Higgs decay to two photons¶
The Standard Model predicted the decay of the [Higgs bosons](https://en.wikipedia.org/wiki/Higgs_boson) into photons. The process is depicted by the diagrams below:
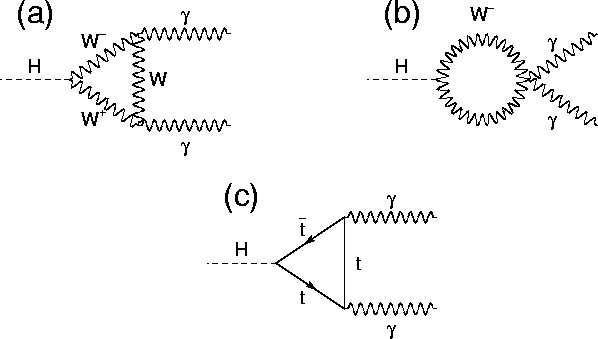 At the [Large Hadron Collider](https://en.wikipedia.org/wiki/Large_Hadron_Collider), this process has been measured. This figure shows how an Higgs boson decay looks in the CMS detector:
At the [Large Hadron Collider](https://en.wikipedia.org/wiki/Large_Hadron_Collider), this process has been measured. This figure shows how an Higgs boson decay looks in the CMS detector:
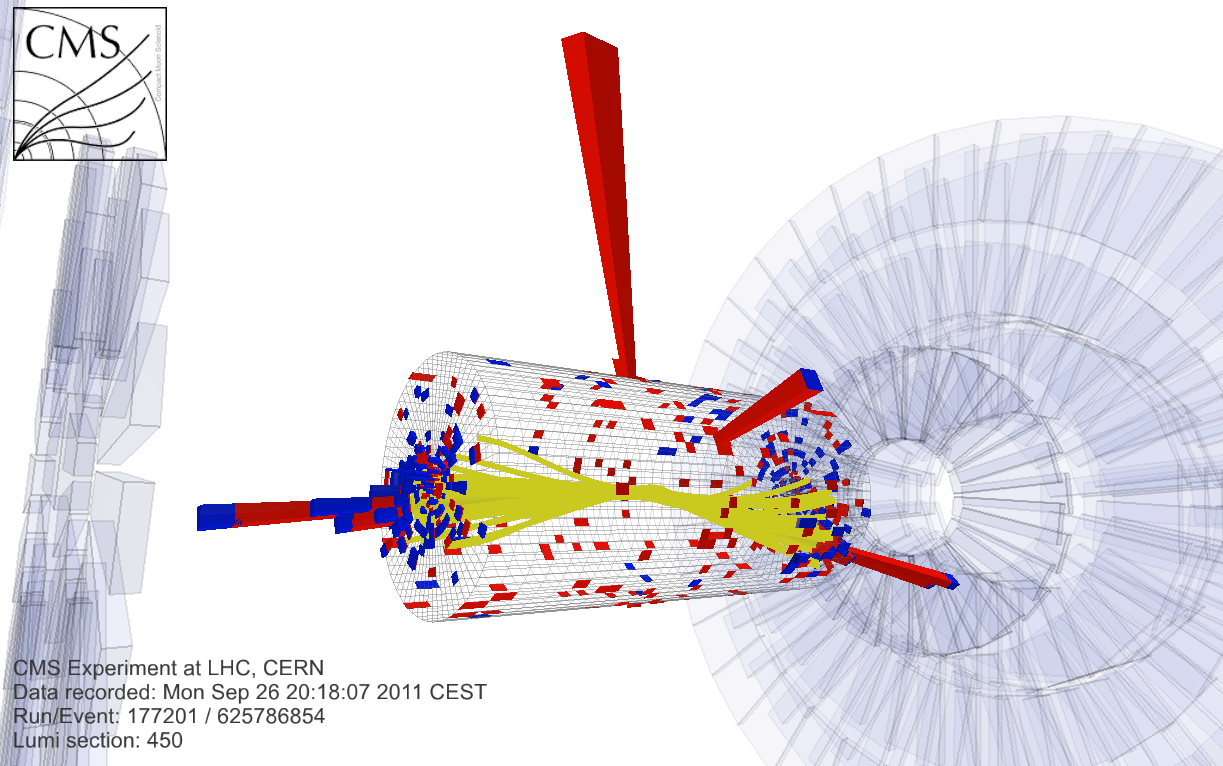 This ROOTbook illustrates a simplified fitting procedure aiming to identify the peak due to the Higgs boson decay over the exponentially falling background.
This ROOTbook illustrates a simplified fitting procedure aiming to identify the peak due to the Higgs boson decay over the exponentially falling background.
Importing input data into a ROOT file¶
First of all we import the input data, here simplistically stored into a text file, into a ROOT file.
TTree tree("HiggsTree","The tree cont");
auto nevt = tree.ReadFile("Hgg.txt","x");
if (nevt <= 0) {
Error("fitHgg","Error reading data from input file ");
}
std::cout << "Read " << nevt << " events from the file." << std::endl;
Read 30770 events from the file.
Modelling the distributions with RooFit¶
ROOT provides a tool for rich modelling of statistical distributions: RooFit. We now create a RooFit model using the RooWorkspace factory: a tool which allow to express complex entities with a convenient syntax.
RooWorkspace w("Workspace for the Higgs fit");
w.factory("x[110,160]"); // invariant mass
// Backround Model
w.factory("nbackground[10000, 0, 10000]");
w.var("nbackground")->setVal(nevt);
w.var("nbackground")->setMin(0.1*nevt);
w.var("nbackground")->setMax(10*nevt);
// Create exponential distribution characterised by two components
w.factory("a1[ 7.5, -500, 500]");
w.factory("a2[-1.5, -500, 500]");
w.factory("expr::z('-(a1*x/100 + a2*(x/100)^2)', a1, a2, x)");
w.factory("Exponential::bmodel(z, 1)");
// Signal Model
w.factory("nsignal[100, 0.0, 1000.0]");
w.factory("mass[130, 110, 150]");
w.factory("width[1, 0.5, 5]");
w.factory("Gaussian::smodel(x, mass, width)");
// Signal + Background Model
w.factory("SUM::model(nbackground*bmodel, nsignal*smodel)");
RooFit v3.60 -- Developed by Wouter Verkerke and David Kirkby
Copyright (C) 2000-2013 NIKHEF, University of California & Stanford University
All rights reserved, please read http://roofit.sourceforge.net/license.txt
Extract the signal and signal+background model from the workspace:
auto smodel = w.pdf("smodel");
auto model = w.pdf("model");
auto x = w.var("x");
We create now the RooFit data set importing the data from the ROOT tree
RooDataSet dataset("data","data",*x,RooFit::Import(tree) )
[#1] INFO:Eval -- RooAbsReal::attachToTree(x) TTree Float_t branch x will be converted to double precision (RooDataSet &) Name: data Title: data
The workspace can be printed on screen: this is very useful to inspect the models created.
w.Print();
RooWorkspace(Workspace for the Higgs fit) Workspace for the Higgs fit contents variables --------- (a1,a2,mass,nbackground,nsignal,width,x) p.d.f.s ------- RooExponential::bmodel[ x=z c=1 ] = 0.000616625 RooAddPdf::model[ nbackground * bmodel + nsignal * smodel ] = 0.000610556 RooGaussian::smodel[ x=x mean=mass sigma=width ] = 3.72665e-06 functions -------- RooFormulaVar::z[ actualVars=(a1,a2,x) formula="-(a1*x/100+a2*(x/100)^2)" ] = -7.39125
Fit of the Data¶
The fitting procedure results in quite some output. It is important to learn how to read the output of a fit, for example the value of the parameters, their uncertainties, the correlation matrix and the status of the fitting procedure.
** Be patient: the fit is unbinned and can take a while **
auto r = model->fitTo(dataset, RooFit::Minimizer("Minuit"),RooFit::Save(true), RooFit::Offset(true));
[#1] INFO:Minization -- p.d.f. provides expected number of events, including extended term in likelihood.
[#1] INFO:NumericIntegration -- RooRealIntegral::init(bmodel_Int[x]) using numeric integrator RooIntegrator1D to calculate Int(x)
[#1] INFO:Minization -- RooMinimizer::optimizeConst: activating const optimization
[#1] INFO:NumericIntegration -- RooRealIntegral::init(bmodel_Int[x]) using numeric integrator RooIntegrator1D to calculate Int(x)
[#1] INFO:Minization -- The following expressions will be evaluated in cache-and-track mode: (bmodel,smodel)
**********
** 1 **SET PRINT 1
**********
**********
** 2 **SET NOGRAD
**********
PARAMETER DEFINITIONS:
NO. NAME VALUE STEP SIZE LIMITS
1 a1 7.50000e+00 1.00000e+02 -5.00000e+02 5.00000e+02
2 a2 -1.50000e+00 1.00000e+02 -5.00000e+02 5.00000e+02
3 mass 1.30000e+02 4.00000e+00 1.10000e+02 1.50000e+02
4 nbackground 1.00000e+04 3.46150e+03 3.07700e+03 3.07700e+05
5 nsignal 1.00000e+02 5.00000e+01 0.00000e+00 1.00000e+03
6 width 1.00000e+00 2.50000e-01 5.00000e-01 5.00000e+00
**********
** 3 **SET ERR 0.5
**********
**********
** 4 **SET PRINT 1
**********
**********
** 5 **SET STR 1
**********
NOW USING STRATEGY 1: TRY TO BALANCE SPEED AGAINST RELIABILITY
**********
** 6 **MIGRAD 3000 1
**********
FIRST CALL TO USER FUNCTION AT NEW START POINT, WITH IFLAG=4.
[#1] INFO:Minization -- RooNLLVar::evaluatePartition(nll_model_data) first = 0 last = 30770 Likelihood offset now set to -156454
START MIGRAD MINIMIZATION. STRATEGY 1. CONVERGENCE WHEN EDM .LT. 1.00e-03
FCN=-382.008 FROM MIGRAD STATUS=INITIATE 121 CALLS 122 TOTAL
EDM= unknown STRATEGY= 1 NO ERROR MATRIX
EXT PARAMETER CURRENT GUESS STEP FIRST
NO. NAME VALUE ERROR SIZE DERIVATIVE
1 a1 7.50000e+00 1.00000e+02 0.00000e+00 8.25702e+04
2 a2 -1.50000e+00 1.00000e+02 0.00000e+00 2.24355e+05
3 mass 1.24328e+02 4.00000e+00 0.00000e+00 4.66358e+01
4 nbackground 1.00000e+04 3.46150e+03 0.00000e+00 -9.08535e+04
5 nsignal 3.12951e+02 5.00000e+01 3.64250e-01 -6.58428e+02
6 width 1.00000e+00 2.50000e-01 0.00000e+00 -1.31987e+02
ERR DEF= 0.5
MIGRAD MINIMIZATION HAS CONVERGED.
MIGRAD WILL VERIFY CONVERGENCE AND ERROR MATRIX.
COVARIANCE MATRIX CALCULATED SUCCESSFULLY
FCN=-13661.8 FROM MIGRAD STATUS=CONVERGED 393 CALLS 394 TOTAL
EDM=2.73773e-06 STRATEGY= 1 ERROR MATRIX ACCURATE
EXT PARAMETER STEP FIRST
NO. NAME VALUE ERROR SIZE DERIVATIVE
1 a1 8.62159e+00 7.99131e-01 6.71813e-06 3.95343e+00
2 a2 -2.02240e+00 3.02257e-01 2.53440e-06 1.05738e+01
3 mass 1.24043e+02 5.15995e-01 2.10145e-03 -5.18608e-02
4 nbackground 3.04968e+04 1.94972e+02 1.62355e-04 1.89450e-01
5 nsignal 2.73230e+02 8.78269e+01 1.19000e-02 4.11368e-03
6 width 1.48353e+00 4.52588e-01 1.61020e-02 1.06792e-03
ERR DEF= 0.5
EXTERNAL ERROR MATRIX. NDIM= 25 NPAR= 6 ERR DEF=0.5
6.386e-01 -2.412e-01 4.880e-03 -2.017e+01 -2.020e+01 5.434e-02
-2.412e-01 9.136e-02 -1.149e-03 7.853e+00 7.866e+00 -2.129e-02
4.880e-03 -1.149e-03 2.663e-01 8.075e+00 8.083e+00 -5.954e-02
-2.017e+01 7.853e+00 8.075e+00 3.801e+04 7.529e+03 -2.255e+01
-2.020e+01 7.866e+00 8.083e+00 7.529e+03 7.816e+03 -2.257e+01
5.434e-02 -2.129e-02 -5.954e-02 -2.255e+01 -2.257e+01 2.090e-01
PARAMETER CORRELATION COEFFICIENTS
NO. GLOBAL 1 2 3 4 5 6
1 0.99864 1.000 -0.999 0.012 -0.129 -0.286 0.149
2 0.99865 -0.999 1.000 -0.007 0.133 0.294 -0.154
3 0.26833 0.012 -0.007 1.000 0.080 0.177 -0.252
4 0.43699 -0.129 0.133 0.080 1.000 0.437 -0.253
5 0.66859 -0.286 0.294 0.177 0.437 1.000 -0.558
6 0.57982 0.149 -0.154 -0.252 -0.253 -0.558 1.000
**********
** 7 **SET ERR 0.5
**********
**********
** 8 **SET PRINT 1
**********
**********
** 9 **HESSE 3000
**********
COVARIANCE MATRIX CALCULATED SUCCESSFULLY
FCN=-13661.8 FROM HESSE STATUS=OK 40 CALLS 434 TOTAL
EDM=2.78595e-06 STRATEGY= 1 ERROR MATRIX ACCURATE
EXT PARAMETER INTERNAL INTERNAL
NO. NAME VALUE ERROR STEP SIZE VALUE
1 a1 8.62159e+00 8.80308e-01 1.34363e-06 1.72440e-02
2 a2 -2.02240e+00 3.33009e-01 5.06880e-07 -4.04480e-03
3 mass 1.24043e+02 5.19935e-01 8.40581e-05 -3.02425e-01
4 nbackground 3.04968e+04 1.96490e+02 3.24710e-05 -9.61368e-01
5 nsignal 2.73230e+02 9.09340e+01 4.76002e-04 3.61233e+00
6 width 1.48353e+00 4.67716e-01 6.44080e-04 -5.97862e-01
ERR DEF= 0.5
EXTERNAL ERROR MATRIX. NDIM= 25 NPAR= 6 ERR DEF=0.5
7.749e-01 -2.928e-01 2.922e-03 -2.578e+01 -2.578e+01 7.361e-02
-2.928e-01 1.109e-01 -3.724e-04 9.988e+00 9.989e+00 -2.867e-02
2.922e-03 -3.724e-04 2.704e-01 9.088e+00 9.089e+00 -6.868e-02
-2.578e+01 9.988e+00 9.088e+00 3.861e+04 8.112e+03 -2.562e+01
-2.578e+01 9.989e+00 9.089e+00 8.112e+03 8.386e+03 -2.563e+01
7.361e-02 -2.867e-02 -6.868e-02 -2.562e+01 -2.563e+01 2.235e-01
PARAMETER CORRELATION COEFFICIENTS
NO. GLOBAL 1 2 3 4 5 6
1 0.99888 1.000 -0.999 0.006 -0.149 -0.320 0.177
2 0.99889 -0.999 1.000 -0.002 0.153 0.328 -0.182
3 0.29339 0.006 -0.002 1.000 0.089 0.191 -0.279
4 0.45101 -0.149 0.153 0.089 1.000 0.451 -0.276
5 0.69618 -0.320 0.328 0.191 0.451 1.000 -0.592
6 0.61576 0.177 -0.182 -0.279 -0.276 -0.592 1.000
[#1] INFO:Minization -- RooMinimizer::optimizeConst: deactivating const optimization
Visualisation of the Result¶
auto plot = x->frame();
dataset.plotOn(plot);
model->plotOn(plot);
model->plotOn(plot, RooFit::Components("bmodel"),RooFit::LineStyle(kDashed));
model->plotOn(plot, RooFit::Components("smodel"),RooFit::LineColor(kRed));
model->paramOn(plot);
[#1] INFO:NumericIntegration -- RooRealIntegral::init(bmodel_Int[x]) using numeric integrator RooIntegrator1D to calculate Int(x) [#1] INFO:Plotting -- RooAbsPdf::plotOn(model) directly selected PDF components: (bmodel) [#1] INFO:Plotting -- RooAbsPdf::plotOn(model) indirectly selected PDF components: (z) [#1] INFO:NumericIntegration -- RooRealIntegral::init(bmodel_Int[x]) using numeric integrator RooIntegrator1D to calculate Int(x) [#1] INFO:Plotting -- RooAbsPdf::plotOn(model) directly selected PDF components: (smodel) [#1] INFO:Plotting -- RooAbsPdf::plotOn(model) indirectly selected PDF components: () [#1] INFO:NumericIntegration -- RooRealIntegral::init(bmodel_Int[x]) using numeric integrator RooIntegrator1D to calculate Int(x)
%jsroot on
TCanvas c;
plot->Draw();
c.Draw();
Persistification of the Model on Disk¶
The ROOT I/O is used to write on disk full RooFit models in order to ease their sharing among scientists, also from different experiments - this establishes an important common language which allows to compare and combine analyses.
w.writeToFile("HiggsModel.root",true);