ChEn-3170: Computational Methods in Chemical Engineering Fall 2018 UMass Lowell; Prof. V. F. de Almeida 05Nov2018
18. Continuous Stirred-Tank Reactor w/ Heating¶
$ \newcommand{\Amtrx}{\boldsymbol{\mathsf{A}}} \newcommand{\Bmtrx}{\boldsymbol{\mathsf{B}}} \newcommand{\Mmtrx}{\boldsymbol{\mathsf{M}}} \newcommand{\Imtrx}{\boldsymbol{\mathsf{I}}} \newcommand{\Pmtrx}{\boldsymbol{\mathsf{P}}} \newcommand{\Lmtrx}{\boldsymbol{\mathsf{L}}} \newcommand{\Umtrx}{\boldsymbol{\mathsf{U}}} \newcommand{\Smtrx}{\boldsymbol{\mathsf{S}}} \newcommand{\xvec}{\boldsymbol{\mathsf{x}}} \newcommand{\uvar}{\boldsymbol{u}} \newcommand{\fvar}{\boldsymbol{f}} \newcommand{\avec}{\boldsymbol{\mathsf{a}}} \newcommand{\bvec}{\boldsymbol{\mathsf{b}}} \newcommand{\cvec}{\boldsymbol{\mathsf{c}}} \newcommand{\rvec}{\boldsymbol{\mathsf{r}}} \newcommand{\mvec}{\boldsymbol{\mathsf{m}}} \newcommand{\gvec}{\boldsymbol{\mathsf{g}}} \newcommand{\zerovec}{\boldsymbol{\mathsf{0}}} \newcommand{\norm}[1]{\bigl\lVert{#1}\bigr\rVert} \newcommand{\transpose}[1]{{#1}^\top} \DeclareMathOperator{\rank}{rank} $
Table of Contents¶
- Objectives
- Problem Statement
- Input Data
- Total Mass Balance
- Species Mole Balance
- Energy Balance
- Vector ODE System
- Degrees of Freedom
- Steady-State Concentration of A
- Steady-State Concentration of B
- Steady-State Heating Power
- Time Integration Function
- Results
- Final Comments
- Interactive Results
Objectives¶
- Introduce the computational notebook approach to problem solving.
- Combine content, problem setup, computational methods, calculations, and interactive analysis in one live, exchangeable, portable (web-based) document.
- Use chemical reaction engineering methods applied to a chemical reactor problem example: open system, time-dependent, non-isothermal, single, endothermic chemical reaction, and two species.
Problem Statement¶
This problem was provided by Prof. Donatelli (2018) while teaching the Fall ChEn-4130 Process Control class.
An irreversible first-order endothermic reaction
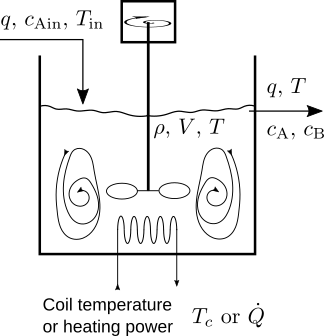
\begin{equation*} \text{A} \overset{k}{\longrightarrow} 2\,\text{B} , \end{equation*}is performed in a continuous-stirred-tank reactor (CSTR; sketch below). A feed stream containing reactant A enters the constant volume CSTR at a mass flow rate $w$, temperature $T_\text{in}$ and molar concentration $c_\text{Ain}$. The product stream leaves the reactor at a mass flow rate $w$, temperature $T$, and molar concentration $c_\text{A}$ for the reactant and $c_\text{B}$ for the product. The heat power addition rate to the CSTR is $\dot{Q}$, and the experimental rate of reaction per unit volume is
\begin{equation*} r_1 = k\,c_\text{A}\,c_\text{B}^0 , \end{equation*}where the rate constant is equal to an Arrhenius equation: $k = k_0\,\exp(\frac{-E}{R\,T})$.
- Derive a dynamic model for the CSTR to determine the concentrations of reactant A and product B and the CSTR temperature as a function of time.
- Perform a degrees of freedom analysis. Identify all input and output variables.
- Determine the steady-state effluent concentrations for reactant A and product B and the required heating rate in kW for the above conditions.
- Suppose a switch is made to a new feed stream with $c_\text{Ain}$ = 1.0 kgmol/m$^3$ and $T_\text{in}$ = 440 K, and the heat addition rate $\dot{Q}$ remains unchanged. Determine the reactant and product concentrations and the CSTR temperature as a function of time. Tabulate and plot the dynamic results. Analyze the results, providing a plausible explanation based on your engineering knowledge.
- Added calculation. Compute, interactively, the effect of simultaneous changes in $c_\text{Ain}$ and $T_\text{in}$ for a range of values in the interval 0.5 to 1.5 kgmol/L, and 400 to 450 K, respectively.
Input Data¶
| Name | Parameter | Value | Unit |
|---|---|---|---|
| mass flow rate | $w$ | 10 | kg/s |
| inflow concentration of A | $c_\text{Ain}$ | 1.2 | kgmol/$\text{m}^3$ |
| inflow concentration of B | $c_\text{Bin}$ | 0.0 | kgmol/$\text{m}^3$ |
| inflow temperature | $T_\text{in}$ | 425 | K |
| holdup volume | $V$ | 0.2 | $\text{m}^3$ |
| mass density | $\rho$ | 1000 | kg/$\text{m}^3$ |
| heat capacity | $C$ | 4184 | J/(kg K) |
| heat of reaction | $\Delta H_\text{R}$ | $6.69 \times 10^8$ | J/kgmol |
| Arrhenius frequency | $k_0$ | $1.97\times 10^{24}$ | $\text{s}^{-1}$ |
| activation temperature | $E/R$ | 20000 | K |
| S.S. reactor temperature | $T_\text{ss}$ | 350 | K |
Note that this is an endothermic reaction (A → 2B) and the reactor temperature at steady state is lower than the temperature of the inflow stream.
'''Parameters'''
w_flow = 10.0 # kgmol/s
c_a_in = 1.2 # kgmol/m^3
c_b_in = 0.0 # kgmol/m^3
temp_in = 425.0 # K
volume = 0.2 # m^3
rho = 1000.0 # kg/m^3
heat_capacity = 4184.0 # J/kg/K
enthalpy_rxn = 6.69e8 # J/kgmol
k_0 = 1.97e24 # 1/s
energy_a_over_r = 20000.0 # K
temp_ss = 350.0 # K
'''Parameters dictionary'''
params = dict()
params['w_flow'] = w_flow
params['c_a_in'] = c_a_in
params['c_b_in'] = c_b_in
params['temp_in'] = temp_in
params['volume'] = volume
params['rho'] = rho
params['heat_capacity'] = heat_capacity
params['enthalpy_rxn'] = enthalpy_rxn
params['k_0'] = k_0
params['energy_a_over_r'] = energy_a_over_r
params['temp_ss'] = temp_ss
Total Mass Balance¶
The volumetric flow rate is a constant because the fluid is incompressible
\begin{equation*} q = q_\text{in} = \text{constant} \end{equation*}where $q = \frac{w}{\rho}$.
Species Mole Balance¶
\begin{equation*} \frac{dc_\text{A}}{dt} = -\frac{1}{\tau}\bigl(c_\text{A}-c_\text{Ain}\bigr) + g_\text{A}(t) ,\\ \frac{dc_\text{B}}{dt} = -\frac{1}{\tau}\bigl(c_\text{B}-c_\text{Bin}\bigr) + g_\text{B}(t) , \end{equation*}where $\tau = \frac{V}{q}$ is the flow residence time in the reactor, $g_\text{A}(t)$ is the species A production rate; similarly for species B, $g_\text{B}(t)$. Note that these equations are the classical relaxation process with a generation term where the relaxation time is the flow residence time $\tau$.
The species production rates are obtained from the stoichiometric relation
\begin{equation*} \Smtrx^\top\,\rvec = \gvec , \end{equation*}for this problem
\begin{equation*} \text{A} \overset{k}{\longrightarrow} 2\,\text{B} , \end{equation*}the stoichiometric relation gives
\begin{equation*} \begin{pmatrix} -1 & 2 \end{pmatrix}^\top \, \begin{pmatrix} k\,c_\text{A} \end{pmatrix} = \begin{pmatrix} -k\,c_\text{A} \\ 2\,k\,c_\text{A} \end{pmatrix} = \begin{pmatrix} g_\text{A} \\ g_\text{B} \end{pmatrix} \end{equation*}where the reaction rate constant is expressed in the Arrhenius form $k = k_0\,e^{\frac{-E}{R\,T}}$.
'''Stoichiometric matrix'''
# (A → 2B)
import numpy as np
stoic_mtrx = np.zeros((1,2)) # stoichiometric matrix
stoic_mtrx[0,0] = -1.0
stoic_mtrx[0,1] = 2.0
params['stoic_mtrx'] = stoic_mtrx
Energy Balance¶
\begin{equation*} \frac{dT}{dt} = -\frac{1}{\tau}\,\bigl(T-T_\text{in}\bigr) - \frac{1}{\rho C} \Bigl(\Delta H_\text{R}\,r_1 - \frac{\dot{Q}}{V}\Bigr) \end{equation*}where $r_1 = k\,c_\text{A}$ is the reaction rate. Note that $\frac{\dot{Q}}{V}$ is the power density of heating or cooling. Likewise $\Delta H_\text{R}\,r_1$ is the power density of chemical reaction heating or cooling. Here again, this is a classical relaxation model with relaxation time equal to the residence time $\tau$ in the presence of a generation term which is a combination of the external heating/cooling power and the chemical reaction heating/cooling power.
Degrees of Freedom¶
There exists three equations (below), the total mass balance forces the volumetric flow rate to be a constant because the flow is incompressible, therefore $q$ must be specified. There exists three unknowns, namely the molarity of species A, $c_\text{A}(t)$, and B, $c_\text{B}(t)$, and the temperature of the reactor, $T(t)$. Therefore there are as many unknowns as there are equations and the problem is potentially solvable given the initial conditions for the unknowns, and all the values of the parameters in the foregoing input data table.
Vector ODE System¶
A vector notation for the foregoing system of equations greatly improves the generality of the derived computer code. First, the usage of the stoichiometric matrix is instrumental for the case with multiple chemical reactions, where the reaction rates can be related to the species production rates that appear as sources in the species mass balances
Towards this goal define
\begin{equation*} \frac{d\uvar}{dt} = \fvar( \uvar, t ) \end{equation*}where $\uvar(t) = (u_1,u_2,u_3)$ and assign
\begin{align*} u_1(t)&=c_\text{A}(t),\\ u_2(t)&=c_\text{B}(t),\\ u_3(t)&=T . \end{align*}Also for $\fvar(\uvar,t) = \bigl(f_1(\uvar,t), f_2(\uvar,t), f_3(\uvar,t)\bigr)$ assign
\begin{align*} f_1 & = - \frac{1}{\tau}\,(u_1-c_\text{Ain}) + g_\text{A}, \\ f_2 & = - \frac{1}{\tau}\,(u_2-c_\text{Bin}) + g_\text{B}, \\ f_3 & = - \frac{1}{\tau}\,(u_3-T_\text{in}) - \frac{1}{\rho C} \Bigl(\Delta H_\text{R}\,r_1 - \frac{\dot{Q}}{V}\Bigr) . \end{align*}Finally, the initial conditions given are as follows:
\begin{align*} u_1(0)&=c_\text{A}(0),\\ u_2(0)&=c_\text{B}(0),\\ u_3(0)&=T_\text{ss}. \end{align*}'''ODE RHS function for integrator'''
def f_vec( t, u_vec, params ):
c_a = u_vec[0]
c_b = u_vec[1]
temp = u_vec[2]
assert c_a >= 0.0
assert c_b >= 0.0
assert temp >= 0.0
c_a_in = params['c_a_in']
c_b_in = params['c_b_in']
temp_in = params['temp_in']
volume = params['volume']
rho = params['rho']
heat_capacity = params['heat_capacity']
enthalpy_rxn = params['enthalpy_rxn']
k_0 = params['k_0']
energy_a_over_r = params['energy_a_over_r']
stoic_mtrx = params['stoic_mtrx']
q_flow = params['q_flow']
q_dot = params['q_dot']
import math
k = k_0 * math.exp(-energy_a_over_r/temp) # reaction rate constant
import numpy as np
r_vec = np.zeros(1)
r_vec[0] = k * c_a # reaction rate vector
g_vec = stoic_mtrx.transpose() @ r_vec # species production rates S^T r = g
tau = volume/q_flow # flow residence time
f_0 = - 1/tau*(c_a - c_a_in) + g_vec[0]
f_1 = - 1/tau*(c_b - c_b_in) + g_vec[1]
f_2 = - 1/tau*(temp - temp_in) \
- 1/rho/heat_capacity*( enthalpy_rxn * r_vec[0] - q_dot/volume )
return np.array([f_0, f_1, f_2])
Steady-State Concentration of A¶
From the species mole balance for A, set the rate of change of $c_\text{A}$ to zero to get
\begin{equation*} c_\text{A}(\infty) = \frac{c_\text{Ain}}{\tau\,k + 1} \end{equation*}'''Steady state concentration of A'''
import math
q_flow = w_flow/rho
tau = volume / q_flow
k_cte = k_0 * math.exp( -energy_a_over_r/temp_ss )
c_a_ss = c_a_in / (tau*k_cte + 1)
params['c_a_ss'] = c_a_ss # save s.s. value in params
params['q_flow'] = q_flow # save s.s. value in params
print('c_A(S.S.) = %5.5e [kgmol/m^3]'%c_a_ss)
print('flow residence time = %5.5e [min]'%(tau/60.0))
c_A(S.S.) = 1.71254e-01 [kgmol/m^3] flow residence time = 3.33333e-01 [min]
Steady-State Concentration of B¶
From the species mole balance for B, set the rate of change of $c_\text{B}$ to zero to get
\begin{equation*} c_\text{B}(\infty) = c_\text{Bin} + \tau\,2\,k\,c_\text{A}(\infty) \end{equation*}'''Steady state concentration of B'''
c_b_ss = c_b_in + tau * 2 * k_cte * c_a_ss
params['c_b_ss'] = c_b_ss # save s.s. value in params
print('c_B(S.S.) = %5.5e [kgmol/m^3]'%c_b_ss)
c_B(S.S.) = 2.05749e+00 [kgmol/m^3]
Steady-State Heating Power¶
From the energy balance
\begin{equation*} \frac{\dot{Q}_\text{ss} }{V} = \frac{\rho C}{\tau} \bigl(T_\text{ss} - T_\text{in}\bigr) + \Delta H_\text{R}\,k\,c_\text{A}(\infty) \end{equation*}'''Steady state heating power'''
q_heat_power_dens_ss = rho*heat_capacity/tau * (temp_ss - temp_in) + enthalpy_rxn * k_cte * c_a_ss
params['q_dot'] = q_heat_power_dens_ss * volume # save s.s. value in params
print('Q(S.S.) = %5.5e [kW]'%(q_heat_power_dens_ss*volume/1000))
Q(S.S.) = 3.74431e+03 [kW]
Heat is provided to the system to keep the reactor temperature from dropping too far below the inflow temperature.
Time Integration Function¶
'''Create the CSTR run function'''
def run_cstr(f_vec, time_stamps, params):
# ODE integrator
from scipy.integrate import odeint
max_n_steps_per_time_step = 100 # max number of nonlinear algebraic solver iterations per time step
u_vec_0 = np.zeros(3,dtype=np.float64)
u_vec_0[0] = params['c_a_ss']
u_vec_0[1] = params['c_b_ss']
u_vec_0[2] = params['temp_ss']
# ODE integrator call
(u_vec_history, info_dict) = odeint( f_vec, u_vec_0, time_stamps,
args=( params, ),
rtol=1e-7, atol=1e-7, mxstep=max_n_steps_per_time_step,
full_output=True, tfirst=True )
assert info_dict['message']=='Integration successful.',\
'Fatal: scipy.integrate.odeint failed %r'%info_dict['message']
return u_vec_history
Results ¶
'''Evolve the CSTR'''
# inflow condition change
params['c_a_in'] = 1.0
params['temp_in'] = 440.0
time_final = 10 * tau # number of residence flow times to evolve
n_time_steps = 200 # number of solution values in time
import numpy as np
time_stamps = np.linspace(0.0, time_final, num=n_time_steps) # create time stamps
u_vec_history = run_cstr(f_vec, time_stamps, params)
print(' Time [s] c_A [kgmol/m^3] c_B [kgmol/m^3] T [K]' )
idx = 0
for (t,u) in zip(time_stamps,u_vec_history):
if idx%5 == 0:
print('%15.3e %15.3e %15.3e %15.3e'%(t,u[0],u[1],u[2]))
idx += 1
Time [s] c_A [kgmol/m^3] c_B [kgmol/m^3] T [K]
0.000e+00 1.713e-01 2.057e+00 3.500e+02
5.025e+00 1.205e-01 2.070e+00 3.523e+02
1.005e+01 8.347e-02 2.075e+00 3.545e+02
1.508e+01 5.791e-02 2.072e+00 3.567e+02
2.010e+01 4.079e-02 2.065e+00 3.589e+02
2.513e+01 2.958e-02 2.055e+00 3.610e+02
3.015e+01 2.232e-02 2.044e+00 3.628e+02
3.518e+01 1.758e-02 2.034e+00 3.643e+02
4.020e+01 1.445e-02 2.025e+00 3.656e+02
4.523e+01 1.232e-02 2.017e+00 3.667e+02
5.025e+01 1.085e-02 2.011e+00 3.675e+02
5.528e+01 9.811e-03 2.006e+00 3.682e+02
6.030e+01 9.061e-03 2.001e+00 3.687e+02
6.533e+01 8.512e-03 1.998e+00 3.692e+02
7.035e+01 8.106e-03 1.996e+00 3.695e+02
7.538e+01 7.802e-03 1.994e+00 3.698e+02
8.040e+01 7.572e-03 1.992e+00 3.700e+02
8.543e+01 7.398e-03 1.991e+00 3.701e+02
9.045e+01 7.265e-03 1.990e+00 3.702e+02
9.548e+01 7.162e-03 1.989e+00 3.703e+02
1.005e+02 7.084e-03 1.988e+00 3.704e+02
1.055e+02 7.023e-03 1.988e+00 3.705e+02
1.106e+02 6.977e-03 1.988e+00 3.705e+02
1.156e+02 6.940e-03 1.987e+00 3.706e+02
1.206e+02 6.912e-03 1.987e+00 3.706e+02
1.256e+02 6.891e-03 1.987e+00 3.706e+02
1.307e+02 6.874e-03 1.987e+00 3.706e+02
1.357e+02 6.861e-03 1.987e+00 3.706e+02
1.407e+02 6.850e-03 1.987e+00 3.707e+02
1.457e+02 6.843e-03 1.987e+00 3.707e+02
1.508e+02 6.836e-03 1.987e+00 3.707e+02
1.558e+02 6.832e-03 1.987e+00 3.707e+02
1.608e+02 6.828e-03 1.986e+00 3.707e+02
1.658e+02 6.825e-03 1.986e+00 3.707e+02
1.709e+02 6.823e-03 1.986e+00 3.707e+02
1.759e+02 6.821e-03 1.986e+00 3.707e+02
1.809e+02 6.820e-03 1.986e+00 3.707e+02
1.859e+02 6.819e-03 1.986e+00 3.707e+02
1.910e+02 6.818e-03 1.986e+00 3.707e+02
1.960e+02 6.817e-03 1.986e+00 3.707e+02
'''Plot Concentration versus Time'''
import matplotlib.pyplot as plt
(fig, ax1) = plt.subplots(1, figsize=(7, 7))
ax1.plot(time_stamps/tau,u_vec_history[:,0],'b-',label='$T_{in}=$ '+str(params['temp_in']) )
ax1.set_xlabel(r'Time [$\tau$] ($\tau=$%5.1f s)'%tau,fontsize=16)
ax1.set_ylabel(r'$c_A$ [mol/L]',fontsize=16,color='blue')
ax1.tick_params(axis='y', labelcolor='blue', labelsize=14)
ax1.tick_params(axis='x', labelsize=14)
ax1.legend(loc='best',fontsize=12)
#ax1.set_ylim(0,1)
ax1.grid(True)
ax2 = ax1.twinx()
ax2.plot(time_stamps/tau,u_vec_history[:,1],'g-.',label='$T_{in}=$ '+str(params['temp_in']) )
ax2.set_ylabel(r'$c_B$ [mol/L]',fontsize=16,color='green')
ax2.tick_params(axis='y', labelcolor='green', labelsize=14)
ax2.legend(loc='center right',fontsize=12)
#ax2.set_ylim(0,2)
#ax2.grid(True)
plt.title('CSTR w/ Heating (endothermic rxn)',fontsize=20)
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.show()
print('')
<Figure size 700x700 with 2 Axes>
import matplotlib.pyplot as plt
(fig, ax1) = plt.subplots(1, figsize=(10, 7))
ax1.plot(time_stamps/tau,u_vec_history[:,2],'r-',label='$T_{in}=$ '+str(params['temp_in']) )
ax1.set_xlabel(r'Time [$\tau$] ($\tau=$%5.1f s)'%tau,fontsize=16)
ax1.set_ylabel(r'$T$ [K]',fontsize=16,color='red')
ax1.tick_params(axis='y', labelcolor='red', labelsize=14)
ax1.tick_params(axis='x', labelsize=14)
ax1.legend(loc='upper left',fontsize=12)
ax1.grid(True)
ax2 = ax1.twinx()
k_cte = params['k_0']*np.exp(-params['energy_a_over_r']/u_vec_history[:,2])
ax2.plot(time_stamps/tau,k_cte,'-',color='navy',label='$T_{in}=$ '+str(params['temp_in']) )
ax2.set_ylabel(r'$k$ [s$^{-1}$]',fontsize=16,color='navy')
ax2.tick_params(axis='y', labelcolor='navy', labelsize=14)
#ax2.legend(loc='best',fontsize=14)
#ax2.grid(True)
ax3 = ax1.twinx()
ax3.plot(time_stamps/tau,q_heat_power_dens_ss/1000/1000*np.ones(time_stamps.size),'-',color='green',label='$T_{in}=$ '+str(params['temp_in']) )
ax3.set_ylabel(r'$\dot{Q}/V$ [kW/L]',fontsize=16,color='green')
ax3.tick_params(axis='y', labelcolor='green', labelsize=14)
#ax3.legend(loc='best',fontsize=14)
#ax2.grid(True)
ax3.spines["right"].set_position(("axes", 1.2))
plt.title('CSTR w/ Heating (endothermic rxn)',fontsize=20)
fig.tight_layout() # otherwise the right y-label is slightly clipped
plt.show()
print('')
Final Comments¶
After the change in feed conditions the reactor reaches another steady state after 125 seconds as shown in the table and plots above.
Interactive Results ¶
'''Create interactive plot'''
import plotly.offline as py # check plotly website
py.init_notebook_mode()
import plotly.graph_objs as go
from ipywidgets import interact # ipython notebook
layout = go.Layout(title="CSTR w/ Heating (endothermic rxn)",
xaxis={'title':'Time [min]'},
yaxis=dict(side='left',title='HELLO'),
yaxis2=dict(overlaying='y',anchor='x',side='right',title='HELLO')
)
fig = go.FigureWidget(layout=layout)
scatt = fig.add_scatter()
fig.layout.titlefont.size = 22
fig.layout.titlefont.family = 'Rockwell'
fig.layout.xaxis.title = 'Time [tau]'
fig.layout.yaxis.title = 'c_A [kgmol/L]'
fig.layout.yaxis2.title = 'T [K]'
'''Vary inflow temperature and molarity of A'''
@interact(inflow_T=(400.0, 450.0, 1.0), inflow_A=(0.7, 1.5, 0.05),
select=['Temperature','Molarity_A','Molarity_B'])
def update(inflow_T=425.0, inflow_A=1.2, select='Molarity_A'):
with fig.batch_update():
scatt.x=time_stamps/tau
params['c_a_in'] = inflow_A
params['temp_in'] = inflow_T
history = run_cstr(f_vec, time_stamps, params)
if select == 'Molarity_A':
scatt.y=history[:,0]
scatt.line.color='blue'
fig.layout.yaxis.title = 'c_A [mol/L]'
elif select == 'Molarity_B':
scatt.y=history[:,1]
scatt.line.color='green'
fig.layout.yaxis.title = 'c_B [mol/L]'
else:
scatt.y=history[:,2]
scatt.line.color='red'
fig.layout.yaxis.title = 'T [K]'
fig
interactive(children=(FloatSlider(value=425.0, description='inflow_T', max=450.0, min=400.0, step=1.0), FloatS…
FigureWidget({
'data': [{'line': {'color': 'blue'},
'type': 'scatter',
'uid': …