Symbols and Symbol-like representation in neurons¶
- We've seen how to represent vectors in neurons
- And how to compute functions on those vectors
- And dynamical systems
- But how can we do anything like human language?
- How could we represent the fact that "the number after 8 is 9"
- Or "dogs chase cats"
- Or "Anne knows that Bill thinks that Charlie likes Dave"
- Does the NEF help us at all with this?
- Or is this just too hard a problem yet?
Traditional Cognitive Science¶
Lots of theories that work with structured information like this
Pretty much all of them use some representation framework like this:
after(eight, nine)chase(dogs, cats)knows(Anne, thinks(Bill, likes(Charlie, Dave)))
Cognitive models manipulate these sorts of representations
- mental arithmetic
- driving a car
- using a GUI
- parsing language
- etc etc
Seems to match well to behavioural data, so something like this should be right
So how can we do this in neurons?
- This is a hard problem
- Jackendoff (2002) posed this as a major problem
- Four linguistic challenges for cognitive neuroscience
- The Binding Problem
- The Problem of 2
- The Problem of Variables
- Working Memory vs Long-term memory
- The Binding Problem
- Suppose you see a red square and a blue circle
- How do you keep these ideas separate? How does "red" get bound with "square", and how is it kept separate from "blue" and "circle"?
- The Problem of 2
- "The little star's beside the big star"
- How do we keep those two uses of "star" separate?
- The Problem of Variables
- Words seem to have types (nouns, verbs, adjectives, etc, etc)
- How do you make use of that?
- E.g. "blue NOUN" is fine, but "blue VERB" is not (or is very different)
- Working memory vs Long-term memory
- We can both use sentences (working memory) and store them indefinitely (long-term memory)
- How do these transfer back and forth?
- What are they in the brain? This seems to require moving from storing something in neural activity to storing it in connection weights
Possible solutions¶
- Oscillations
- "red square and blue circle"
- Different patterns of activity for RED, SQUARE, BLUE, and CIRCLE
- Have the patterns for RED and SQUARE happen, then BLUE and CIRCLE, then back to RED and SQUARE
- More complex structures possible too:
- E.g. the LISA architecture
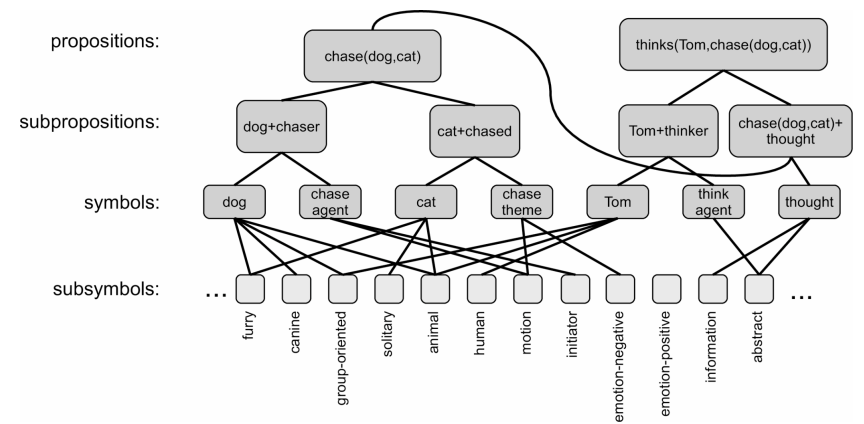
- Problems
- What controls this oscillation?
- How is it parsed? (i.e., mapped to sets of oscillators)
- How do we deal with the exponentional explosion of nodes needed?
'2. Implementing Symbol Systems in Neurons
- Build a general-purpose symbol-binding system
- Lots of temporary pools of neurons
- Ways to temporarily associate them with particular concepts
- Ways to temporarily associate pools together
- Neural Blackboard Architecture

- Problems
- Very particular structure (doesn't seem to match biology)
- Uses a very large number of neurons (~500 million) to be flexible enough for simple sentences
- And that's just to represent the sentence, never mind controlling and manipulating it
'3. Vector operators
- Paul Smolensky 1990 suggests using a mathematical operator called a 'tensor product' to bind vectors together.
- The idea is that this operator can 'bind' and its inverse 'unbind' these vectors together.
- It provides an algebraic way of specifying symbolic structures in a vector space.
- If you can represent vector spaces and operators in neurons (e.g. NEF), then it provides a way of working with symbolic structures in neurons.

- Problems
- Every time you bind vectors, the result has $D^2$ as many dimensions.
- It scales extremely poorly: the dimensionality of a structure representation is the base space to the power of depth, i.e. $D_S = D^{depth+1}$
A deeper issue¶
- If we're just implementing something like the original symbol system in neurons, then we're reducing the neural aspects to mere implementational details
- That is, if we had a perfectly good way to implement symbol structures in neurons, then we could take existing cognitive theories based on symbol structures and implement them in neurons
- But so what? That'd just be conceeding the point that the neurons don't matter -- you don't need them to develop the theory.
- They're useful for testing some aspects, like what firing patterns should be
- But it's more for completeness sake than for understanding
- No more interesting than the question of "well, how does that neuron get implemented in terms of chemistry" or atoms. Or quarks.
- This is why the majority of cognitive scientists don't worry about neurons
Semantic pointers¶
- Tensor products are on the right track, if we can solve the scaling problem.
- 'How to build a brain' exploits a compression operator introduced by Tony Plate called 'circular convolution' $\circledast$
- In the book, I suggest that neurally realized compressed vector representations are prevalent in the brain, and call such representations 'semantic pointers'
- Pointers: because they can be used like computer science pointers to be efficient references to complex representations
- Semantic: because (unlike CS pointers), their content is derived (through compression) from semantically related representations
- This provides something that's similar to the symbolic approach, but much more tied to biology
- Most of the same capabilities as the classic symbol systems
- But not all
- Based on vectors and functions on those vectors
- There is a vector for each concept
- Build up structures by doing math on those vectors
- Example
- blue square and red circle
- can't just do BLUE+SQUARE+RED+CIRCLE
- why?
- need some other operation as well
- Requirements
- input 2 vectors, get a new vector as output
- reversible (given the output and one of the input vectors, generate the other input vector)
- output vector is highly dissimilar to either input vector
- unlike addition, where the output is highly similar
- Circular convoluation can act as such a function
- (There are many other such functions: XOR, Multiply, etc... collectively this kind of vector space binding to represent structures are sometimes called 'Vector Symbolic Architectures' (Gayler, 2003))
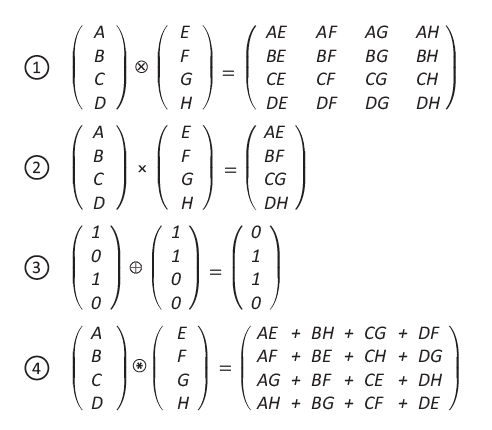

Why circular convolution?
- Extensively studied (Plate, 1997: Holographic Reduced Representations)
- Easy to approximately invert (circular correlation)
Examples:
BLUE$\circledast$SQUARE + RED$\circledast$CIRCLEDOG$\circledast$AGENT + CAT$\circledast$THEME + VERB$\circledast$CHASE

- unbinding:
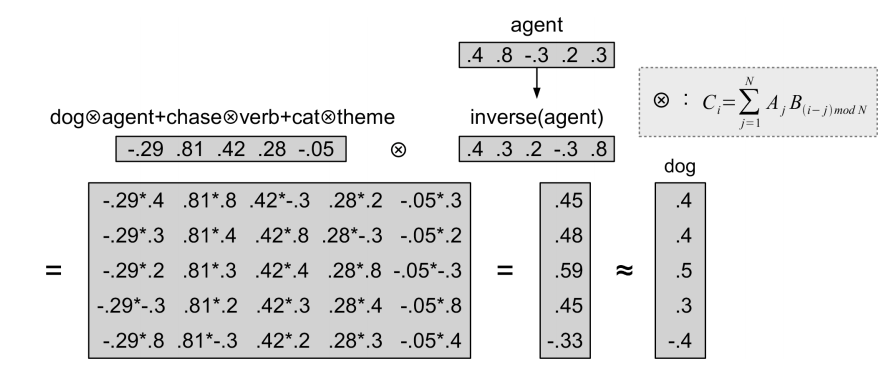
- Can also think of this operator as a compressed outer product

Lots of nice properties
- Can store complex structures
after(eight, nine)NUMBER$\circledast$EIGHT + NEXT$\circledast$NINEknows(Anne, thinks(Bill, likes(Charlie, Dave)))SUBJ$\circledast$ANNE + ACT$\circledast$KNOWS + OBJ$\circledast$(SUBJ$\circledast$BILL + ACT$\circledast$THINKS + OBJ$\circledast$(SUBJ$\circledast$CHARLIE + ACT$\circledast$LIKES + OBJ$\circledast$DAVE))
- But gracefully degrades!
- as representation gets more complex, the accuracy of breaking it apart decreases
- Keeps similarity information
- if
REDis similar toPINKthenRED$\circledast$CIRCLEis similar toPINK$\circledast$CIRCLE
- if
- Can store complex structures
But rather complicated
- Seems like a weird operation for neurons to do
Circular convolution in the NEF¶
- Or is it?
- Circular convolution is a whole bunch ($D^2$) of multiplies
- Like any convolution, it can also be written as a Fourier transform, an elementwise multiply, and another Fourier transform
- i.e. $A \circledast B = IDFT(DFT(A).DFT(B)))$
- The discrete fourier transform is just a linear operation, hence a matrix
- So that's just $D$ pairwise multiplies
- In fact, circular convolution turns out to be exactly what the NEF shows neurons are good at
Jackendoff's Challenges¶
As pointed out in Vector Symbolic Architectures Answer Jackendoff's Challenges for Cognitive Neuroscience
The Binding Problem
- There is a lot of "binding" (structurally combining items) in linguistics (more than vision)
RED$\circledast$CIRCLE + BLUE$\circledast$TRIANGLE- After it is bound, we can ask "what color is the circle by doing $\circledast$
CIRCLE'- where
'is "inverse" - (
RED$\circledast$CIRCLE + BLUE$\circledast$TRIANGLE) $\circledast$CIRCLE' - =
RED$\circledast$CIRCLE$\circledast$CIRCLE' + BLUE$\circledast$TRIANGLE$\circledast$CIRCLE' - =
RED + BLUE$\circledast$TRIANGLE$\circledast$CIRCLE' - =
RED + noise - $\approx$
RED
- where
The Problem of 2
- How can we distinguish two uses of the same concept in a sentence?
- "The little star's beside the big star"
OBJ1$\circledast$(TYPE$\circledast$STAR + SIZE$\circledast$LITTLE) + OBJ2$\circledast$(TYPE$\circledast$STAR + SIZE$\circledast$BIG) + BESIDE$\circledast$OBJ1$\circledast$OBJ2- notice that
BESIDE$\circledast$OBJ1$\circledast$OBJ2=BESIDE$\circledast$OBJ2$\circledast$OBJ1 - and that we can distinguish the two uses of star
- notice that
The Problem of Variables
- How can we manipulate abstract place holders?
S=RED$\circledast$NOUNVAR=BALL$\circledast$NOUN'S$\circledast$VAR=RED$\circledast$BALL
Binding in Working Memory vs Long-term memory
- How can we use and manipulate the same structures in both WM and LTM?
- vectors are what we work with (activity of neurons)
- functions are long-term memory (connection weights)
- functions return (and modify) vectors
Symbol-like manipulation¶
- Can do a lot of standard symbol stuff
- Have to explicitly bind and unbind to manipulate the data
- Less accuracy for more complex structures
- But we can also do more with these representations
Raven's Progressive Matrices¶
- An IQ test that's generally considered to be the best at measuring general-purpose "fluid" intelligence
- nonverbal (so it's not measuring language skills, and fairly unbiased across cultures, hopefully)
- fill in the blank
- given eight possible answers; pick one

- This is not an actual question on the test
- The test is copyrighted
- They don't want the test to leak out, since it's been the same set of 60 questions since 1936
- But they do look like that
How can we model people doing this task?
A fair number of different attempts
- None neural
- Generally use the approach of building in a large set of different types of patterns to look for, and then trying them all in turn
- Which seems wrong for a test that's supposed to be about flexible, fluid intelligence
Does this vector approach offer an alternative?
- First we need to represent the different patterns as a vector
- This is a hard image interpretation problem
- Still ongoing work here
- So we'll skip it and start with things in vector form

How do we represent a picture?
SHAPE$\circledast$ARROW + NUMBER$\circledast$ONE + DIRECTION$\circledast$UP- can do variations like this for all the pictures
- fairly standard with most assumptions about how people represent complex scenes
- but that part is not being modelled (yet!)
We have shown that it's possible to build these sorts of representations up directly from visual stimuli
- With a very simple vision system that can only recognize a few different shapes
- And where items have to be shown sequentially as it has no way of moving its eyes
Examples in Spaun¶
- Let's look at a simpler but related example:
- Binding used for building a list to remember
from IPython.display import YouTubeVideo
YouTubeVideo('U_Q6Xjz9QHg', width=720, height=400, loop=1, autoplay=0, playlist='U_Q6Xjz9QHg')
- The memory of the list is built up by using a basal ganglia action selection system to control feeding values into an integrator
- The thought bubble shows how close the decoded values are to the ideal
- Notice the forgetting!
- The same system can be used to do a version of the Raven's Matrices task
from IPython.display import YouTubeVideo
YouTubeVideo('Q_LRvnwnYp8', width=720, height=400, loop=1, autoplay=0, playlist='Q_LRvnwnYp8')
S1 = ONE$\circledast$P1S2 = ONE$\circledast$P1 + ONE$\circledast$P2S3 = ONE$\circledast$P1 + ONE$\circledast$P2 + ONE$\circledast$P3S4 = FOUR$\circledast$P1S5 = FOUR$\circledast$P1 + FOUR$\circledast$P2S6 = FOUR$\circledast$P1 + FOUR$\circledast$P2 + FOUR$\circledast$P3S7 = FIVE$\circledast$P1S8 = FIVE$\circledast$P1 + FIVE$\circledast$P2what is
S9?
How does Spaun make a guess at the end?
Let's figure out what the transformation is
T1 = S2$\circledast$S1'T2 = S3$\circledast$S2'T3 = S5$\circledast$S4'T4 = S6$\circledast$S5'T5 = S8$\circledast$S7'T = (T1 + T2 + T3 + T4 + T5)/5S9 = S8$\circledast$TS9 = FIVE$\circledast$P1 + FIVE$\circledast$P2 + FIVE$\circledast$P3This becomes a novel way of manipulating structured information
- Exploiting the fact that it is a vector underneath
- A spiking neural model applied to the study of human performance and cognitive decline on Raven's Advanced Progressive Matrices
Using Nengo¶
- How can we use Nengo to perform these kinds of operations?
- There's a 'SPA' module you can import, and it lets you do all kinds of this stuff
- Let's try answering simple questions about the bindings in a representation
- I just stole this from the 'Question answering' example in Nengo
%pylab inline
import nengo
from nengo import spa
# Number of dimensions for the Semantic Pointers
dimensions = 32
model = spa.SPA(label="Simple question answering")
with model:
model.color_in = spa.Buffer(dimensions=dimensions)
model.shape_in = spa.Buffer(dimensions=dimensions)
model.conv = spa.Buffer(dimensions=dimensions)
model.cue = spa.Buffer(dimensions=dimensions)
model.out = spa.Buffer(dimensions=dimensions)
# Connect the buffers
cortical_actions = spa.Actions(
'conv = color_in * shape_in',
'out = conv * ~cue'
)
model.cortical = spa.Cortical(cortical_actions)
The input will switch every 0.5 seconds between RED and BLUE. In the same way the shape input switches between CIRCLE and SQUARE. Thus, the network will bind alternatingly RED * CIRCLE and BLUE * SQUARE for 0.5 seconds each.
The cue for deconvolving bound semantic pointers cycles through CIRCLE, RED, SQUARE, and BLUE within one second.
def color_input(t):
if (t // 0.5) % 2 == 0:
return 'RED'
else:
return 'BLUE'
def shape_input(t):
if (t // 0.5) % 2 == 0:
return 'CIRCLE'
else:
return 'SQUARE'
def cue_input(t):
sequence = ['0', 'CIRCLE', 'RED', '0', 'SQUARE', 'BLUE']
idx = int((t // (1. / len(sequence))) % len(sequence))
return sequence[idx]
with model:
model.inp = spa.Input(color_in=color_input, shape_in=shape_input, cue=cue_input)
with model:
model.config[nengo.Probe].synapse = nengo.Lowpass(0.03)
color_in = nengo.Probe(model.color_in.state.output)
shape_in = nengo.Probe(model.shape_in.state.output)
cue = nengo.Probe(model.cue.state.output)
conv = nengo.Probe(model.conv.state.output)
out = nengo.Probe(model.out.state.output)
from nengo_gui.ipython import IPythonViz
IPythonViz(model, "configs/blue_red_spa.py.cfg")
sim = nengo.Simulator(model)
sim.run(3.)
plt.figure(figsize=(10, 10))
vocab = model.get_default_vocab(dimensions)
plt.subplot(5, 1, 1)
plt.plot(sim.trange(), model.similarity(sim.data, color_in))
plt.legend(model.get_output_vocab('color_in').keys, fontsize='x-small')
plt.ylabel("color")
plt.subplot(5, 1, 2)
plt.plot(sim.trange(), model.similarity(sim.data, shape_in))
plt.legend(model.get_output_vocab('shape_in').keys, fontsize='x-small')
plt.ylabel("shape")
plt.subplot(5, 1, 3)
plt.plot(sim.trange(), model.similarity(sim.data, cue))
plt.legend(model.get_output_vocab('cue').keys, fontsize='x-small')
plt.ylabel("cue")
plt.subplot(5, 1, 4)
for pointer in ['RED * CIRCLE', 'BLUE * SQUARE']:
plt.plot(sim.trange(), vocab.parse(pointer).dot(sim.data[conv].T), label=pointer)
plt.legend(fontsize='x-small')
plt.ylabel("convolved")
plt.subplot(5, 1, 5)
plt.plot(sim.trange(), spa.similarity(sim.data[out], vocab))
plt.legend(model.get_output_vocab('out').keys, fontsize='x-small')
plt.ylabel("output")
plt.xlabel("time [s]");
sum(ens.n_neurons for ens in model.all_ensembles) #Total number of neurons
35200