Diabetes related genes expressed in pancreas¶
This notebook shows how to integrate genomic and image data resources. This notebook looks at the question Which diabetes related genes are expressed in the pancreas? Tissue and disease can be modified.
Steps:
- Query humanmine.org, an integrated database of Homo sapiens genomic data using the intermine API to find the genes.
- Using the list of found genes, search in the Image Data Resource (IDR) for images linked to the genes, tissue and disease.
We use the intermine API and the IDR API. This notebook is inspired by Workshop_Pax6Workflow.
Summary:¶
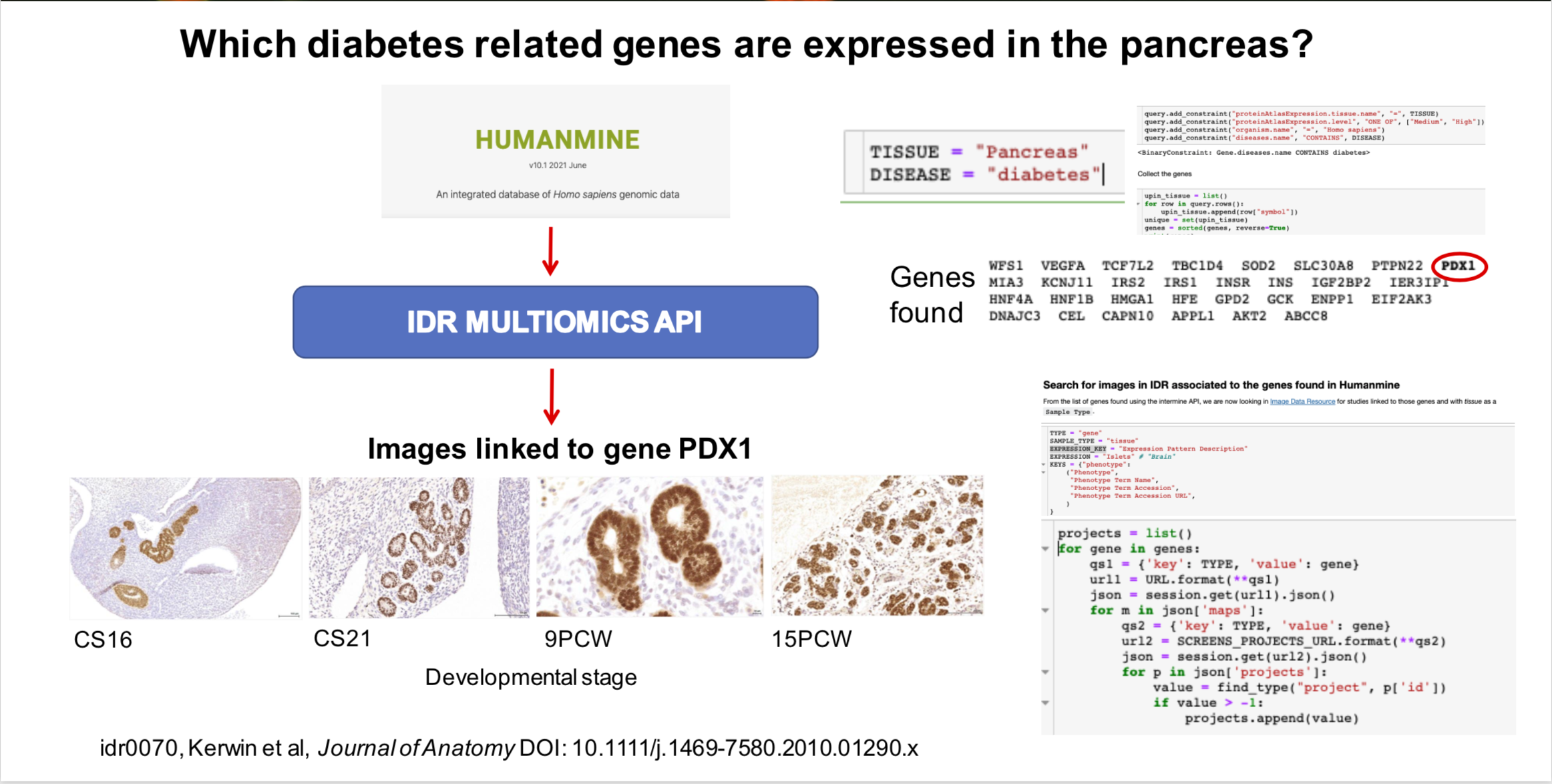
Settings:¶
Auxiliary libraries used¶
- nb_conda_kernels: Enables a Jupyter Notebook or JupyterLab application in one conda environment to access kernels for Python, R, and other languages found in other environments.
- jupyter_contrib_nbextensions: Package containing a collection of community-contributed unofficial extensions that add functionality to the Jupyter notebook.
Launch¶
binder¶
If not already running, you can launch by clicking on the logo
run locally using Docker and repo2docker¶
With jupyter-repo2docker installed, run:
git clone https://github.com/IDR/idr-notebooks.git
cd idr-notebooks
repo2docker .
Install dependencies if required¶
The cell below will install dependencies if you choose to run the notebook in Google Colab.
%pip install intermine
Requirement already satisfied: intermine in /Users/jmarie/opt/anaconda3/envs/stardist-1/lib/python3.9/site-packages (1.13.0) Note: you may need to restart the kernel to use updated packages.
Import libraries¶
# libraries to interact with intermine
from intermine.webservice import Service
# libraries to interact with IDR
import requests
import json
Search for genes in HumanMine¶
We first define the output columns, then add the constraints i.e. specify the tissue and the disease.
service = Service("https://www.humanmine.org/humanmine/service")
query = service.new_query("Gene")
query.add_view(
"primaryIdentifier", "symbol", "proteinAtlasExpression.cellType",
"proteinAtlasExpression.level", "proteinAtlasExpression.reliability",
"proteinAtlasExpression.tissue.name"
)
<intermine.query.Query at 0x111b1b220>
We look for those genes in the specified tissue and that are also associated with the specified disease.
TISSUE = "Pancreas"
DISEASE = "diabetes"
query.add_constraint("proteinAtlasExpression.tissue.name", "=", TISSUE)
query.add_constraint("proteinAtlasExpression.level", "ONE OF", ["Medium", "High"])
query.add_constraint("organism.name", "=", "Homo sapiens")
query.add_constraint("diseases.name", "CONTAINS", DISEASE)
<BinaryConstraint: Gene.diseases.name CONTAINS diabetes>
Collect the genes
upin_tissue = set()
for row in query.rows():
upin_tissue.add(row["symbol"])
genes = sorted(upin_tissue, reverse=True)
Print out the list of genes
for i, a in enumerate(genes):
print(a, end=' ')
if i % 8 == 7:
print("")
YIPF5 WFS1 VEGFA TCF7L2 TBC1D4 SOD2 SLC30A8 PTPN22 PDX1 MIA3 KCNJ11 IRS2 IRS1 INSR INS IGF2BP2 IER3IP1 HNF4A HNF1B HMGA1 HFE GPD2 GCK ENPP1 EIF2AK3 DNAJC3 CEL CAPN10 APPL1 AKT2 ABCC8
Search for images in IDR associated to the genes found in HumanMine¶
From the list of genes found using the intermine API, we are now looking in Image Data Resource for studies linked to those genes and with TISSUE.
Set up where to query and session¶
INDEX_PAGE = "https://idr.openmicroscopy.org/webclient/?experimenter=-1"
# create http session
with requests.Session() as session:
request = requests.Request('GET', INDEX_PAGE)
prepped = session.prepare_request(request)
response = session.send(prepped)
if response.status_code != 200:
response.raise_for_status()
Search studies¶
Search the studies related to the list of genes found in the HumanMine resource.
SEARCH_URL = "https://idr.openmicroscopy.org/searchengine/api/v1/resources/{type}/search/"
KEY_VALUE_SEARCH = SEARCH_URL + "?key={key}&value={value}"
KEY = "Gene Symbol"
%%time
import collections
from collections import defaultdict
results = {}
for gene in genes:
qs1 = {'type': 'image', 'key': KEY, 'value': gene}
url = KEY_VALUE_SEARCH.format(**qs1)
json = session.get(url).json()
images = json['results']['results']
results[gene] = images
CPU times: user 1.02 s, sys: 335 ms, total: 1.36 s Wall time: 10.3 s
First we filter images with the development stage¶
# Annotation key in IDR to find and filter by.
EXPRESSION_KEY = "Expression Pattern Description"
EXPRESSION = "Islets"
STAGE = "Developmental Stage"
development_stage = {}
for k in results:
images = results[k]
result_images = defaultdict(list)
for image in images:
values = image["key_values"]
stage = ""
for v in values:
name = v["name"]
value = v['value']
if name == STAGE:
stage = value
if name == EXPRESSION_KEY and EXPRESSION in value:
result_images[stage].append(image["id"])
development_stage[k] = result_images.items()
print(development_stage)
{'YIPF5': dict_items([]), 'WFS1': dict_items([]), 'VEGFA': dict_items([]), 'TCF7L2': dict_items([]), 'TBC1D4': dict_items([]), 'SOD2': dict_items([]), 'SLC30A8': dict_items([]), 'PTPN22': dict_items([]), 'PDX1': dict_items([('15PCW', [9841210, 9841211]), ('9PCW', [9841212, 9841213, 9841218, 9841219]), ('CS21', [9841214, 9841215]), ('CS16', [9841216, 9841217])]), 'MIA3': dict_items([]), 'KCNJ11': dict_items([]), 'IRS2': dict_items([]), 'IRS1': dict_items([]), 'INSR': dict_items([]), 'INS': dict_items([('15PCW', [9839153, 9839154])]), 'IGF2BP2': dict_items([]), 'IER3IP1': dict_items([]), 'HNF4A': dict_items([]), 'HNF1B': dict_items([]), 'HMGA1': dict_items([]), 'HFE': dict_items([]), 'GPD2': dict_items([]), 'GCK': dict_items([]), 'ENPP1': dict_items([]), 'EIF2AK3': dict_items([]), 'DNAJC3': dict_items([]), 'CEL': dict_items([]), 'CAPN10': dict_items([]), 'APPL1': dict_items([]), 'AKT2': dict_items([]), 'ABCC8': dict_items([])}
Display the images¶
Display the images associated to the genes.
# URLs to retrieve the thumbnails and link to the images in IDR
BASE_URL = "https://idr.openmicroscopy.org/webclient"
IMAGE_DATA_URL = BASE_URL + "/render_thumbnail/{id}"
LINK_URL = BASE_URL + "/?show=image-{id}"
Display the images with development stage¶
Click on the thumbnail to open the image in IDR.
# Display the images
from ipywidgets import AppLayout, widgets
table_widget = widgets.HTML("")
html = "<table>"
for gene in development_stage:
images = development_stage[gene]
if len(images) > 0:
html += '<tr><td><h2>Gene: '+gene+'</h2></td></tr><tr>'
for k, v in images:
html += '<tr><td><h4>Developmental stage: '+k+'</h4></td></tr><tr>'
for i in v:
qs = {'id': i}
url = IMAGE_DATA_URL.format(**qs)
url_link = LINK_URL.format(**qs)
html += '<td><a href="'+url_link+'" target="_blank"><img src="'+url+'"/></a></td>'
html += "</tr>"
html += "</tr>"
html += "</table>"
table_widget.value = html
AppLayout(header=None,
left_sidebar=None,
center=table_widget,
right_sidebar=None,
footer=None)
AppLayout(children=(HTML(value='<table><tr><td><h2>Gene: PDX1</h2></td></tr><tr><tr><td><h4>Developmental stag…
Find the image with the following key(s) and value(s)¶
PART_KEY = "Organism Part"
PATHOLOGY_KEY = "Pathology"
PATHOLOGY_NORMAL_VALUE = "Normal"
pathology_images = {}
for k in results:
images = results[k]
result_images = defaultdict(list)
for image in images:
values = image["key_values"]
part = None
for v in values:
name = v["name"]
value = v['value']
if PART_KEY in name and (TISSUE or EXPRESSION in value):
part = value
for v in values:
name = v["name"]
value = v['value']
if part is not None and name == PATHOLOGY_KEY:
if PATHOLOGY_NORMAL_VALUE in value:
result_images[PATHOLOGY_NORMAL_VALUE].append(image["id"])
else:
result_images[value].append(image["id"])
pathology_images[k] = result_images.items()
Plot the disease vs number of images found
Filter images with a given Organism Part value¶
We explore the images associated to the gene PDX1 with abnormal pathology status.
import pandas as pd
import matplotlib.pyplot as plt
disease_map = {}
gene = "PDX1"
images = pathology_images[gene]
if len(images) == 0:
print("No images found")
else:
for k, v in images:
if k != PATHOLOGY_NORMAL_VALUE:
disease_map[k] = len(v)
disease_ordered = collections.OrderedDict(sorted(disease_map.items()))
df = pd.DataFrame({'disease':disease_ordered.items(),
'number of images':disease_ordered.values()})
df.plot(kind='barh', x='disease', y='number of images', figsize=(10,10))
Select the disease and display the associated images¶
Click on the thumbnail to open the image in IDR.
from ipywidgets import GridspecLayout, widgets
increase = 8
max_value = increase
min_value = 0
disease = ""
def display_images(images, min, max):
html = "<table>"
html += '<tr>'
if min < 0:
min = 0
if max >= len(images):
max = len(images)
for i in images[min:max]:
qs = {'id': i}
url = IMAGE_DATA_URL.format(**qs)
url_link = LINK_URL.format(**qs)
html += '<td><a href="'+url_link+'" target="_blank"><img src="'+url+'"/></a> </td>'
html += "</tr>"
html += "</table>"
html_widget.value = html
# Set the number of images found
count_widget.value = "<b>Number of images found: " + str(len(images)) + "</b>"
def on_selection_change(change):
global disease
if change['name'] == 'value':
values = get_images(change['new'])
if values is None:
return
disease = change['new']
min_value = 0
max_value = increase
display_images(values, min_value, max_value)
def get_images(disease):
for k, v in images:
if k == disease:
return v
return None
def on_click_next(b):
global min_value
global max_value
max_value = max_value + increase
min_value = min_value + increase
values = get_images(disease)
button_previous.disabled = False
if values is None:
return
if max_value > len(values):
button_next.disabled = True
display_images(values, min_value, max_value)
def on_click_previous(b):
global min_value
global max_value
max_value = max_value - increase
min_value = min_value - increase
button_next.disabled = False
if min_value <= 0: # reset
min_value = 0
max_value = increase
button_previous.disabled = True
values = get_images(disease)
if values is not None:
display_images(values, min_value, max_value)
def dropdown_widget(disease_list,
dropdown_widget_name,
displaywidget=False):
selection = widgets.Dropdown(
options=disease_list,
value=disease_list[0],
description=dropdown_widget_name,
disabled=False,
)
selection.observe(on_selection_change)
display_images(get_images(selection.value), min_value, max_value)
return selection
disease_list = list(disease_ordered.keys())
disease = disease_list[0]
gene_widget = widgets.HTML("")
count_widget = widgets.HTML("")
html_widget = widgets.HTML("")
disease_box = dropdown_widget(
disease_list,
'Disease: ', True
)
button_next = widgets.Button(description="Next>>")
button_next.on_click(on_click_next)
button_previous = widgets.Button(description="<<Previous", disabled=True)
button_previous.on_click(on_click_previous)
gene_widget.value = "Gene: <b>" + gene + "</b>"
grid = GridspecLayout(3, 3)
grid[0, 0] = gene_widget
grid[0, 1] = disease_box
grid[0, 2] = count_widget
grid[2, 0] = button_previous
grid[1, :] = html_widget
grid[2, 2] = button_next
grid
GridspecLayout(children=(HTML(value='Gene: <b>PDX1</b>', layout=Layout(grid_area='widget001')), Dropdown(descr…
License (BSD 2-Clause)¶¶
Copyright (C) 2021-2022 University of Dundee. All Rights Reserved.
Redistribution and use in source and binary forms, with or without modification, are permitted provided that the following conditions are met:
Redistributions of source code must retain the above copyright notice, this list of conditions and the following disclaimer. Redistributions in binary form must reproduce the above copyright notice, this list of conditions and the following disclaimer in the documentation and/or other materials provided with the distribution. THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS" AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT OWNER OR CONTRIBUTORS BE LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.