Documentation on dimensions and indexing in Parcels¶
Parcels supports Fields on any curvilinear Grid. For velocity Fields (U, V and W), there are some additional restrictions if the Grid is discretised as an Arakawa-B or Arakawa-C grid. That is because under the hood, Parcels uses a specific interpolation scheme for each of these grid types. This is described in Section 2 of Delandmeter and Van Sebille (2019) and summarised below.
The summary is: for Arakawa-B and -C grids, Parcels requires the locations of the corner points of the grid cells for the dimensions dictionary of velocity Fields. In other words, on an Arakawa-C grid, the [k, j, i] node of U will not be located at position [k, j, i] of U.grid.
Barycentric coordinates and indexing in Parcels¶
Arakawa-C grids¶
When calling FieldSet.from_c_grid_dataset() or a method that is based on it like FieldSet.from_nemo(), Parcels automatically sets the interpolation method for the U, V and W Fields to cgrid_velocity. With this interpolation method, the velocity interpolation follows the description in Section 2.1.2 of Delandmeter and Van Sebille (2019).
Importantly, the interpolation of velocities on Arakawa-C grids is done in barycentric coordinates: $\xi$, $\eta$ and $\zeta$ instead of longitude, latitude and depth. These coordinates are always between 0 and 1, measured from the corner of the grid cell. This is more accurate than simple linear interpolation.
from parcels import FieldSet
from glob import glob
import numpy as np
from os import path
from datetime import timedelta as delta
Let's see how this works. We'll use the NemoNorthSeaORCA025-N006_data data, which is on an Arakama-C grid. If we create the FieldSet using the coordinates in the U, V and W files, we get an Error as seen below:
data_path = 'NemoNorthSeaORCA025-N006_data/'
ufiles = sorted(glob(data_path+'ORCA*U.nc'))
vfiles = sorted(glob(data_path+'ORCA*V.nc'))
wfiles = sorted(glob(data_path+'ORCA*W.nc'))
filenames = {'U': ufiles, 'V': vfiles, 'W': wfiles}
variables = {'U': 'uo', 'V': 'vo', 'W': 'wo'}
dimensions = {'U': {'lon': 'nav_lon', 'lat': 'nav_lat', 'depth': 'depthu', 'time': 'time_counter'},
'V': {'lon': 'nav_lon', 'lat': 'nav_lat', 'depth': 'depthv', 'time': 'time_counter'},
'W': {'lon': 'nav_lon', 'lat': 'nav_lat', 'depth': 'depthw', 'time': 'time_counter'}}
fieldset = FieldSet.from_nemo(filenames, variables, dimensions)
--------------------------------------------------------------------------- ValueError Traceback (most recent call last) <ipython-input-2-551c32df110d> in <module> 10 'W': {'lon': 'nav_lon', 'lat': 'nav_lat', 'depth': 'depthw', 'time': 'time_counter'}} 11 ---> 12 fieldset = FieldSet.from_nemo(filenames, variables, dimensions) ~/Codes/PARCELScode/parcels/fieldset.py in from_nemo(cls, filenames, variables, dimensions, indices, mesh, allow_time_extrapolation, time_periodic, tracer_interp_method, field_chunksize, **kwargs) 463 fieldset = cls.from_c_grid_dataset(filenames, variables, dimensions, mesh=mesh, indices=indices, time_periodic=time_periodic, 464 allow_time_extrapolation=allow_time_extrapolation, tracer_interp_method=tracer_interp_method, --> 465 field_chunksize=field_chunksize, **kwargs) 466 if hasattr(fieldset, 'W'): 467 fieldset.W.set_scaling_factor(-1.) ~/Codes/PARCELScode/parcels/fieldset.py in from_c_grid_dataset(cls, filenames, variables, dimensions, indices, mesh, allow_time_extrapolation, time_periodic, tracer_interp_method, field_chunksize, **kwargs) 524 525 if 'U' in dimensions and 'V' in dimensions and dimensions['U'] != dimensions['V']: --> 526 raise ValueError("On a C-grid, the dimensions of velocities should be the corners (f-points) of the cells, so the same for U and V. " 527 "See also https://nbviewer.jupyter.org/github/OceanParcels/parcels/blob/master/parcels/examples/documentation_indexing.ipynb") 528 if 'U' in dimensions and 'W' in dimensions and dimensions['U'] != dimensions['W']: ValueError: On a C-grid, the dimensions of velocities should be the corners (f-points) of the cells, so the same for U and V. See also https://nbviewer.jupyter.org/github/OceanParcels/parcels/blob/master/parcels/examples/documentation_indexing.ipynb
We can still load the data this way, if we use the FieldSet.from_netcdf() method. But because this method assumes the data is on an Arakawa-A grid, this will mean less accurate interpolation
fieldsetA = FieldSet.from_netcdf(filenames, variables, dimensions)
Instead, we need to provide the coordinates of the $f$-points. In NEMO, these are called glamf, gphif and depthw. The first two are in the coordinates.nc file, the last one is in the W file
mesh_mask = data_path + 'coordinates.nc'
filenames = {'U': {'lon': mesh_mask, 'lat': mesh_mask, 'depth': wfiles[0], 'data': ufiles},
'V': {'lon': mesh_mask, 'lat': mesh_mask, 'depth': wfiles[0], 'data': vfiles},
'W': {'lon': mesh_mask, 'lat': mesh_mask, 'depth': wfiles[0], 'data': wfiles}}
dimensions = {'U': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw', 'time': 'time_counter'},
'V': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw', 'time': 'time_counter'},
'W': {'lon': 'glamf', 'lat': 'gphif', 'depth': 'depthw', 'time': 'time_counter'}}
fieldsetC = FieldSet.from_nemo(filenames, variables, dimensions)
WARNING: File NemoNorthSeaORCA025-N006_data/coordinates.nc could not be decoded properly by xarray (version 0.15.1).
It will be opened with no decoding. Filling values might be wrongly parsed.
WARNING: Casting lon data to np.float32
WARNING: Casting lat data to np.float32
WARNING: Trying to initialize a shared grid with different chunking sizes - action prohibited. Replacing requested field_chunksize with grid's master chunksize.
We can plot the different grid points to see that indeed, the longitude and latitude of fieldsetA.U and fieldsetA.V are different.
%matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # noqa
fig, ax = plt.subplots()
nind = 3
ax1 = ax.plot(fieldsetA.U.grid.lon[:nind, :nind], fieldsetA.U.grid.lat[:nind, :nind], '.r', label='U points assuming A-grid')
ax2 = ax.plot(fieldsetA.V.grid.lon[:nind, :nind], fieldsetA.V.grid.lat[:nind, :nind], '.b', label='V points assuming A-grid')
ax3 = ax.plot(fieldsetC.U.grid.lon[:nind, :nind], fieldsetC.U.grid.lat[:nind, :nind], '.k', label='C-grid points')
ax.legend(handles=[ax1[0], ax2[0], ax3[0]], loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
Further information about the NEMO C-grids is available in the NEMO 3D tutorial.
Arakawa-B grids¶
Interpolation for Arakawa-B grids is detailed in Section 2.1.3 of Delandmeter and Van Sebille (2019). Again, the dimensions that need to be provided are those of the barycentric cell edges (i, j, k).
Background info: How does Parcels deal with B and C grids?¶
To keep it simple, let's take the case of a 2D Arakawa-C grid.
Why doesn't Parcels need the exact location of the U and V nodes? This is because in C grids, U and V nodes are always located the same way: for each grid cell, U nodes are located at the middle of the left and right edges, and V nodes are located at the bottom and top edges (see right panel in the Figure below).
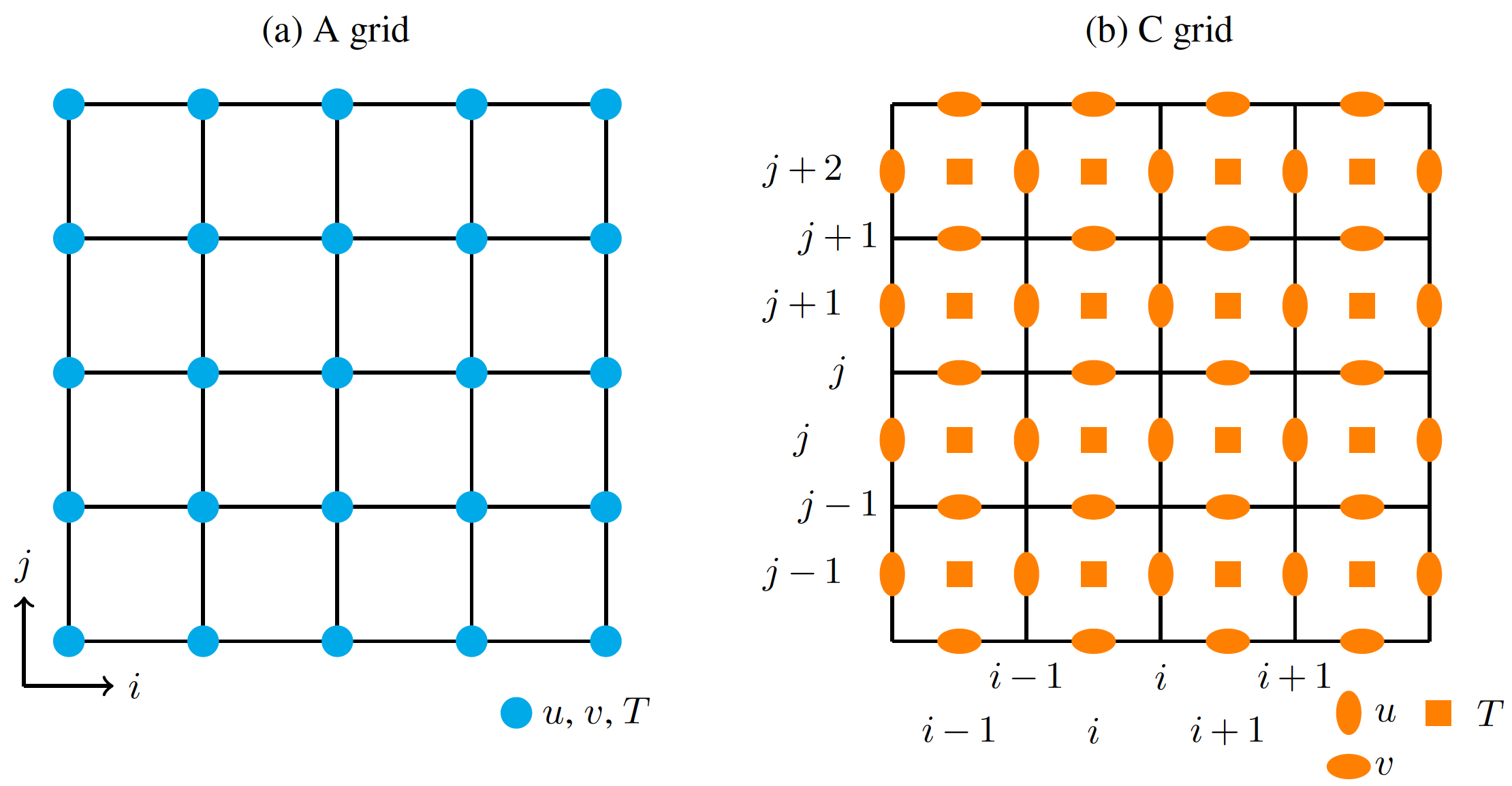
The only information that Parcels needs is the location of the 4 corners of the cell, which are called the f-nodes. Those are the ones provided by U.grid (and are equal to the ones in V.grid).
Now, as you've noticed on the grid illustrated on the figure, there are 4x4 cells. The grid providing the cell corners is a 5x5 grid. But there are only 4x5 U nodes and 5x4 V nodes, since the grids are staggered.
The indexing notation chosen in Parcels is the same as for Nemo, U[j, i] is located between the cell corners [j-1, i] and [j, i], and V[j, i] is located between cell corners [j, i-1] and [j, i]. This implies that the first row of U data and the first column of V data is never used (and do not physically exist), but the U and V fields are correctly provided on a 5x5 table. If your original data are provided on a 4x5 U grid and a 5x4 V grid, you need to regrid your table to follow Parcels notation before creating a FieldSet!
For 3D C grids and B grids, the idea is the same. It is important to follow the indexing notation, which is defined in Parcels and in Delandmeter and Van Sebille (2019).
Note that Parcels can take care of loading these B and C grid data for you, through the general FieldSet.from_c_grid_dataset and FieldSet.from_b_grid_dataset methods, and the model-specific methods FieldSet.from_nemo and FieldSet.from_pop.