ChEn-3170: Computational Methods in Chemical Engineering Fall 2018 UMass Lowell; Prof. V. F. de Almeida 20Dec2018
Final Exam 20Dec2018 Session 801/802¶
$ \newcommand{\Amtrx}{\boldsymbol{\mathsf{A}}} \newcommand{\Bmtrx}{\boldsymbol{\mathsf{B}}} \newcommand{\Cmtrx}{\boldsymbol{\mathsf{C}}} \newcommand{\Mmtrx}{\boldsymbol{\mathsf{M}}} \newcommand{\Imtrx}{\boldsymbol{\mathsf{I}}} \newcommand{\Pmtrx}{\boldsymbol{\mathsf{P}}} \newcommand{\Lmtrx}{\boldsymbol{\mathsf{L}}} \newcommand{\Umtrx}{\boldsymbol{\mathsf{U}}} \newcommand{\Smtrx}{\boldsymbol{\mathsf{S}}} \newcommand{\xvec}{\boldsymbol{\mathsf{x}}} \newcommand{\yvec}{\boldsymbol{\mathsf{y}}} \newcommand{\zvec}{\boldsymbol{\mathsf{z}}} \newcommand{\avec}{\boldsymbol{\mathsf{a}}} \newcommand{\bvec}{\boldsymbol{\mathsf{b}}} \newcommand{\cvec}{\boldsymbol{\mathsf{c}}} \newcommand{\rvec}{\boldsymbol{\mathsf{r}}} \newcommand{\gvec}{\boldsymbol{\mathsf{g}}} \newcommand{\norm}[1]{\bigl\lVert{#1}\bigr\rVert} \DeclareMathOperator{\rank}{rank} \DeclareMathOperator{\abs}{abs} $
Name:¶
Guidance:¶
- This is a closed-book, closed-note, individual exam.
- You may use an off-line, simple calculator, that does not store documents or notes.
- You may use scratch paper if you wish (paper will be provided).
- All pages must be submitted with your completed exam.
- Make sure to answer the question asked.
- Show your work and be clear.
Rubric Panel¶
| Show No. | Now Showing (3 - 6) | Value | Score |
|---|---|---|---|
| 1-5 | Jupythonic Park | 50 | |
| 6 | Relax...Relax... | 20 | |
| 7 | Just One More Time | 20 | |
| 8 | Fast and furious | 10 | |
| Total | 100 |
Show Problem 1 (10 pts)¶
Name and give an example of the four main data structures in Python used in this course. Write your example as you would do in a Jupyter program cell below and initialize each of the data structures as empty.
'''1.1 Data structure'''
# this is a list
xmas = list() # this is an empty xmas gift list
xmas = ['gift1','gift2','gift3']
'''1.2 Data structure'''
# this is a ??? data structure
velocity =
'''1.3 Data structure'''
# this is a ??? data structure
species_molar_masses =
'''1.4 Data structure'''
# this is a ??? data structure
species =
Show Problem 2 (10 pts)¶
In each of the cells below compute the result as it would show if the cell were executed by the Python interpreter.
'''2.1 (4pts)'''
vec_a = (2,1,0)
vec_b = (8,6,7)
vec_c = vec_a + vec_b
print(vec_c)
print(len(vec_c))
'''2.2 (4pts)'''
row_1 = [2,1,6]
row_2 = [0,-1,5]
row_3 = row_1 + row_2
print(row_3)
print(len(row_3))
'''2.3 (2pts)'''
text = 'compute the result'
print(len(text))
Show Problem 3 (10 pts)¶
3.1 (2pts) Given matrices $\Amtrx$ and $\Bmtrx$ compute $\Cmtrx = \Amtrx\,\Bmtrx$. Write the simplest Python code you can imagine to performe this operation and compute and print the result. Note: $\Amtrx = \begin{pmatrix} 1 & 2 & -4 \\ 0 & 3 & 4 \\ 1 & 2 & 5 \end{pmatrix}$ and $\Bmtrx = \begin{pmatrix} 1& 7 & 3 \\ 3& 2 & 6 \\ 0& 2 & 4 \end{pmatrix}$.
'''3.1 (2pts) Code and result'''
import numpy as np
a_mtrx = np.array([???])
b_mtrx = np.array([???])
c_mtrx = a_mtrx ? b_mtrx
print(c_mtrx)
3.2 (4pts) Given matrix $\Amtrx$ and vector $\bvec$ compute the solution $\xvec$ of $\Amtrx\,\xvec=\bvec$ and write the simplest possible Python code to compute and print the result. Note: $\Amtrx = \begin{pmatrix} 1 & 0 & 0 \\ 1/2 & 1 & 0 \\ -1/2 & 1 & 1 \end{pmatrix}$ and $\bvec = \begin{pmatrix} 1 \\ 3 \\ 2 \end{pmatrix}$.
'''3.2 (4pts) Code and result'''
import numpy as np
a_mtrx = np.array([???])
b_vec = np.array([???])
x_vec = np.linalg.?(?,?)
print(x_vec)
3.3 (4pts) Given matrix $\Amtrx$ and vector $\bvec$ compute the solution $\xvec$ of $\Amtrx\,\xvec=\bvec$ and write the simplest possible Python code to compute and print the result. Note: $\Amtrx = \begin{pmatrix} 2 & 3 & 1 \\ 0 & 1/2 & 1/2 \\ 0 & 0 & 1 \end{pmatrix}$ and $\bvec = \begin{pmatrix} 1 \\ 5/2 \\ 0 \end{pmatrix}$.
'''3.3 (4pts) Code and result'''
import numpy as np
a_mtrx = np.array([???])
b_vec = np.array([???])
x_vec = np.linalg.?(?,?)
print(x_vec)
Show Problem 4 (10 pts)¶
a) (5pts) Given matrix $\Amtrx$ compute the solution $\Amtrx\,\xvec=\bvec$. Write the simplest Python code you can imagine to performe this operation and compute and print the result. Note: $\Amtrx = \begin{pmatrix} 2 & 3 & 1 \\ 1 & 2 & 1 \\ -1 & -1 & 1 \end{pmatrix}$ and $\bvec = \begin{pmatrix} 1 \\ 3 \\ 2 \end{pmatrix}$.
'''4.1 Solve A x = b'''
import numpy as np
a_mtrx = np.array([???])
b_vec = np.array([???])
x_vec = np.linalg.?(?,?)
print(x_vec)
b) (5pts) Given matrix $\Amtrx$ compute the solution $\Amtrx\,\xvec=\bvec$. Write the simplest Python code you can imagine to performe this operation and compute and print the result. Note: $\Amtrx = \begin{pmatrix} 2 & 3 & 1 \\ 1 & 2 & 1 \\ -1 & -1 & 0 \end{pmatrix}$ and $\bvec = \begin{pmatrix} 1 \\ 3 \\ 2 \end{pmatrix}$.
'''4.2 Solve A x = b'''
import numpy as np
a_mtrx = np.array([???])
b_vec = np.array([???])
x_vec = np.linalg.?(?,?)
print(x_vec)
Show Problem 5 (10 pts)¶
Given the following reaction mechanism for hydrogen bromide:
\begin{align*} \text{H}_2 + \text{Br}_2 &\longleftrightarrow 2\,\text{HBr} \\ \text{Br}_2 &\longleftrightarrow 2\,\text{Br} \\ \text{Br} + \text{H}_2 &\longleftrightarrow \text{HBr} + \text{H} \\ \text{H} + \text{Br}_2 &\longleftrightarrow \text{HBr} + \text{Br} \\ \text{H} + \text{HBr} &\longleftrightarrow \text{H}_2 + \text{Br} \\ 2\,\text{Br} &\longleftrightarrow \text{Br}_2 \end{align*}write a simple code to compute the number of independent reactions and compute this number.
'''5. Independent reactions'''
import numpy as np
stoic_mtrx = np.array([???])
num_independent_rxn = np.linalg.?(?)
print(num_independent_rxn)
Show Problem 6 (20 pts)¶
A concentration signal is believed to follow the model
\begin{equation*} c(t) = a + b\,e^{-t/\tau_1} + c\,e^{-t/\tau_2} \end{equation*}which has two modes of relaxation with relaxation times, $\tau_1=2$ s, and $\tau_2=4$ s. Given the scarce measurements of $c(t)$ below:
- Calculate the best values of $a,b,c$ that fit the model.
- Calculate the residual of your best approximation.
- Is this a good approximation for the data? (hint: calculate $c(1)$ and/or graph the approximant $c(t)$ on the plot)
- Explain why or why not the approximation is good and the conditions that control the goodness of the approximation.
'''Data'''
'''6.1 Calculate optimum a, b, c'''
This show problem does not ask for coding.
- Build the matrix problem
\begin{align*} c(0) = 2.0 &= a + b\,e^{-0/\tau_1} + c\,e^{-0/\tau_2}\\ c(6) = 1.5 &= a + b\,e^{-6/\tau_1} + c\,e^{-6/\tau_2}\\ c(12) = 1 &= a + b\,e^{-12/\tau_1} + c\,e^{-12/\tau_2}\\ \end{align*}
$\Amtrx = $?
$ \bvec = \begin{pmatrix} 2 \\ 1.5 \\ 1 \end{pmatrix} $
$ \xvec = \begin{pmatrix} a \\ b \\ c \end{pmatrix}. $
$\Amtrx\,\xvec = \bvec$
- Solve for the coefficients
'''6.2 Calculate the residual'''
This show problem does not ask for coding.
- Calculate the residual of the matrix problem
'''6.3 Is this a good approximation of the data?'''
This show problem does not ask for coding.
Hint: calculate $c(1)$ and/or graph the approximant $c(t)$ on the plot
'''6.4 Explain why or why not the approximation is good and the conditions that control the goodness of the approximation'''
This show problem does not ask for coding.
Show Problem 7 (20 pts)¶
A reversible first-order endothermic reaction
\begin{equation*} \text{A} \underset{k_\text{b}}{\underset{\leftarrow}{\overset{k_\text{f}}{\longrightarrow}}} 2\,\text{B} , \end{equation*}is performed in a continuous-stirred-tank reactor.
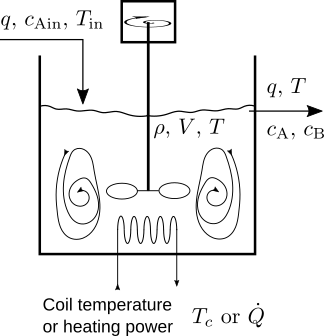
The experimental forward rate of reaction per unit volume is
\begin{equation*} r_{\text{f},1} = k_\text{f}\,c_\text{A}\,c_\text{B}^0 , \end{equation*}and the reverse is
\begin{equation*} r_{\text{b},1} = \lambda\,k_\text{f} \,c_\text{A}^0\,c^\beta_\text{B} . \end{equation*}The governing equations for the time variation of the concentrations of species A and B are:
\begin{equation*} \frac{dc_\text{A}}{dt} = -\frac{1}{\tau}\bigl(c_\text{A}-c_\text{Ain}\bigr) + g_\text{A}(t) ,\\ \frac{dc_\text{B}}{dt} = -\frac{1}{\tau}\bigl(c_\text{B}-c_\text{Bin}\bigr) + g_\text{B}(t) , \end{equation*}where $\tau = \frac{V}{q} = \frac{V\,\rho}{w}$ is the residence time of the flow in the reactor tank, and $\gvec = \begin{pmatrix} g_\text{A} \\ g_\text{B} \end{pmatrix}$ is the species production vector obtained from the stoichiometric relation
\begin{equation*} \Smtrx^\top\,\rvec = \gvec , \end{equation*}using $\rvec= r_{\text{f},1} - r_{\text{b},1}$.
The following values are given
| Name | Parameter | Value | Unit |
|---|---|---|---|
| mass flow rate | $w$ | 10 | kg/s |
| inflow concentration of A | $c_\text{Ain}$ | 1.2 | kgmol/$\text{m}^3$ |
| inflow concentration of B | $c_\text{Bin}$ | 0.0 | kgmol/$\text{m}^3$ |
| holdup volume | $V$ | 0.2 | $\text{m}^3$ |
| mass density | $\rho$ | 1000 | kg/$\text{m}^3$ |
| Arrhenius frequency | $k_0$ | $1.97\times 10^{24}$ | $\text{s}^{-1}$ |
| activation temperature | $E/R$ | 20000 | K |
| reverse reaction factor | $\lambda$ | 0.1 | |
| reverse reaction order | $\beta$ | 2.5 | |
| S.S. reactor temperature | $T_\text{ss}$ | 350 | K |
When the reactor reaches steady state (temperature $T_\text{ss} = 350$ K), the forward reaction constant $k_\text{f}$ is given by the Arrhenius relation: $k_\text{f} = k_0\,\exp(\frac{-E}{R\,T})$ at the steady state temperature. Compute the concentration of A and B at steady state.
'''7 Compute the steady state concentration of A and B'''
This show problem does not ask for coding.
Steady state: \begin{equation*} 0 = -\frac{1}{\tau}\bigl(c_\text{A}-c_\text{Ain}\bigr) + g_\text{A} ,\\ 0 = -\frac{1}{\tau}\bigl(c_\text{B}-c_\text{Bin}\bigr) + g_\text{B} , \end{equation*} Compute $\gvec = \begin{pmatrix} g_\text{A} \\ g_\text{B} \end{pmatrix}$ at steady state ($T_\text{ss}=350$ K), $\gvec = \Smtrx^\top\,\rvec$, and substitute into right side of equations and solve for $c_\text{A}$ and $c_\text{B}$.Build your $\Smtrx^\top$ and multiply $\rvec$ to find $\gvec$.
Show Problem 8 (10 pts)¶
If the flow rate through the tank in Problem 7 is too high, the species generation terms are negligible. In this limit, calculate:
- The time-dependent behavior of the concentrations of A and B assuming that the starting concentrations in the tank are the values of the steady state computed in Problem 7.
- What would be the temperature in the tank?
'''8.1a (4pts) Solve for c_a(t)'''
This show problem does not ask for coding.
Fast and furious flow rate: \begin{equation*} \frac{dc_\text{A}}{dt} = -\frac{1}{\tau}\bigl(c_\text{A}-c_\text{Ain}\bigr) + 0 ,\\ \frac{dc_\text{B}}{dt} = -\frac{1}{\tau}\bigl(c_\text{B}-c_\text{Bin}\bigr) + 0 , \end{equation*}'''8.1b (4pts) Solve for c_b(t)'''
This show problem does not ask for coding.
'''8.2 (2pts) Temperature in the tank'''
This show problem does not ask for coding.