Disclaimer: Most of the content in this notebook is coming from www.scipy-lectures.org
NumPy¶
NumPy is the fundamental package for scientific computing with Python. It is the basic building block of most data analysis in Python and contains highly optimized routines for creating and manipulating arrays.
Everything revolves around numpy arrays¶
Scipyadds a bunch of useful science and engineering routines that operate on numpy arrays. E.g. signal processing, statistical distributions, image analysis, etc.pandasadds powerful methods for manipulating numpy arrays. Like data frames in R - but typically faster.scikit-learnsupports state-of-the-art machine learning over numpy arrays. Inputs and outputs of virtually all functions are numpy arrays.- If you want many more short exercises than the ones in this notebook - you can find 100 of them here
NumPy arrays vs Python arrays¶
NumPy arrays look very similar to Pyhon arrays.
import numpy as np
a = np.array([0, 1, 2, 3])
a
So, why is this useful? NumPy arrays are memory-efficient containers that provide fast numerical operations. We can show this very quickly by running a simple computation on the same two arrays.
L = range(1000)
# Computing the power of the first 1000 numbers with Python arrays
%timeit [i**2 for i in L]
a = np.array(L)
# Computing the power of the first 1000 numbers with Numpy arrays
%timeit a**2
a = np.array([0, 1, 2, 3])
a
a.shape
2-D, 3-D, ...¶
b = np.array([[0, 1, 2], [3, 4, 5]]) # 2 x 3 array
b
b.shape
c = np.array([[[1], [2]], [[3], [4]]])
c
c.shape
a = np.arange(10) # 0 .. n-1 (!)
a
b = np.arange(1, 9, 2) # start, end (exclusive), step
b
... or by number of points¶
c = np.linspace(0, 1, 6) # start, end, num-points
c
d = np.linspace(0, 1, 5, endpoint=False)
d
Common arrays¶
a = np.ones((3, 3)) # reminder: (3, 3) is a tuple
a
b = np.zeros((2, 2))
b
c = np.eye(3)
c
d = np.diag(np.array([1, 2, 3, 4]))
d
np.random: random numbers (Mersenne Twister PRNG)¶
a = np.random.rand(4) # uniform in [0, 1]
a
b = np.random.randn(4) # Gaussian
b
np.random.seed(1234) # Setting the random seed
Basic data types¶
You may have noticed that, in some instances, array elements are displayed with a trailing dot (e.g. 2. vs 2). This is due to a difference in the data-type used:
a = np.array([1, 2, 3])
a.dtype
b = np.array([1., 2., 3.])
b.dtype
Different data-types allow us to store data more compactly in memory, but most of the time we simply work with floating point numbers. Note that, in the example above, NumPy auto-detects the data-type from the input.
You can explicitly specify which data-type you want:
c = np.array([1, 2, 3], dtype=float)
c.dtype
The default data type is floating point:
a = np.ones((3, 3))
a.dtype
There are also other types:
# Complex
d = np.array([1+2j, 3+4j, 5+6*1j])
d.dtype
# Bool
e = np.array([True, False, False, True])
e.dtype
# Strings
f = np.array(['Bonjour', 'Hello', 'Hallo',])
f.dtype # <--- strings containing max. 7 letters
And much more...
int32int64uint32uint64
Indexing and slicing¶
The items of an array can be accessed and assigned to the same way as other Python sequences (e.g. lists):
a = np.arange(10)
a
a[0], a[2], a[-1]
Warning: Indices begin at 0, like other Python sequences (and C/C++). In contrast, in Fortran or Matlab, indices begin at 1.
The usual python idiom for reversing a sequence is supported:
a[::-1]
For multidimensional arrays, indexes are tuples of integers:
a = np.diag(np.arange(3))
a
a[1, 1]
a[2, 1] = 10 # third line, second column
a
a[1]
Note¶
- In 2D, the first dimension corresponds to rows, the second to columns.
- For multidimensional
a,a[0]is interpreted by taking all elements in the unspecified dimensions.
Slicing: Arrays, like other Python sequences can also be sliced¶
a = np.arange(10)
a
a[2:9:3] # [start:end:step]
Note that the last index is not included!
a[:4]
All three slice components are not required: by default, start is 0,
end is the last and step is 1:
a[1:3]
a[::2]
a[3:]
A small illustrated summary of NumPy indexing and slicing...
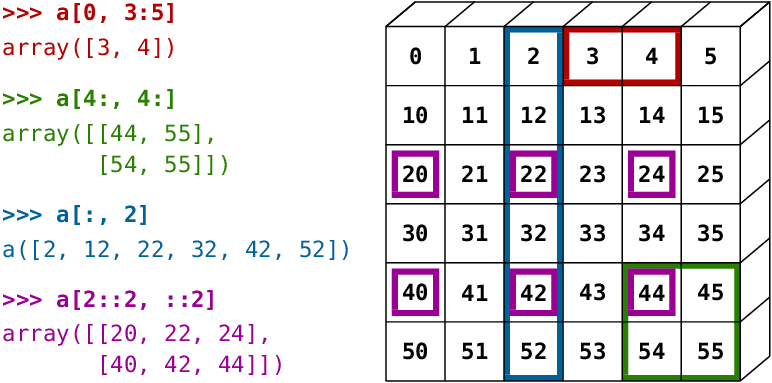
You can also combine assignment and slicing:
a = np.arange(10)
a[5:] = 10
a
b = np.arange(5)
a[5:] = b[::-1]
a
np.random.seed(3)
a = np.random.randint(0, 21, 15)
a
(a % 3 == 0)
mask = (a % 3 == 0)
extract_from_a = a[mask] # or, a[a%3==0]
extract_from_a # extract a sub-array with the mask
Indexing with a mask can be very useful to assign a new value to a sub-array:
a[a % 3 == 0] = -1
a
Indexing with an array of integers¶
a = np.arange(0, 100, 10)
a
Indexing can be done with an array of integers, where the same index is repeated several time:
a[[2, 3, 2, 4, 2]] # note: [2, 3, 2, 4, 2] is a Python list
New values can be assigned with this kind of indexing:
a[[9, 7]] = -100
a
The image below illustrates various fancy indexing applications:
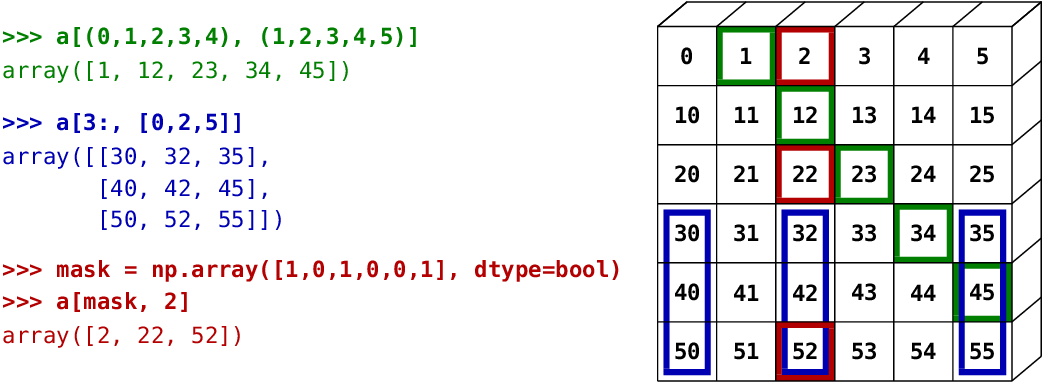

a = np.array([1, 2, 3, 4])
a + 1
2**a
All arithmetic operates elementwise:
b = np.ones(4) + 1
a - b
a * b
j = np.arange(5)
2**(j + 1) - j
Array multiplication is not matrix multiplication¶
c = np.ones((3, 3))
c * c # NOT matrix multiplication!
# Matrix multiplication
c.dot(c)
a = np.array([1, 2, 3, 4])
b = np.array([4, 2, 2, 4])
a == b
a > b
Array-wise comparisons:
a = np.array([1, 2, 3, 4])
b = np.array([4, 2, 2, 4])
c = np.array([1, 2, 3, 4])
np.array_equal(a, b)
np.array_equal(a, c)
Logical operations¶
a = np.array([1, 1, 0, 0], dtype=bool)
b = np.array([1, 0, 1, 0], dtype=bool)
np.logical_or(a, b)
np.logical_and(a, b)
Transcendental functions¶
a = np.arange(5)
np.sin(a)
np.log(a)
np.exp(a)
Shape mismatches¶
a = np.arange(4)
# NBVAL_SKIP
a + np.array([1, 2])
Broadcasting? We'll return to that later.
Transposition¶
a = np.triu(np.ones((3, 3)), 1)
a
a.T
The transposition is a view¶
As a result, the following code is wrong and will not make a matrix symmetric:
>>> a += a.T
It will work for small arrays (because of buffering) but fail for large one, in unpredictable ways.
x = np.array([1, 2, 3, 4])
np.sum(x)
Note: Certain NumPy functions can be also written at the end of an Numpy array.
x.sum()
Sum by rows and by columns:
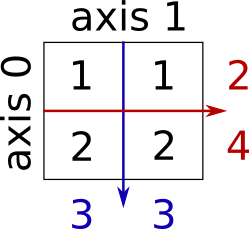
x = np.array([[1, 1], [2, 2]])
x
x.sum(axis=0) # columns (first dimension)
x.sum(axis=1) # rows (second dimension)
Other reductions¶
Like, mean, std, cumsum etc. works the same way (and take axis=).
Extrema¶
x = np.array([1, 3, 2])
x.min()
x.max()
x.argmin() # index of minimum
x.argmax() # index of maximum
Logical operations¶
np.all([True, True, False])
np.any([True, True, False])
Can be used for array comparisons:
a = np.zeros((100, 100))
np.any(a != 0)
np.all(a == a)
a = np.array([1, 2, 3, 2])
b = np.array([2, 2, 3, 2])
c = np.array([6, 4, 4, 5])
((a <= b) & (b <= c)).all()
Statistics¶
x = np.array([1, 2, 3, 1])
y = np.array([[1, 2, 3], [5, 6, 1]])
x.mean()
np.median(x)
np.median(y, axis=-1) # last axis
x.std() # full population standard dev.
... and many more (best to learn as you go).
Broadcasting¶
Basic operations on
numpyarrays (addition, etc.) are elementwiseThis works on arrays of the same size. *Nevertheless, It's also possible to do operations on arrays of different sizes if NumPy* can transform these arrays so that they all have the same size: this conversion is called broadcasting.
The image below gives an example of broadcasting:
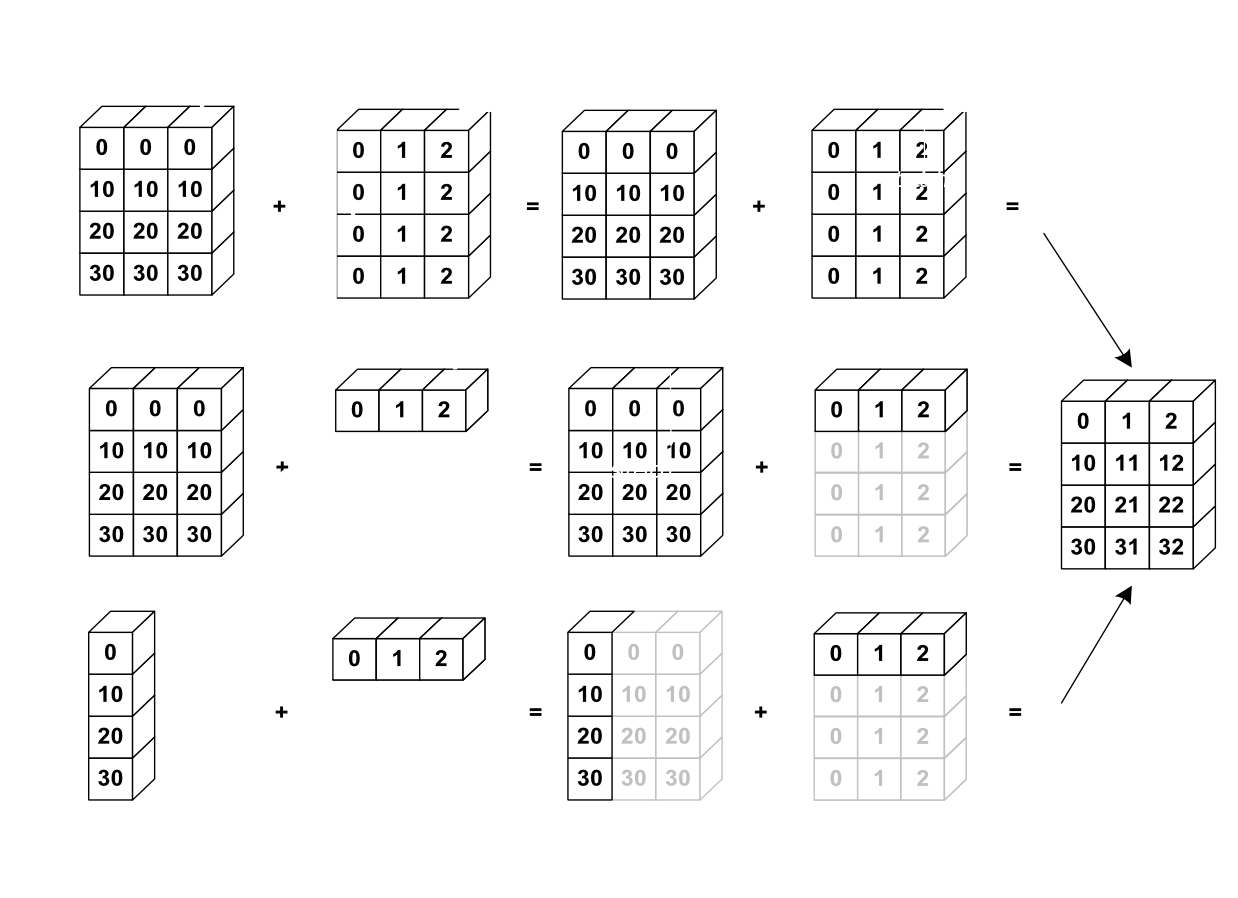
Let's verify this:
a = np.tile(np.arange(0, 40, 10), (3, 1)).T
a
b = np.array([0, 1, 2])
a + b
We have already used broadcasting without knowing it!
a = np.ones((4, 5))
a[0] = 2 # we assign an array of dimension 0 to an array of dimension 1
a
An useful trick:
a = np.arange(0, 40, 10)
a.shape
a = a[:, np.newaxis] # adds a new axis -> 2D array
a.shape
a
a + b
Broadcasting seems a bit magical, but it is actually quite natural to use it when we want to solve a problem whose output data is an array with more dimensions than input data.
A lot of grid-based or network-based problems can also use broadcasting. For instance, if we want to compute the distance from the origin of points on a 10x10 grid, we can do:
x, y = np.arange(5), np.arange(5)[:, None]
distance = np.sqrt(x ** 2 + y ** 2)
distance
Or in color:
a = np.array([[1, 2, 3], [4, 5, 6]])
a
a.ravel()
a.T
a.T.ravel()
Higher dimensions: last dimensions ravel out "first".
Reshaping¶
The inverse operation to flattening:
a = np.array([[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]])
a
a.reshape(6, 2)
Or,
a.reshape((6, -1)) # unspecified (-1) value is inferred
Adding a dimension¶
Indexing with the np.newaxis or None object allows us to add an axis to an array (you have seen this already above in the broadcasting section):
z = np.array([1, 2, 3])
z
z[:, np.newaxis]
z[np.newaxis, :]
Dimension shuffling¶
a = np.arange(4*3*2).reshape(4, 3, 2)
a
a.shape
b = a.transpose(1, 2, 0)
b
b.shape
Resizing¶
Size of an array can be changed with ndarray.resize:
a = np.arange(4)
a.resize((8,))
a
Sorting data¶
Sorting along an axis:
a = np.array([[4, 3, 5], [1, 2, 1]])
b = np.sort(a, axis=1)
b
Important: Note that the code above sorts each row separately!
In-place sort:
a.sort(axis=1)
a
Sorting with fancy indexing:
a = np.array([4, 3, 1, 2])
j = np.argsort(a)
j
a[j]
Finding minima and maxima:
a = np.array([4, 3, 1, 2])
j_max = np.argmax(a)
j_min = np.argmin(a)
j_max, j_min
npy - NumPy's own data format¶
NumPy has its own binary format, not portable but with efficient I/O:
data = np.ones((3, 3))
np.save('pop.npy', data)
data3 = np.load('pop.npy')
Summary - What do you need to know to get started?¶
- Know how to create arrays :
array,arange,ones,zeros. - Know the shape of the array with
array.shape, then use slicing to obtain different views of the array:array[::2], etc. Adjust the shape of the array usingreshapeor flatten it withravel. - Obtain a subset of the elements of an array and/or modify their values with masks
a[a < 0] = 0 - Know miscellaneous operations on arrays, such as finding the mean or max (
array.max(),array.mean()). - For advanced use: master the indexing with arrays of integers, as well as broadcasting.