The IPython Notebook: A Comprehensive Tool for Data Science¶
- revised 03 december 2015
- revised 17 december 2014
- revised 16 setembre 2014
- created 17 june 2014
Patrick BROCKMANN - LSCE (Climate and Environment Sciences Laboratory)
<img align="left" width="40%" src="http://www.lsce.ipsl.fr/Css/img/banniere_LSCE_75.png" >

A very good introduction to what is IPython Notebook by Brian Granger¶
from IPython.display import YouTubeVideo
YouTubeVideo('nRHBdkxVn48')
The script from 0:00 to 1:32¶
[...] Today, I want to introduce you to the IPython Notebook and in particuliar describe how the notebook is emerging as an important tool for Data Science.
If you have a bunch of data, the first thing you'll notice it that data is useless on its own. To be usefull, you need to leverage the data to tell a story.
In the process of telling that story, there are many different things that are coming into play :
- you'll need to write code to process and analyze the data
- you'll need to visualize the data
- you'll need to write narrative text
- and possibly perform mathematical derivations
- you'll need to record everything
- share everything with collaborators and finally,
- you'll need to present the story to different audiences.
The IPython Notebook is an open source web-based interactive computing environment for python and other languages that helps you tell stories using data.
As its core, the notebook is designed for writing and running code. For that, you can use the full power of python and its many libraries or even collate other languages such as R (Author Note: or ferret) but in addition to that, the notebook allows you to build shareable documents that combine that
- Code with
- Text
- Equations
- Visualisations
- Images and
- Video
In other words everything that is part of the story you are telling. [...]

Your first example¶
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
t = np.arange(0.0, 5.0, 0.01)
s = np.sin(3*np.pi*t)
plt.plot(t, s, linewidth=4.0, color='b')
plt.xlabel('time (s)')
plt.ylabel('voltage (mV)')
plt.title('About as simple as it gets, folks')
plt.grid(True)
plt.savefig("test.png")
plt.show()

The IPython Notebook: A web-based UI for writing and running code¶
The IPython Notebook is an open source (BSD) tool for telling stories with code and data that are:
- Interactive
- Exploratory
- Collaborative
- Open
- Reproducible
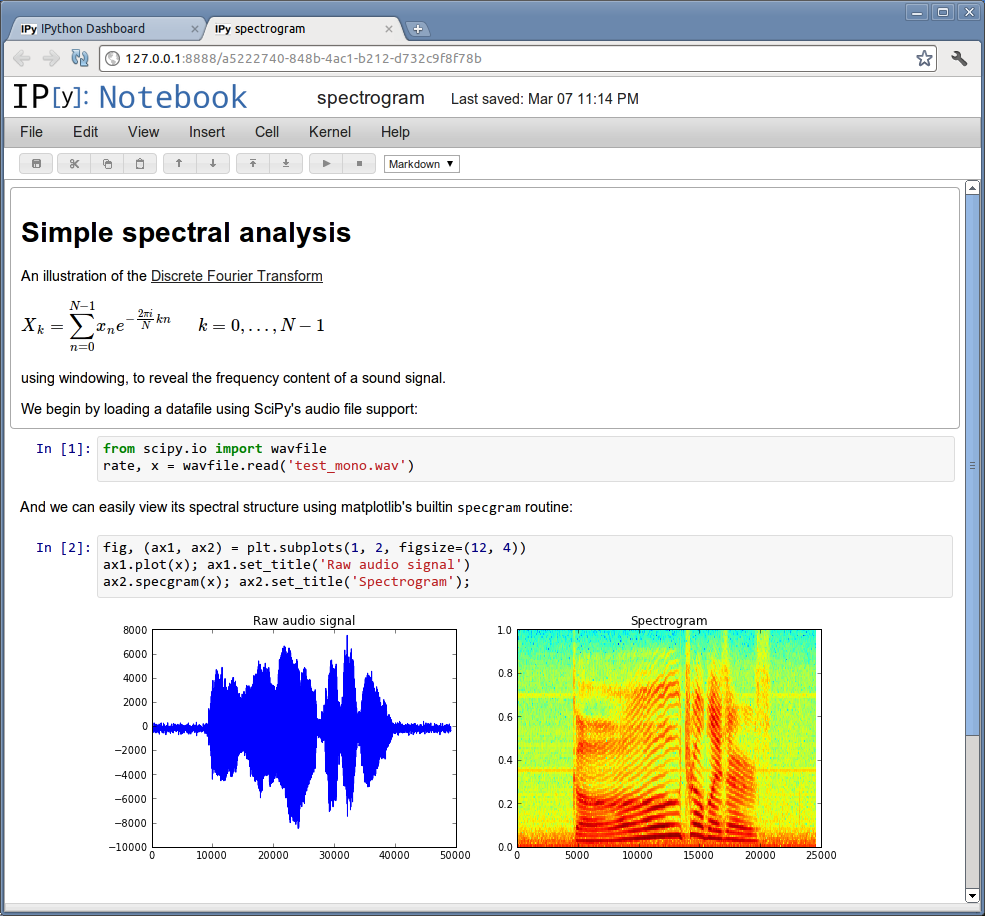
Start the IPython Notebook¶
To start the notebook, type
$ ipython notebook
or with the option --notebook-dir to specify that your directory of notebooks is ~/IPy_Notebooks
$ ipython notebook --notebook-dir=~/IPy_Notebooks
Loading Notebook Files¶
You can also load IPython Notebooks that other people have created, saved as IPython Notebook files (File extension .ipynb.) Try downloading and opening some notebook files from http://nbviewer.ipython.org/github/Unidata/unidata-python-workshop/tree/master/ or http://nbviewer.ipython.org/.

After you download the Notebook file, move it into your IPython Notebook working directory and then choose File -> Open in Notebook to open it.
That Notebook contains some additional code, and some suggestions for changes you can make by going back and editing the existing files. Take a few moments to play with the Notebook - rerun the cells, edit the cells to change them, don't be afraid to break things!
Loading Python Files¶
You can also load a pre-existing Python file into an IPython Notebook cell by typing
%load "myprogram.py"
Into a cell and running it. This loads up a new cell containing the contents of myprogram.py.
Test this feature out by loading one of the scripts you wrote during the recap session. You may have to specify the full path to the script file, depending on the directory IPython Notebook started up from.
There is one other useful built-in tool for working with Python files:
%run "myprogram.py"
This will run myprogram.py and load the output into a Notebook cell.

The IPython project¶

- IPython: open source (BSD) interactive computing environment in Python
- History:
- Started in 2001 by Fernando Perez (BDFL)
- The project now has an extremely dedicated and talented team
- $>$ 20 person years of development, $>$ 150 contributors
- IPython is the de facto standard environment for interactive work in Python
- Funded by:
- Mostly by volunteers
- NASA, DOD/DRC, NIH
- Microsoft, Enthought
- Alfred P. Sloan Foundation ($1.15 million dollar grant starting in Jan. 2013)
- Components:
- IPython Kernel
- Stateful computation engine
- Runs code and returns results
- Uses language agnostic JSON based message protocol over ZeroMQ/WebSockets
- Frontends:
- Terminal Console
- Qt Console
- Notebook
- Parallel computing framework
- IPython Kernel

What are IPython Notebook documents ?¶
- JSON files
- Are stored as files in your local directory
- Can store:
- Code in any language
- Text (Markdown)
- Equations (LaTeX)
- Images
- Links to video
- HTML
- Can be version controlled
- Change 1 line of code, get a 1 line diff
- Can be viewed by anyone online without IPython installed (http://nbviewer.ipython.org/)
- Can be exported to HTML, Markdown, reStructured Text, LaTeX, PDF
- Can be viewed as slideshows with live computations
We try to make writing code pleasant:
- Tab completion
- Integrated help
- Syntax highlighting
- Civilized multiline editing
- Interactive shorthands (aliases, magics)
Not just Python code though! Though cell magics (%%) the Notebook supports running code in other languages:
- R
- ferret (read https://github.com/ipython/ipython/wiki/Extensions-Index#ferret)
- Octave
- Cython
- Bash
- Perl
- Ruby
- Julia
- etc.
%%bash
echo "Hello bash world"
Hello bash world
You can enter latex directly with the %%latex cell magic:
%%latex
\begin{aligned}
\nabla \times \vec{\mathbf{B}} -\, \frac1c\, \frac{\partial\vec{\mathbf{E}}}{\partial t} & = \frac{4\pi}{c}\vec{\mathbf{j}} \\
\nabla \cdot \vec{\mathbf{E}} & = 4 \pi \rho \\
\nabla \times \vec{\mathbf{E}}\, +\, \frac1c\, \frac{\partial\vec{\mathbf{B}}}{\partial t} & = \vec{\mathbf{0}} \\
\nabla \cdot \vec{\mathbf{B}} & = 0
\end{aligned}
You can do the same with R or octave.

Essential Shortcuts¶
Esc/Enter: Mode Switchj/k: Move up/down- Execute Cells
Shift-Enter: Run and go downAlt-Enter: Run and make newControl-Enter: Run in place
a/b: Insert cell above/belowx: cut cell- Cell mode switch:
r: rawm: markdowny: python code


Building slides¶
- Turn on the 'slideshow' cell toolbar
- Types:
- Slide: start a new slide
-: Continue a slide- Sub-Slide: Make a 'down' slide
- Fragment: Make a 'bullet' type incoming slide
- Skip: keep in the notebook, not the deck
- Notes: speaker notes
Then type ipython nbconvert Presentation.ipynb --to slides

Installation and use¶
Standalone installation¶
- My recommendation is to install the completely free Python distribution called Anaconda

Then just type: ipython notebook
From your local machine to asterixN, obelixN (LSCE), curie (TGCC) or ciclad (IPSL) machines¶
 Chained SSH connections through a proxy gateway
Chained SSH connections through a proxy gateway
Read more at http://www.onlamp.com/pub/a/onlamp/excerpt/ssh_11/index1.html
- Make an ssh tunnel from your local machine (through a gateway or not) to the remote machine (replace XXXXX,... by appropriate logins)
- Launch IPython Notebook in the remote terminal
- Open a browser on your local machine
*From your local machine to obelixN (LSCE)*
- ssh -X -t -L70xx:localhost:70xx XXXXX@idefix1.extra.cea.fr ssh -L70xx:localhost:70xx YYYYY@obelix.lscelb.extra.cea.fr
- export PATH="/home/share/unix_files/anaconda/anaconda3/bin:$PATH"
to get the python from the shared anaconda distribution
- module load R/3.1.2 (if you want to use the R magic extension, use in IPython Notebook %load_ext rpy2.ipython)
- jupyter notebook --no-browser --port=70xx --notebook-dir=~/IPy_Notebooks
*From your local machine to curie (TGCC)*
Forwarding ssh port has been forbidden since 10/02/2015
ssh -X -t -L70xx:localhost:70xx XXXXX@idefix1.extra.cea.fr ssh -L70xx:localhost:70xx YYYYY@curie-ccrt.ccc.cea.frmodule load python/2.7.8module load R/3.0.2 (if you want to use the R magic extension, use in IPython Notebook %load_ext rpy2.ipython)module load octave/3.6.3 (if you want to use octave magic extensions)- ~~ipython notebook --no-browser --port=70xx --notebook-dir=~/IPy_Notebooks~~
*From your local machine to ciclad-ng (IPSL)*
- ssh -X -t -L70xx:localhost:70xx XXXXX@ciclad-ng.ipsl.jussieu.fr
- module load python/3.6-anaconda50
- ipython notebook --no-browser --port=70xx --notebook-dir=~/IPy_Notebooks
Beware not sharing the same port number, indicated above as 70xx. Please visit and update http://wiki.ipsl.jussieu.fr/IGCMG/Outils/IPython_Notebook with an available port number.
Now, you should be able to work directly on the remote file system from the browser of your local machine (NOT the remote cluster or marchine) by opening http://localhost:70xx
Remember that ressources from cluster machines are shared among many users. So please shutdown notebooks that are not used anymore (press the red button to do so) and shutdown the notebook server when finish your work (press twice CTRL+C in the console you launch the server). The notebook server has an autosave system and you will certainly find back your notebook in the state you let it.

Publish to share your notebooks with gist or gitHub¶
When ready to publish:
- choose File -> Save in the notebook menu
- in the directory where you run the notebook, find .ipynb file (or do File -> Download as -> IPython (.ipynb)); it is a JSON file
- paste this JSON on http://gist.github.com/ or other pastebin service
- go to http://nbviewer.ipython.org/ and insert Gist's URL (any public URL works)
You can also use gitHub and structure all your notebooks in a repository as https://github.com/PBrockmann/IPy_Notebooks and browsable from http://nbviewer.ipython.org/github/PBrockmann/IPy_Notebooks

A very good talk from Josh Barratt at last OSCON 2014 (Open Source Conference - July 20–24, 2014 Portland)¶
from IPython.display import YouTubeVideo
YouTubeVideo('XkXXpaVpNSc')

An article from Nature that describes the IPython notebook in a study on Workflow software Platforms
http://www.nature.com/naturejobs/2014/140327/pdf/nj7493-523a.pdf

Original workflow
- slow (read whole data file each time, lots of context switching)
- version controlled analysis, but not commentary, difficult to 'go back to'
- Automating requires non-trivial additional dev

Now workflow
- Speedups primarily from no context switching, interactivity, and reusable data loading.
- Reproducible, literate, annotatable, auditable.


Conclusion¶
The IPython Notebook provides an open source environment and foundation for telling stories with code and data.
It puts the fun back into working with code and data
It is finally all about a documented and reproductible workflow that brings you a fluid transition between:
- Exploratory work
- Collaborative development
- Production
- Publication
- Communication


My selection of lectures or interesting notebooks¶
Official website
- Visit the IPython website
A good introduction
Lectures organized by Unidata team
The ferret extension for pyferret
Must have python packages
- numpy
- netCDF4
- matplotlib
- pandas
- cdo.py
- cartopy
Promising packages
- mpld3
- http://mpld3.github.io/
- Jake Vanderplas - Python in the Browser Age: Data Exploration in the IPython Notebook https://www.youtube.com/watch?v=NzX7DDRkecU
Collection of notebooks

Issues of interest¶
- IPython Notebook run as a server
- Cluster usage and parallel computing from notebooks
- Setting a private nbviewer
Please contact me Patrick.Brockmann@lsce.ipsl.fr