Agent based or Individual based stochastic models¶
model conceptualization¶
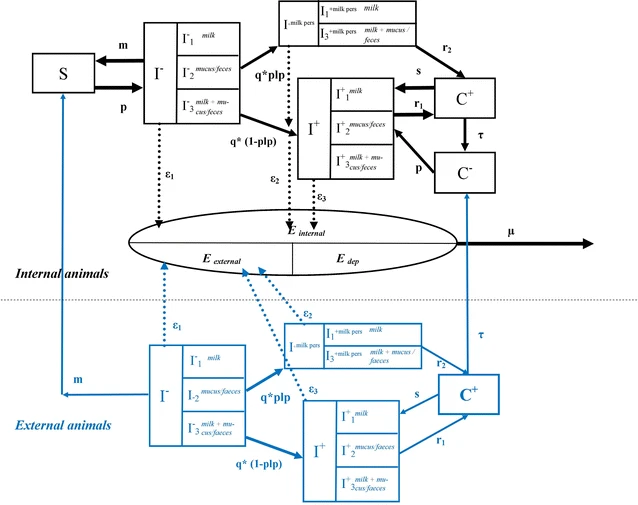
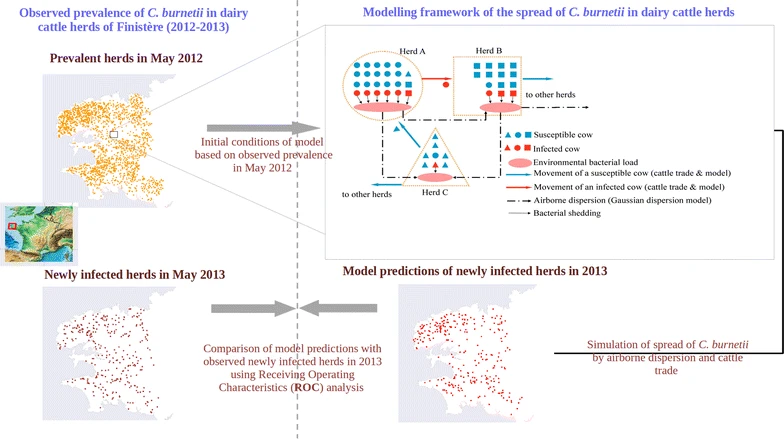
- This is a model for describing Q fever transmission within a dairy herd.
- The idea of the model is simple
- the probability of a susceptible cow to get infected depends on the environmental bacterial load in the herd
- an infected cow sheds bacteria in environment through various routes
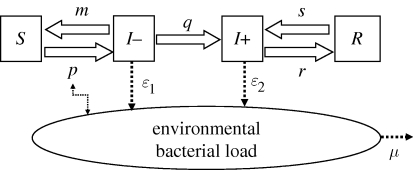
We are developing a simpler version of the model.
We will assume SIR transmission
importing packages¶
In [1]:
import matplotlib.pyplot as plt
import numpy as np
import pylab as pl
import math as math
import random as random
from scipy import stats as stats
import math as ma
import scipy.special as sp
import pandas as pd
import seaborn as sns
import matplotlib.style as style
style.use('fivethirtyeight')
plt.rcParams['font.family'] = 'Times New Roman'
sns.set_context("notebook", font_scale=1.30, rc={"lines.linewidth": 0.8})
%matplotlib inline
C:\Users\falco\anaconda3\lib\site-packages\numpy\_distributor_init.py:30: UserWarning: loaded more than 1 DLL from .libs:
C:\Users\falco\anaconda3\lib\site-packages\numpy\.libs\libopenblas.NOIJJG62EMASZI6NYURL6JBKM4EVBGM7.gfortran-win_amd64.dll
C:\Users\falco\anaconda3\lib\site-packages\numpy\.libs\libopenblas.WCDJNK7YVMPZQ2ME2ZZHJJRJ3JIKNDB7.gfortran-win_amd64.dll
warnings.warn("loaded more than 1 DLL from .libs:"
Lets understand stochasticity¶
- Stochastic processes are randomly determined processes.
- They are random but still have some central tendency.
coin toss¶
In [2]:
probability = 0.5
In [3]:
np.random.multinomial(1, [probability, 1-probability])
Out[3]:
array([1, 0])
In [4]:
def toss_coin():
result = np.random.multinomial(1, [probability, 1-probability])
if result[0] == 1:
print('Heads')
elif result[1] == 1:
print('Tails')
In [5]:
toss_coin()
Tails
multiple outcomes¶
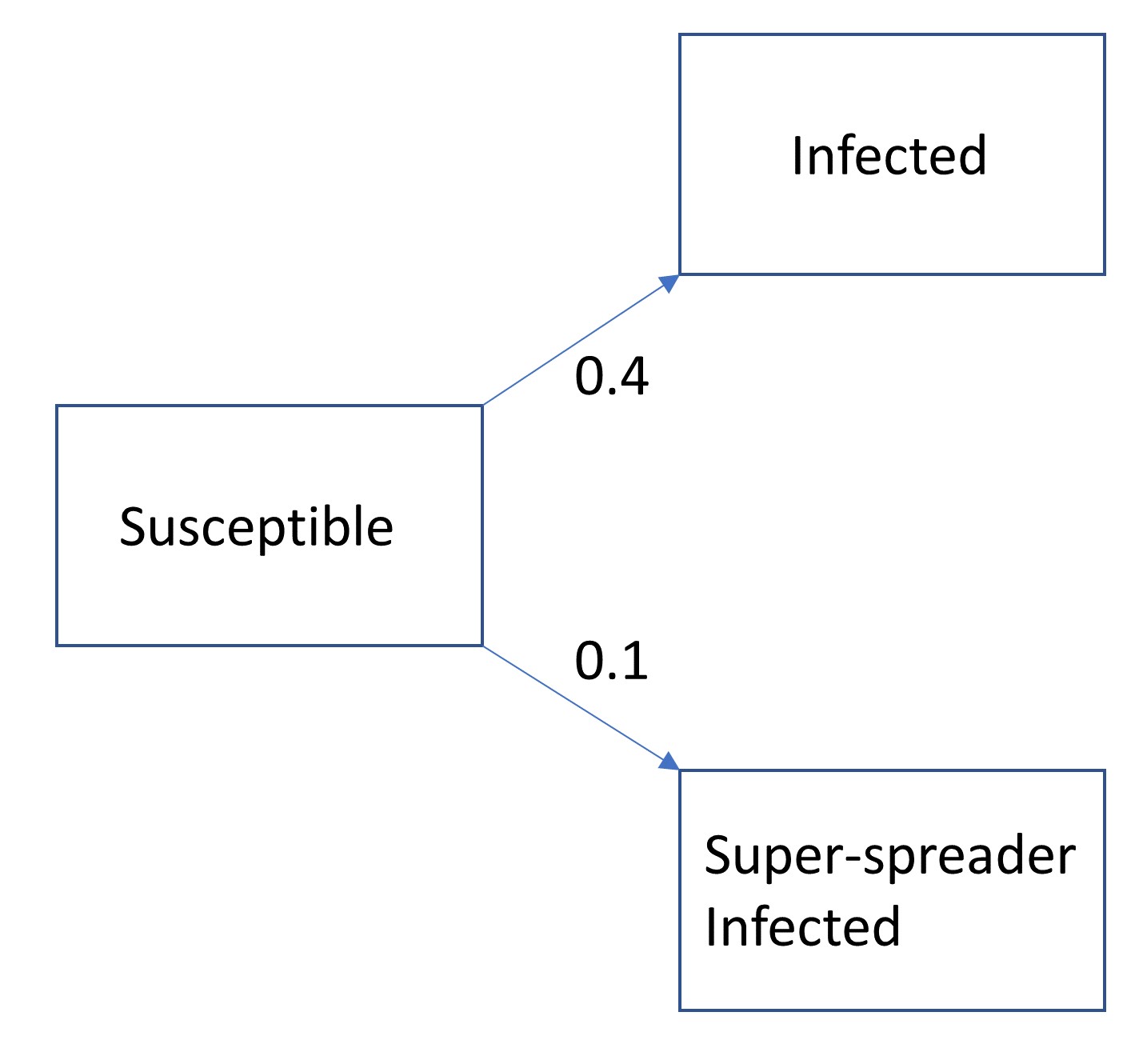
- at time t
- probability of an individual to get infected with normal shedding is 0.4
- and to become a super spreader is 0.1
- hence for it to remain susceptible is 1-(0.1+0.4) = 0.5
In [6]:
p_infection = 0.4 ## probability to get infected
p_superspreader = 0.1 ## probability to become super spreader
def change_susceptibles():
result = np.random.multinomial(1, [(1-(p_infection+ p_superspreader)), p_infection, p_superspreader])
if result[0] == 1:
print('remain susceptible')
elif result[1] == 1:
print('become infected')
elif result[2] == 1:
print('become super-spreader')
In [7]:
np.random.multinomial(1, [(1-(p_infection+ p_superspreader)), p_infection, p_superspreader])
Out[7]:
array([0, 1, 0])
In [8]:
change_susceptibles()
become infected
In [9]:
for i in range(100):
change_susceptibles()
become infected become infected remain susceptible remain susceptible remain susceptible become infected remain susceptible remain susceptible become infected remain susceptible remain susceptible remain susceptible become infected remain susceptible become infected become super-spreader become super-spreader remain susceptible remain susceptible become infected remain susceptible become infected become infected become infected remain susceptible remain susceptible become infected remain susceptible become super-spreader remain susceptible become infected remain susceptible become infected become super-spreader become infected become super-spreader become infected remain susceptible become infected become infected become infected become infected become infected become infected become infected become super-spreader become infected become infected become infected become infected remain susceptible remain susceptible become infected remain susceptible remain susceptible remain susceptible remain susceptible become infected become infected become infected become infected remain susceptible remain susceptible become infected become super-spreader become super-spreader remain susceptible remain susceptible become infected remain susceptible become super-spreader remain susceptible remain susceptible remain susceptible become infected become infected become infected become super-spreader remain susceptible remain susceptible become super-spreader become infected become infected become infected remain susceptible remain susceptible remain susceptible become infected become infected remain susceptible remain susceptible remain susceptible become infected remain susceptible become super-spreader remain susceptible become infected become infected become infected remain susceptible
Object oriented modeling¶
In [15]:
n = 20
class Person:
name = ""
workplace = ""
age = n
def __init__(self, personName, workplace, n):
self.name = personName
self.workplace = workplace
self.age = n
def __repr__(self):
return str(self.name)+', '+str(self.workplace)
In [16]:
Pranav = Person('Pranav', "UC Davis")
--------------------------------------------------------------------------- TypeError Traceback (most recent call last) <ipython-input-16-84c8dd6b3370> in <module> ----> 1 Pranav = Person('Pranav', "UC Davis") TypeError: __init__() missing 1 required positional argument: 'n'
In [17]:
Pranav
--------------------------------------------------------------------------- NameError Traceback (most recent call last) <ipython-input-17-0f309a7818e1> in <module> ----> 1 Pranav NameError: name 'Pranav' is not defined
Model Parameters¶
In [18]:
pi= ma.pi
##Transmission probabilities
m = 0.7; ##Transition probability from I- to S
q = 0.02; ##Transition Probability from I- to I+
pIp = 0.5; ## Proportion of Cows going from I- to I + and becoming I+ milk pers
r1 = 0.2; ##Transition Probability I= to C=
r2 = 0.02; ## Transition Probability I+ to C+
s = 0.15; ##Transition Probability C+ to I+
tau = 0.0096; ##Transition probability C+ to C-
pc = 1; ##Transition Probability from C- to I+
mu = 0.2 ## DEATH OF BACTERIA IN ENVIRONMENT COMPARTMENT
pmf = 0.28; ##Probability of bacteria shed hrough muucs/ feces filing the environement compartment
## Shedding level probabilities for Iminus
qte1 = [0.85,0.15,0];
rho = 0.28; ##fraction of excreted bacteria in mucus / feces contaminating the environment
High = 1/float(30); ##qunatity of bacteria shed by high shedding animal durin one week
#High = 1
intitialheardsize = 50; #Animals in the herd
Agent characteristics¶
In [19]:
def enum(enumName, *listValueNames):
# A sequence of integers, we create as
# As there are values in the enum.
listValueNumbers = range(len(listValueNames))
# Creation definately attributes.
# Fill with initial correspondence: enum value -> integer
dictAttrib = dict( zip(listValueNames, listValueNumbers) )
# Create the reverse dictionary. whole -> enum value
dictReverse = dict( zip(listValueNumbers, listValueNames) )
# Adding the inverse dictionary in attributes
dictAttrib["dictReverse"] = dictReverse
# Create and renvoyage type
mainType = type(enumName, (), dictAttrib)
return mainType
Health = enum("Health", "Susceptible", "Infected", "Recovered")
Herdinfectedornot = enum("Herdinfectedornot","InfectionFree", "Infected")
HerdExposedtoBacteria = enum("HerdExposedtoBacteria","NotExposed","Exposed")
In [ ]:
Agents¶
In [20]:
class Cow(object):
def __init__(self, healthstate = Health.Susceptible):
self.healthstate = healthstate
def __repr__(self):
return str(self.healthstate)
def setHealthState(self, state):
self.healthstate = state
def changeSusceptible(self, Multinomial):
if Multinomial[1] == 1:
self.healthstate = Health.Infected
return True
return False
def changeInfected(self, Multinomial):
if Multinomial[0] == 1:
self.setHealthState(Health.Susceptible)
return True
elif Multinomial[1] == 1:
self.setHealthState(Health.Infected)
elif Multinomial[2] ==1:
self.setHealthState(Health.Recovered)
return True
return False
In [21]:
class calendar (object):
def __init__(self, week=random.randint(1,52)):
self.week = week
def ChangeOneWeek (self):
if self.week < 52:
self.week = self.week+1;
if self.week == 52:
self.week = 0
In [22]:
####################################################################################################################################################
#####################################################################################################################################################
#####################################################################################################################################################
#####################################################################################################################################################
######################## Description of Objects: Herd
class Herd (object):
def __init__(self, initialHerdSize=None, time = None, HerdId = None):
self.initialHerdSize = initialHerdSize
self.time = time
self.HerdID = HerdId
self.A = 17*self.initialHerdSize
self.S = 0;
self.I = 0;
self.R = 0;
self.E = 0;
self.Slist = []
self.Ilist = []
self.clist = []
self.Elist = []
self.Elist.append(self.E)
self.Nlist = []
self.incidence = np.zeros(time_length)
self.N = initialHerdSize
self.pinfection = 1-math.exp(-(self.E))
self.Herdinfectionstate = False
self.PopulateHerd()
def PopulateHerd(self):
self.lCows = [Cow(healthstate=Health.Susceptible) for i in range(self.initialHerdSize)]
def __repr__(self):
return str(self.HerdID)
def InitiateInfection(self,TimeStep):
#self.lCows[0].healthstate = Health.Infected;
#self.lCows[1].healthstate = Health.Infected;
#self.Herdinfectionstate = True
self.HerdExposedAir = HerdExposedtoBacteria.Exposed
self.E = 0.01
# self.indexcases = 4
def InfectionDynamics (self, TimeStep):
if self.pinfection<0:
self.pinfection =0
## loop for all cows
for iCow in self.lCows:
if iCow.healthstate==Health.Susceptible:
state=np.random.multinomial(1, [(1-self.pinfection), self.pinfection])
isInfected = iCow.changeSusceptible(state)
#if isInfected:
# print(iCow.healthstate)
elif iCow.healthstate==Health.Infected:
state=np.random.multinomial(1,[m,(1-m-q),q])
becomingSusceptible = iCow.changeInfected(state)
## Live animals, will be counted each time step
self.S=0;
self.I=0;
self.c=0;
self.N=0;
#____________Enumerating number of animals in each healthstate _______________________________________________________________________________________________________________________________________________________________
for iCow in (self.lCows[:]):
if iCow.healthstate==Health.Susceptible:
self.S+=1
elif iCow.healthstate==Health.Infected:
self.I+=1
elif iCow.healthstate==Health.Recovered:
self.c+=1
self.N =self.S+self.I +self.c
#print(self.I)
def BacteriaExcretion (self):
E1=0;
Q1Dl = 0;
## Calculating the bacteria shedd at time "t"
for iCow in self.lCows:
if iCow.healthstate==Health.Infected:# I shedding with feces
sheddinglevelprob=np.random.multinomial(1,qte1)
if sheddinglevelprob[1]==1:
Q1Dl+=1 ## *low
#____________Counting the Bacteria shed accordint to the SEASON _______________________________________________________________________________________________________________________________________________________________
E1=Q1Dl*High*rho
self.E+=(E1-(self.E*mu))
#print(self.E)
def calculationInfectionProbability (self):
self.pinfection=1-math.exp(-(self.E))
#print(self.pinfection)
def appendingTimeLoopResults(self):
self.Slist.append(self.S)
self.Ilist.append(self.I)
self.clist.append(self.c)
self.Nlist.append(self.N)
self.Elist.append(self.E)
if self.Herdinfectionstate == True and self.I == 0:
self.Herdinfectionstate == False
In [23]:
class ResultsInfectionCause (object):
def __init__ (self,lHerds):
self.Prevalence=0
def appendInfectionCauses (self, lHerds):
for iHerd in lHerds:
#if iHerd.Herdinfectionstate:
if iHerd.ExposedBy != infectionCause.Free :
self.Prevalence+=1
class Results (object):
def __init__(self):
self.Susc = [] ##Suscpetibles
self.INFECTED= [] ##Iminus milk shedders
self.CARRIERS = [] ##Carriers
self.Eload=[] ## Environmental bacterial load
def appendingSimuLoopResults (self, iHerd):
self.Susc.append(iHerd.Slist) ##Suscpetibles
self.INFECTED.append(iHerd.Ilist)
self.CARRIERS.append(iHerd.clist) ##Carriers
self.Eload.append(iHerd.Elist) ## Environmental bacterial load
class SavingResultsInfectionCause (object):
def __init__(self):
self.HerdPrevalence=[]
def appendingresultsofinfectioncause (self, HerdPrevalence):
self.HerdPrevalence.append(HerdPrevalence)
class HerdInfectionProbability (object):
def __init__ (self, simu):
self.InfectedTimes= [0 for x in range(Herds)]
self.ExposedByWIND= [0 for x in range(Herds)]
self.ExposedByMOVEMENT= [0 for x in range(Herds)]
self.MeanTimeforINFECTION=[[0 for x in range (simu)] for h in range(Herds)]
def HerdInfectionProb (self,lHerds,si):
for iHerd in lHerds:
if iHerd.Herdinfectionstate== True:
self.InfectedTimes[lHerds.index (iHerd)] +=1
if iHerd.ExposedBy == infectionCause.Wind :
self.ExposedByWIND[lHerds.index (iHerd)] +=1
if iHerd.ExposedBy == infectionCause.Movement :
self.ExposedByMOVEMENT[lHerds.index (iHerd) ]+=1
if iHerd.TimeforInfection!= None:
self.MeanTimeforINFECTION[lHerds.index (iHerd)][si]= iHerd.TimeforInfection
Simulation¶
In [24]:
lResults = Results()
Running simulation¶
In [25]:
%%time
### this command just print out the computer time required ro run this command
years=5;
time_length=52*years; ## number of time steps I am running the model
dt=1; #time step
simu=100; #number of stochastic iteration
#### loop function that will run the model n times (n being the number of stochastic iterations)
for s in range(simu):
### generate a herd
iHerd = Herd(
initialHerdSize=100,
HerdId="UCDavis",
)
## create cows in our herd
iHerd.PopulateHerd()
##inisitate infection
iHerd.InitiateInfection(TimeStep=1)
## Start the time loop
for t in range (time_length):
iHerd.InfectionDynamics(TimeStep=t) ## first infection dynamics will play, cows will change their health states
iHerd.BacteriaExcretion() ## then infected cows will shed bacteria in the environment
iHerd.calculationInfectionProbability() ## estimate the probability of infection for a new cow based on the contamination
iHerd.appendingTimeLoopResults() ## save results in for time t
## save results of one stochastic iteration in Results object
lResults.appendingSimuLoopResults(iHerd)
Wall time: 37.3 s
Plotting results¶
In [26]:
## Convert all results to dataframe for easy plotting
Susceptibles = pd.DataFrame(lResults.Susc).T
Infected = pd.DataFrame(lResults.INFECTED).T
Recovered = pd.DataFrame(lResults.CARRIERS).T
Eload = pd.DataFrame(lResults.Eload).T
Plot for health states¶
In [27]:
fig, (ax1, ax2, ax3) = plt.subplots(1, 3, sharey = False, figsize = [16,6])
#Susceptibles.plot(color = '#808080', ax = ax1)
#Infected.plot(color = '#808080', ax = ax2)
#Recovered.plot(color = '#808080', ax = ax3)
Susceptibles.plot( ax = ax1)
Infected.plot( ax = ax2)
Recovered.plot( ax = ax3)
ax1.get_legend().remove()
ax2.get_legend().remove()
ax3.get_legend().remove()
ax1.set_xlabel('Time (weeks)')
ax2.set_xlabel('Time (weeks)')
ax3.set_xlabel('Time (weeks)')
ax1.set_ylabel('Number of cows')
ax1.set_title('Susceptibles')
ax2.set_title('Infected')
ax3.set_title('Recovered')
plt.show()
plot for environmental contamination¶
In [45]:
fig, (ax1) = plt.subplots(1, 1, sharey = True, figsize = [16,6])
#Eload.plot(color = '#808080', ax = ax1)
Eload.plot(ax = ax1)
ax1.get_legend().remove()
ax1.set_xlabel('Time (weeks)')
ax1.set_ylabel('Environmental bacterial contamination')
plt.show()
In [46]:
fig, ax = plt.subplots(figsize = (6,4))
Infected.mean(axis=1).plot(ax= ax,label='mean infected')
ax.set_ylim(0)
plt.legend(loc='upper left')
ax.set_ylabel('number of cows')
ax.set_xlabel('time (weeks)')
plt.tight_layout()
plt.show()
In [47]:
fig, ax = plt.subplots(figsize = (6,4))
Eload.mean(axis=1).plot(ax= ax,label='mean contamination')
ax.set_ylim(0)
plt.legend(loc='upper left')
ax.set_ylabel('number of cows')
ax.set_xlabel('time (weeks)')
plt.tight_layout()
plt.show()
Making it mode complicated and complex to represent real life heterogeneities¶