
xarray: Data Access
Unidata AMS 2021 Student Conference
Using xarray, you can work with a variety of multi-dimensional array-based data formats and access methods. NetCDF is the first and best option as a well-supported, metadata-rich format, but GRIB and others are also supported. In this notebook, we will be demonstrating various ways to access data in xarray.
Focuses¶
- Open a local NetCDF file into xarray and preview its contents
- Open a remote dataset from a THREDDS server using OPENDAP and look through its metadata
- Open a local GRIB file using cfgrib and learn how to work with the multiple datasets that often arise from a single GRIB file
Objectives¶
Imports¶
import cfgrib
from datetime import datetime, timedelta
from siphon.catalog import TDSCatalog
import xarray as xr
1. Access local NetCDF data¶
Xarray's data model strongly resembles that of NetCDF, so it is no surprise that NetCDF format data is well-supported in xarray. First, let's try opening a local NetCDF file. All you need to do is pass your (relative or absolute) file path to xr.open_dataset:
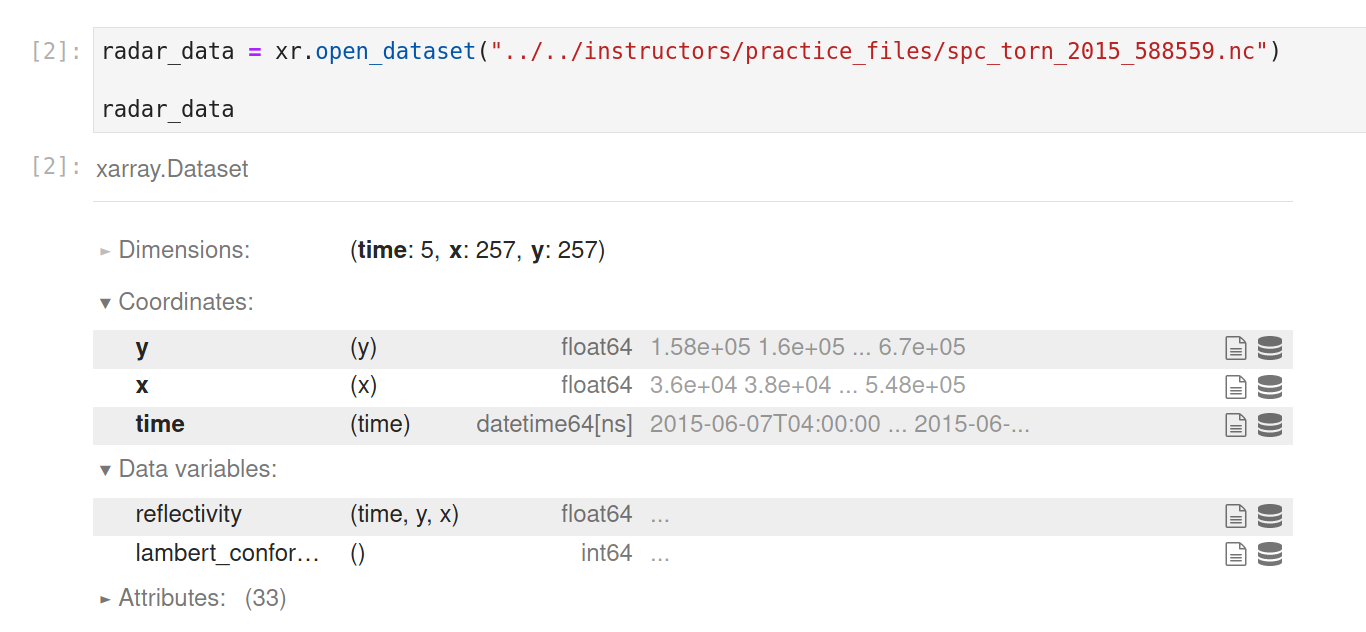
radar_data = xr.open_dataset("../../instructors/practice_files/spc_torn_2015_588559.nc")
radar_data
Notice how xarray identifies all the dimensions, coordinates, data variables and global attributes that are in your NetCDF file. Also, if you're running this yourself, notice the speed of this operation. By default, this operation is just loading in the coordinate and attribute metadata, and not the data variables themselves (which are instead only loaded when they are accessed/used). This is called lazy loading, and is one of the key features that helps make xarray easy to work with.
Let's load in some of those data and preview what it looks like! (For more information on the .isel part of the line of code below, be sure to check out the xarray: indexing training!)
radar_data['reflectivity'].isel(time=2).plot.imshow()
From here, your xarray Dataset is ready to work with (see all the other things you can do with xarray at its documentation or with the other training notebooks linked below).
2. Access remote OPENDAP data¶
What if you don't have your data locally, but instead hosted remotely, on say a THREDDS server? Xarray can still help you here! Instead of specifying a filepath to xr.open_dataset, xarray also natively handles OPENDAP URLs:
gfs_data = xr.open_dataset(
"https://thredds.unidata.ucar.edu/thredds/dodsC/casestudies/python-gallery/"
"GFS_20101026_1200.nc"
)
gfs_data
Notice here how it pulls all of the metadata (dimensions, coordinates, and attributes), but doesn't actually actually download the data variable contents yet. Again, this is the benefit of xarray's lazy-loading approach. Feel free to click around the Dataset HTML object above to see what is contained in this GFS dataset.
One additional word of caution with OPENDAP: sometimes when you copy and past an OPENDAP link from a THREDDS server, you'll get a string that looks like this:
https://thredds.unidata.ucar.edu/thredds/dodsC/casestudies/python-gallery/NARR_19930313_1800.nc.html
Xarray doesn't handle that .html, so you'll have to remove it before you run the Jupyter cell in your code. See how in the example above that has also been done for you.
3. Access local GRIB data¶
On its own, xarray does not natively support GRIB file formats. However, it (fairly seamlessly) integrates with other packages that extend its supported file types to include GRIB. The package we'll be using is called cfgrib.
While you are able to use xr.open_dataset with GRIB files when cfgrib is installed by specifying the engine='cfgrib' keyword argument, because of how cfgrib works, this will often require you to specify custom parameters (e.g., filter_by_keys) to resolve conflicting coordinates. Instead, it is much easier (so long as you have enough capacity to load your entire file into memory) to just use cfgrib.open_datasets, which opens all the data in the GRIB file into separate datasets to resolve any conflicts. You can then search through that list of datasets to find the fields that you need.
Here, we have a GRIB file that produces just one dataset:
mrms_datasets = cfgrib.open_datasets(
"../../instructors/practice_files/"
"MRMS_MergedReflectivityQCComposite_00.50_20150607-060039.grib2",
backend_kwargs={'indexpath':''} # don't use cached idx file, which often gives warning
)
mrms_datasets
See how we have just one dataset in this list, so, to access it, we index into that list:
mrms_data = mrms_datasets[0]
mrms_data['paramId_0'].plot.imshow(vmin=0, vmax=70)
4. Access data with Siphon¶
Siphon can also be used to access remote data and load it into xarray. Two methods are shown below: first, using use_xarray=True in the remote_access method, and second, using the xr.backends.NetCDF4DataStore wrapper to handle other NetCDF4-like data loading methods.
For more details on using Siphon, please glance through all the Siphon remote access trainings available on the course website.
use_xarray=True¶
See the remote access with Siphon training notebook, from which this example comes.
catUrl = "https://thredds-test.unidata.ucar.edu/thredds/catalog/casestudies/harvey/model/gfs/GFS_Global_0p5deg_20170825_1800.grib2/catalog.xml"
datasetName = "GFS_Global_0p5deg_20170825_1800.grib2"
catalog = TDSCatalog(catUrl)
ds = catalog.datasets[datasetName]
dataset = ds.remote_access(use_xarray=True)
dataset
Backend wrapper¶
See the siphon subset notebook, from which this example comes
top_cat = TDSCatalog('http://thredds.ucar.edu/thredds/catalog.xml')
models_cat = top_cat.catalog_refs[0].follow() # follow reaturns a handle to the specified dataset
gfs_cat = models_cat.catalog_refs['GFS Quarter Degree Forecast'].follow()
ds = gfs_cat.latest
ncss = ds.subset()
query = ncss.query()
query.lonlat_point(lon=-105, lat=40) # set coordinates of point of interest.
now = datetime.utcnow() # get current time
query.time_range(now, now + timedelta(days=1)) # create time range of 24 hours
query.variables('Temperature_surface') # request surface temperature variable
query.accept('netcdf4') # return data as a netCDF4 object
grid_data = xr.open_dataset(xr.backends.NetCDF4DataStore(ncss.get_data(query))) # wrap NetCDF4-like object so that xarray accepts it
grid_data
See also¶
There's a lot more to xarray than just opening files! Be sure to take a look at the other xarray training notebooks linked below: