNodes.add(node)
nextNodes = newNextNodes
topTypes = collections.defaultdict(lambda: 0)
for t in topNodes:
topTypes[F.otype.v(t)] += 1
for t in topTypes:
TF.info(
"From {} {} nodes reached {} {} nodes".format(
len(startNodes), kind, topTypes[t], t
),
tm=False,
)
TF.info("Embedding trees")
getTop(leafType, tree.eparent, {}, False)
getTop(rootType, tree.echildren, {}, True)
TF.info("Restructd trees")
getTop(leafType, tree.rparent, tree.elderSister, False)
getTop(rootType, tree.rchildren, tree.sisters, True)
TF.info("Done")
# 7
TF.info("Which types embed which types and how often? ...")
for kind in ("e", "r"):
pLinkedTypes = collections.defaultdict(lambda: 0)
parent = tree.eparent if kind == "e" else tree.rparent
kindRep = "embedding" if kind == "e" else "restructd"
for (c, p) in parent.items():
pLinkedTypes[(F.otype.v(c), F.otype.v(p))] += 1
TF.info("Found {} parent ({}) links between types".format(len(parent), kindRep))
for lt in sorted(pLinkedTypes):
TF.info("{}: {}x".format(lt, pLinkedTypes[lt]), tm=False)
# 8
TF.info("How many mothers can nodes have? ...")
motherLen = {}
for c in N.walk():
lms = list(E.mother.f(c))
nms = len(lms)
if nms:
motherLen[c] = nms
count = collections.defaultdict(lambda: 0)
for c in tree.mother:
count[motherLen[c]] += 1
TF.info("There are {} tree nodes with a mother".format(len(tree.mother)))
for cnt in sorted(count):
TF.info(
"{} nodes have {} mother{}".format(count[cnt], cnt, "s" if cnt != 1 else ""),
tm=False,
)
# 9
TF.info("Which types have mother links to which types and how often? ...")
mLinkedTypes = collections.defaultdict(lambda: set())
for (c, m) in tree.mother.items():
ctype = F.otype.v(c)
mLinkedTypes[(ctype, Fs(rela).v(c), F.otype.v(m))].add(c)
TF.info("Found {} mother links between types".format(len(parent)))
for lt in sorted(mLinkedTypes):
TF.info("{}: {}x".format(lt, len(mLinkedTypes[lt])), tm=False)
# 10
TF.info("Counting {}s with mothers in another {}".format(clauseType, rootType))
expectedOther = {
"3": 2,
"4": 0,
"4b": 0,
"2016": 0,
"2017": 0,
}
TF.info(
"Expecting {} {}s with mothers in another {}".format(
expectedOther.get(VERSION, "??"), clauseType, rootType
)
)
exceptions = set()
for node in tree.mother:
if F.otype.v(node) not in typeTable:
continue
mNode = tree.mother[node]
sNode = tree.getRoot(node, "e")
smNode = tree.getRoot(mNode, "e")
if sNode != smNode:
exceptions.add((node, sNode, smNode))
TF.info("{} nodes have a mother in another {}".format(len(exceptions), rootType))
for (n, sn, smn) in exceptions:
TF.error(
"[{} {}]({}) occurs in {} but has mother in {}".format(
F.otype.v(n), tree.slotss(n), n, sn, smn
),
tm=False,
)
16s Embedding trees
16s Starting from word nodes ...
From 426584 word nodes reached 63711 sentence nodes
17s Starting from sentence nodes ...
From 63711 sentence nodes reached 426584 word nodes
18s Restructd trees
18s Starting from word nodes ...
From 426584 word nodes reached 63711 sentence nodes
18s Starting from sentence nodes ...
From 63711 sentence nodes reached 425419 word nodes
19s Done
# 11
TF.info("Computing lengths and depths")
nTrees = 0
rnTrees = 0
totalDepth = {"e": 0, "r": 0}
rTotalDepth = {"e": 0, "r": 0}
maxDepth = {"e": 0, "r": 0}
rMaxDepth = {"e": 0, "r": 0}
totalLength = 0
for node in F.otype.s(rootType):
nTrees += 1
totalLength += tree.length(node)
thisDepth = {}
for kind in ("e", "r"):
thisDepth[kind] = tree.depth(node, kind)
different = thisDepth["e"] != thisDepth["r"]
if different:
rnTrees += 1
for kind in ("e", "r"):
if thisDepth[kind] > maxDepth[kind]:
maxDepth[kind] = thisDepth[kind]
totalDepth[kind] += thisDepth[kind]
if different:
if thisDepth[kind] > rMaxDepth[kind]:
rMaxDepth[kind] = thisDepth[kind]
rTotalDepth[kind] += thisDepth[kind]
TF.info(
"{} trees seen, of which in {} cases restructuring makes a difference in depth".format(
nTrees,
rnTrees,
)
)
if nTrees > 0:
TF.info(
"Embedding trees: max depth = {:>2}, average depth = {:.2g}".format(
maxDepth["e"],
totalDepth["e"] / nTrees,
)
)
TF.info(
"Restructd trees: max depth = {:>2}, average depth = {:.2g}".format(
maxDepth["r"],
totalDepth["r"] / nTrees,
)
)
if rnTrees > 0:
TF.info("Statistics for cases where restructuring makes a difference:")
TF.info(
"Embedding trees: max depth = {:>2}, average depth = {:.2g}".format(
rMaxDepth["e"],
rTotalDepth["e"] / rnTrees,
)
)
TF.info(
"Restructd trees: max depth = {:>2}, average depth = {:.2g}".format(
rMaxDepth["r"],
rTotalDepth["r"] / rnTrees,
)
)
TF.info(
"Total number of leaves in the trees: {}, average number of leaves = {:.2g}".format(
totalLength,
totalLength / nTrees,
)
)
28s Computing lengths and depths
30s 63711 trees seen, of which in 10904 cases restructuring makes a difference in depth
30s Embedding trees: max depth = 9, average depth = 3.6
30s Restructd trees: max depth = 24, average depth = 3.9
30s Statistics for cases where restructuring makes a difference:
30s Embedding trees: max depth = 8, average depth = 3.7
30s Restructd trees: max depth = 24, average depth = 5.4
30s Total number of leaves in the trees: 426584, average number of leaves = 6.7