OA Oyster Proteome Data VeriCheck¶
Supplemental file check¶
2fold and uniques
5folds
5fold
In [ ]:
In [ ]:
In [ ]:
In [ ]:
In [ ]:
SKYLINE DATA Check¶
uploaded raw to SQLSHare
Query
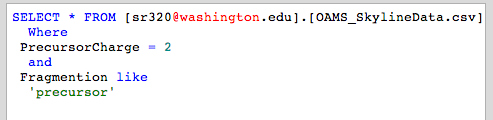
Still has duplicate peptides
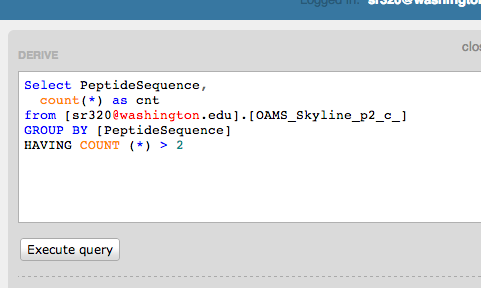
Select PeptideSequence,
count(*) as cnt
from [sr320@washington.edu].[OAMS_Skyline_p2_c_]
GROUP BY [PeptideSequence]
HAVING COUNT (*) > 2
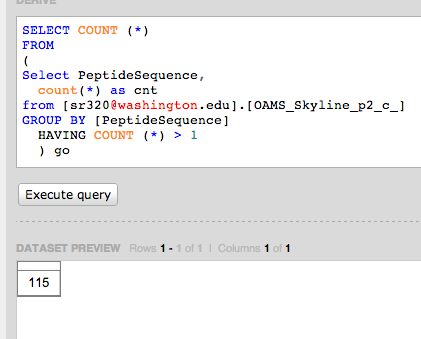
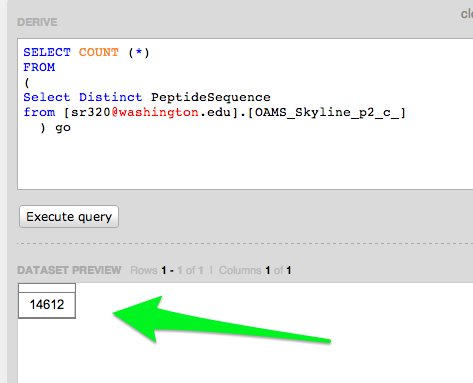
Emma's Data
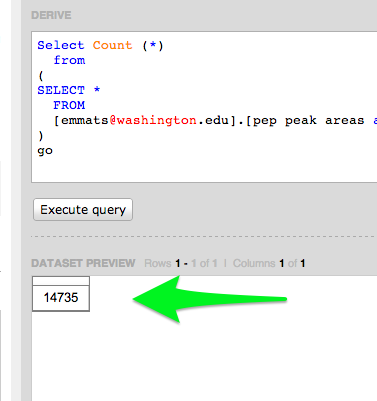
Query to verify
Select Count (*)
from
(
SELECT PeptideSequence,
Count (*) as cnt
FROM
[emmats@washington.edu].[pep peak areas all oysters2.txt]
Group by PeptideSequence
Having Count (*) > 1
)
go
In [2]:
!blastp -query /Volumes/web/cnidarian/OAMS_skyline_pep.fa -db /Volumes/Bay3/Software/ncbi-blast-2.2.26\+/db/oyster_v9_aa -outfmt 6 -max_target_seqs 1 -out /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9.txt
Warning: lcl|Query_1 ID: Warning: One or more U or O characters replaced by X for alignment score calculations at positions 10
In [4]:
!head /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9.txt
1 CGI_10010163 100.00 14 0 0 1 14 108 121 0.010 30.8 2 CGI_10003288 100.00 10 0 0 1 10 228 237 1.0 24.6 3 CGI_10015115 100.00 10 0 0 1 10 762 771 0.62 25.4 4 CGI_10007640 100.00 10 0 0 1 10 1144 1153 1.0 25.0 5 CGI_10018366 100.00 10 0 0 1 10 106 115 1.6 23.9 6 CGI_10000265 100.00 24 0 0 1 24 134 157 7e-11 52.8 7 CGI_10014468 100.00 11 0 0 1 11 1027 1037 0.49 25.8 8 CGI_10014915 100.00 17 0 0 1 17 579 595 6e-06 40.8 9 CGI_10022396 100.00 10 0 0 1 10 315 324 0.36 26.2 10 CGI_10001357 100.00 10 0 0 1 10 241 250 0.034 29.3
In [5]:
!wc /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9.txt
13078 156936 724094 /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9.txt
In [3]:
!blastp -query /Volumes/web/cnidarian/OAMS_skyline_pep.fa -db /Volumes/Bay3/Software/ncbi-blast-2.2.26\+/db/oyster_v9_aa -outfmt 6 -max_target_seqs 4 -out /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9_b.txt
Warning: lcl|Query_1 ID: Warning: One or more U or O characters replaced by X for alignment score calculations at positions 10
In [7]:
!tail /Volumes/web/cnidarian/OAMS_skyline_Blastp_oysterb9_b.txt
14610 CGI_10003195 70.59 17 5 0 1 17 768 784 0.60 26.2 14610 CGI_10001323 70.59 17 5 0 1 17 143 159 0.41 26.6 14610 CGI_10018755 70.59 17 5 0 1 17 659 675 0.49 26.2 14610 CGI_10013230 70.59 17 5 0 1 17 1007 1023 0.52 26.2 14611 CGI_10003521 100.00 17 0 0 1 17 390 406 2e-05 39.7 14611 CGI_10003521 57.14 14 6 0 3 16 1468 1481 4.3 23.5 14611 CGI_10003521 53.85 13 6 0 4 16 606 618 9.2 22.3 14611 CGI_10008439 41.18 17 10 0 1 17 1105 1121 8.9 22.3 14612 CGI_10017178 100.00 17 0 0 1 17 458 474 1e-06 43.1 14612 CGI_10001407 100.00 17 0 0 1 17 385 401 1e-06 42.7
In [8]:
!head /Volumes/web/oyster/proteomics/supplemental\ data/Supp\ Data\ sig\ qvalue\ proteins2.txt
Treatment comparison Protein Fold change under stress p-value q-value SwissProt Accession BLASTx e-value SwissProt Annotation 400 vs. 2800 µatm CGI_10000591 0.68 0.0036 0.19 Q71LX4 1.00E-77 Talin-2 400 vs. 2800 µatm CGI_10001585 2.42 0.0023 0.16 Q9W0E8 4.00E-126 Protein zer-1 homolog 400 vs. 2800 µatm CGI_10002289 0.51 0.0041 0.19 Q9Y490 3.00E-76 Talin-1 400 vs. 2800 µatm CGI_10002361 0.34 0.0015 0.16 P08011 1.00E-39 Microsomal glutathione S-transferase 1 400 vs. 2800 µatm CGI_10003022 0.56 0.0066 0.20 Q9VN14 7.00E-13 Contactin 400 vs. 2800 µatm CGI_10003145 0.56 0.0053 0.20 Q8NDM7 0 WD repeat-containing protein 96 400 vs. 2800 µatm CGI_10003812 0.44 0.0021 0.16 Q32NR4 5.00E-94 Tetratricopeptide repeat protein 29 400 vs. 2800 µatm CGI_10003897 0.74 0.0042 0.19 P18652 0 Ribosomal protein S6 kinase 2 alpha 400 vs. 2800 µatm CGI_10003914 0.67 0.0058 0.20 Q9Y490 2.00E-177 Talin-1
In [6]:
#probably not going to work with %
!head /Volumes/web/oyster/proteomics/supplemental\ data/Supp\ Data\ CGI_KEGG_association.txt
CGI_10028931 hsa:54847 0 642
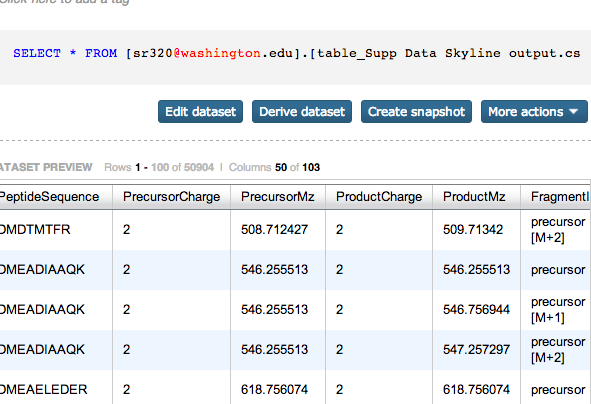
Peptides¶
In [4]:
from pandas import *
# read data from data file into a pandas DataFrame
oams_in = read_csv("/Users/sr320/Desktop/Supp Data Skyline output.csv") # name of the data file
#sep="\t", # what character separates each column?
#na_values=["", " ",]) # what values should be considered "blank" values?
In [5]:
print oams_in
<class 'pandas.core.frame.DataFrame'> Int64Index: 50904 entries, 0 to 50903 Columns: 103 entries, ProteinName to 8_03 BestRetentionTime dtypes: float64(2), int64(48), object(53)
In [6]:
oams_in.head()
Out[6]:
<class 'pandas.core.frame.DataFrame'> Int64Index: 5 entries, 0 to 4 Columns: 103 entries, ProteinName to 8_03 BestRetentionTime dtypes: float64(2), int64(48), object(53)
In [7]:
oams_in.count()
Out[7]:
ProteinName 50904 PeptideSequence 50904 PrecursorCharge 50904 PrecursorMz 50904 ProductCharge 50904 ProductMz 50904 FragmentIon 50904 11_01 TotalArea 50904 11_01 BestRetentionTime 50904 11_02 TotalArea 50904 11_02 BestRetentionTime 50904 11_03 TotalArea 50904 11_03 BestRetentionTime 50904 2_01 TotalArea 50904 2_01 BestRetentionTime 50904 ... 35_02 BestRetentionTime 50904 35_03 TotalArea 50904 35_03 BestRetentionTime 50904 5_01 TotalArea 50904 5_01 BestRetentionTime 50904 5_02 TotalArea 50904 5_02 BestRetentionTime 50904 5_03 TotalArea 50904 5_03 BestRetentionTime 50904 8_01 TotalArea 50904 8_01 BestRetentionTime 50904 8_02 TotalArea 50904 8_02 BestRetentionTime 50904 8_03 TotalArea 50904 8_03 BestRetentionTime 50904 Length: 103, dtype: int64
In [37]:
oams3.sum()
Out[37]:
protein CGI_10000013CGI_10000033CGI_10000045CGI_100000... CG2 1.460698e+11 CG5 1.069723e+11 CG8 9.31888e+10 CG11 1.052842e+11 CG26 1.281632e+11 CG29 7.01565e+10 CG32 5.606221e+10 CG35 8.597555e+10 CG221 1.472379e+11 CG224 1.213833e+11 CG227 1.531762e+11 CG230 9.807976e+10 CG242 7.674892e+10 CG245 1.392999e+11 CG248 1.274987e+11 CG251 1.218495e+11
In [38]:
oams3.mean()
Out[38]:
CG2 54564747.455610 CG5 39959762.291869 CG8 34810907.746420 CG11 39329175.735815 CG26 47875685.008945 CG29 26207134.783028 CG32 20942177.548209 CG35 32116380.094820 CG221 55001079.517204 CG224 45343045.513655 CG227 57219348.992819 CG230 36637939.460258 CG242 28669748.334060 CG245 52035818.414249 CG248 47627464.703441 CG251 45517176.820778
In [39]:
oams3.dropna()
Out[39]:
<class 'pandas.core.frame.DataFrame'> Int64Index: 2677 entries, 0 to 3379 Data columns: protein 2677 non-null values CG2 2677 non-null values CG5 2677 non-null values CG8 2677 non-null values CG11 2677 non-null values CG26 2677 non-null values CG29 2677 non-null values CG32 2677 non-null values CG35 2677 non-null values CG221 2677 non-null values CG224 2677 non-null values CG227 2677 non-null values CG230 2677 non-null values CG242 2677 non-null values CG245 2677 non-null values CG248 2677 non-null values CG251 2677 non-null values dtypes: float64(16), object(1)
Spectral Count Method¶
In [ ]:
48 files that look like.... .
<img src="http://eagle.fish.washington.edu/cnidarian/skitch/SQLShare_-_View_Query_17B02CD9.png" alt="SQLShare_-_View_Query_17B02CD9.png"/>
In [20]:
!fetchdata.py -d ([emmats@washington.edu].[CG2 unique peps > 1]) -f tsv -o /Volumes/web/cnidarian/OA_.txt
/bin/sh: -c: line 0: syntax error near unexpected token `('
/bin/sh: -c: line 0: `fetchdata.py -d ([emmats@washington.edu].[CG2 unique peps > 1]) -f tsv -o /Volumes/web/cnidarian/OA_.txt'
In [48]:
!fetchdata.py -d "[emmats@washington.edu].[CG251 unique peps >1]" -f tsv -o /Volumes/web/cnidarian/OA_MS_CG_251_upep.txt
In [35]:
!head /Volumes/web/cnidarian/OA_MS_CG_11_upep.txt
In [49]:
!fetchdata.py -d "[emmats@washington.edu].[NSAF all oysters]" -f tsv -o /Volumes/web/cnidarian/OA_MS_NSAFall.txt
In [50]:
!head /Volumes/web/cnidarian/OA_MS_NSAFall.txt
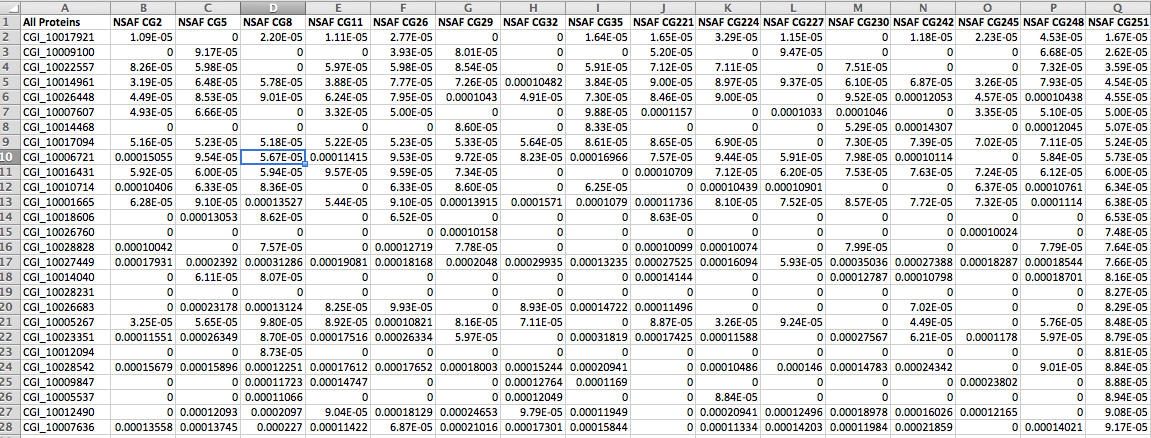
Spectral Count data 080813¶
In [1]:
!head /Users/sr320/Desktop/NSAF\ avg\ SpC.csv
CGI_10028023,0.00126516,0.000769597,0.001270981,0,0,0.001045932,0.001660575,0,0.001526871,0,0,0,0.001903786,0,0,0
DAVID Enrichment Analysis¶
Comparing OA¶
In [2]:
#Supplemental file 4
!head /Volumes/web/oyster/proteomics/supplemental\ data/Supplementary\ Table\ S4.txt
CGI_10028902 0 0.00070663 0 0.000587183 0 0 0 0 0 0 0 0.000492864 0.000624293 0 0 0.000825286 0.000323453 0 0.000123216 0.000362395 2.625091303 2.941133862 0 *
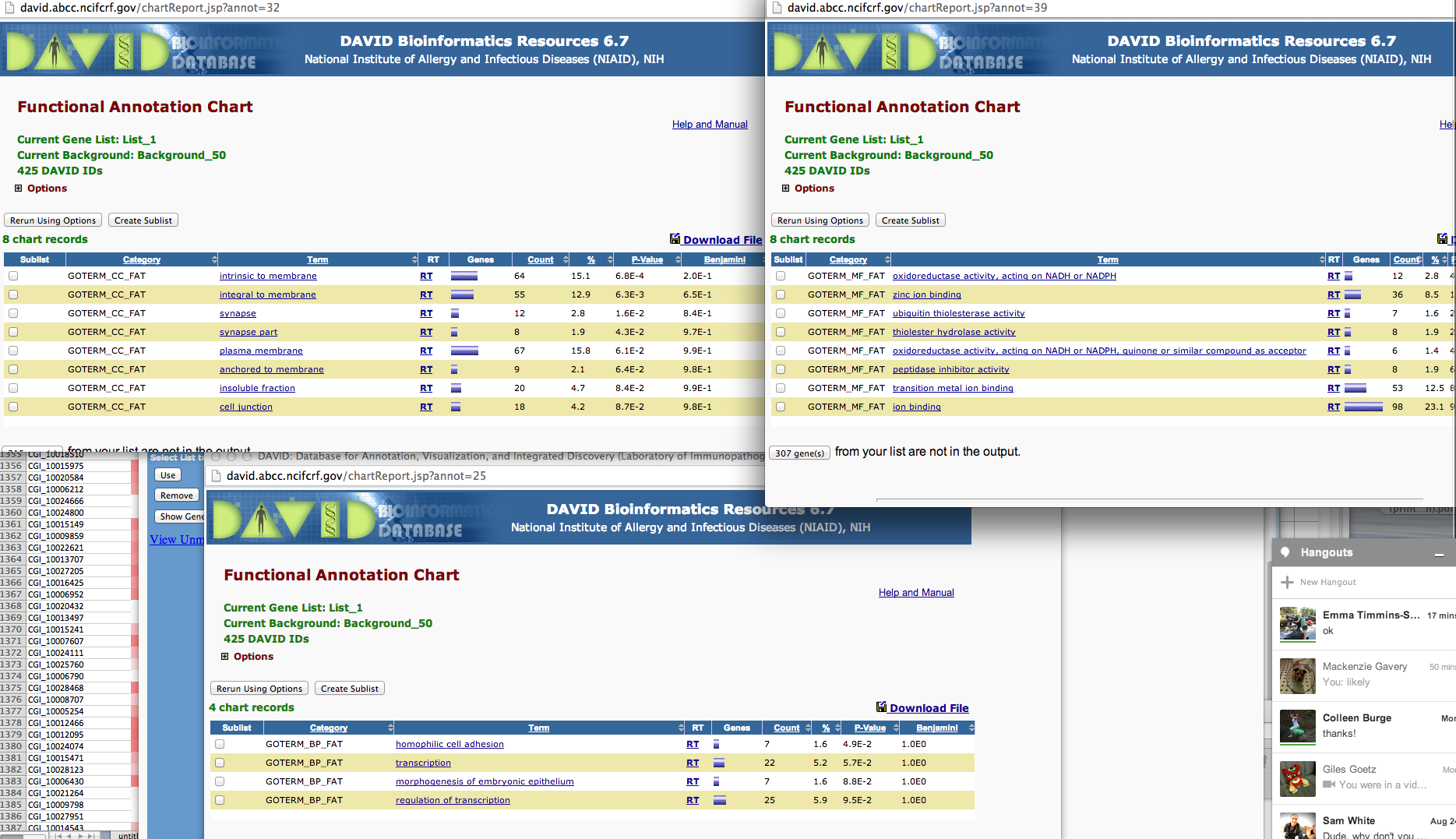
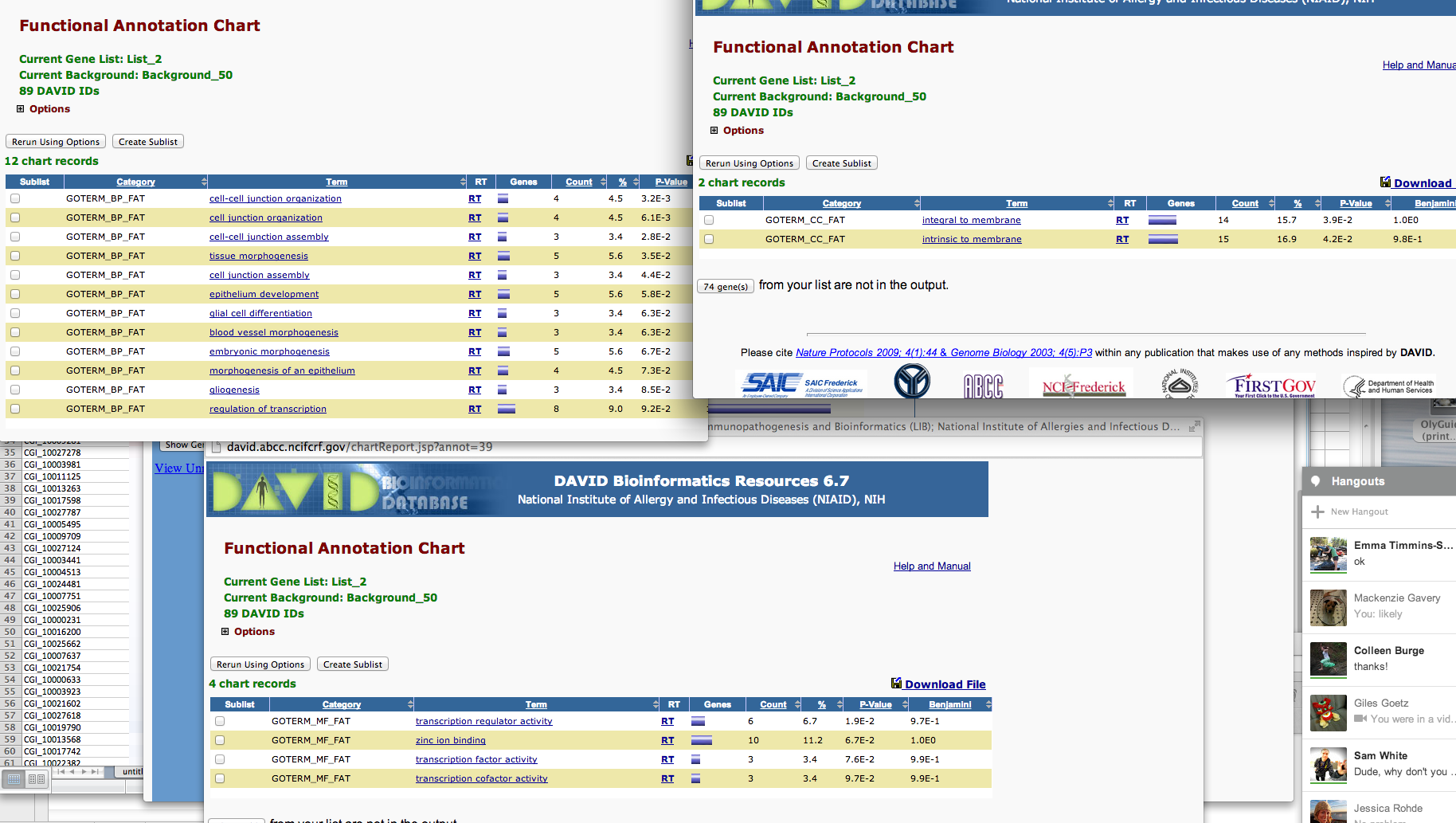
SPIDs from the OA 5-fold column
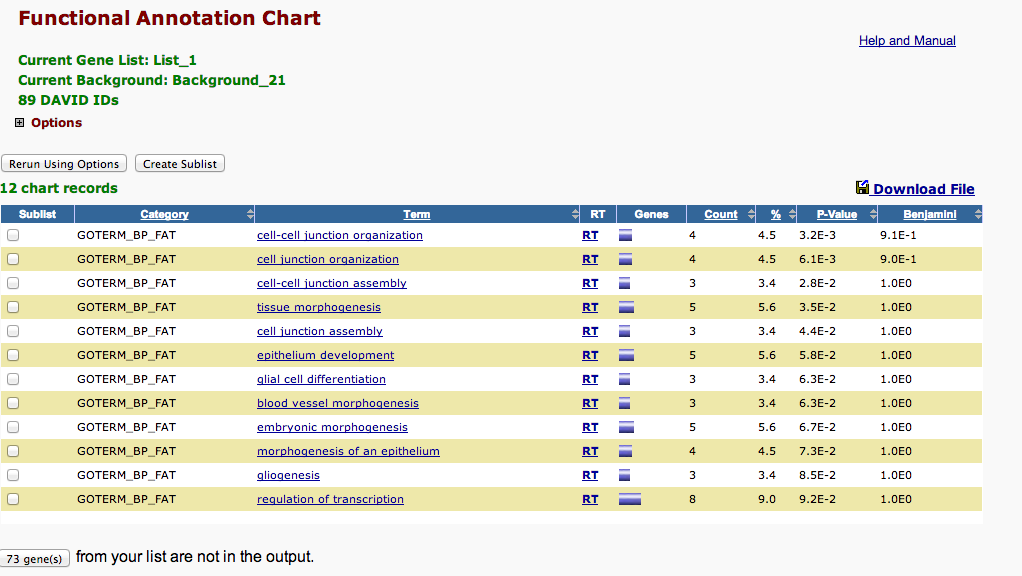
In [42]:
#file
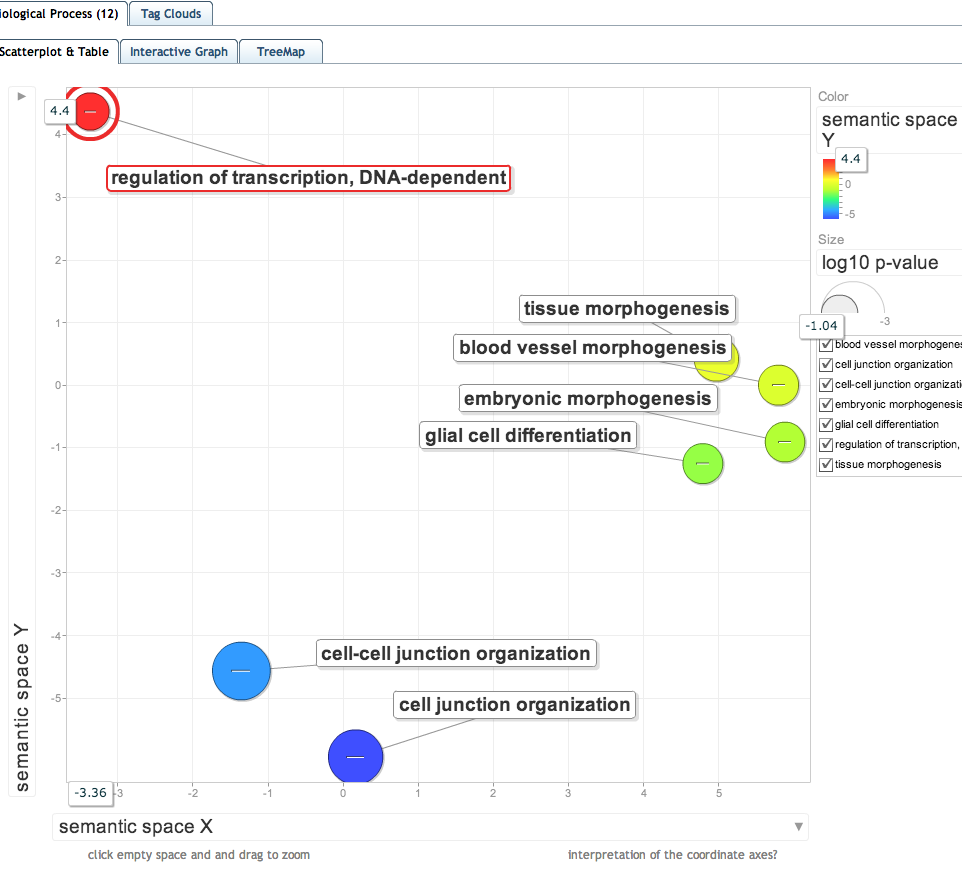
!head /Volumes/web/cnidarian/chart_B1049AF0BD891379525818063.txt
Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_BP_FAT GO:0045216~cell-cell junction organization 4 4.49438202247191 0.003215314108718092 Q01989, Q94887, Q9Y490, Q9VN14 70 5 959 10.96 0.9080385953446322 0.9080385953446322 4.7758884843944305 GOTERM_BP_FAT GO:0034330~cell junction organization 4 4.49438202247191 0.006099557106516472 Q01989, Q94887, Q9Y490, Q9VN14 70 6 959 9.133333333333333 0.9892581980951891 0.8963573354992699 8.877881028273594 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.3707865168539324 0.02785621995858589 Q94887, Q9Y490, Q9VN14 70 4 959 10.275 0.9999999991903415 0.9990679612912189 34.903309791067706 GOTERM_BP_FAT GO:0048729~tissue morphogenesis 5 5.617977528089887 0.03480063151772505 Q01989, P46531, Q94887, Q5XIH7, P11584 70 18 959 3.805555555555555 0.9999999999960077 0.9985864712773175 41.62214279701073 GOTERM_BP_FAT GO:0034329~cell junction assembly 3 3.3707865168539324 0.04428349052518013 Q94887, Q9Y490, Q9VN14 70 5 959 8.22 0.9999999999999973 0.9987844554464399 49.755240844743675 GOTERM_BP_FAT GO:0060429~epithelium development 5 5.617977528089887 0.057813930529541334 Q01989, Q94887, Q9VN14, Q5XIH7, P11584 70 21 959 3.2619047619047614 1.0 0.9993604421596898 59.54296565749193 GOTERM_BP_FAT GO:0010001~glial cell differentiation 3 3.3707865168539324 0.06337545723447832 P46531, Q94887, Q9VN14 70 6 959 6.85 1.0 0.9990227433649204 63.02363443000699 GOTERM_BP_FAT GO:0048514~blood vessel morphogenesis 3 3.3707865168539324 0.06337545723447832 P46531, Q7Z3E1, P19971 70 6 959 6.85 1.0 0.9990227433649204 63.02363443000699 GOTERM_BP_FAT GO:0048598~embryonic morphogenesis 5 5.617977528089887 0.06693586709090873 Q01989, P46531, Q94887, P58365, P11584 70 22 959 3.1136363636363633 1.0 0.9983667515177859 65.1028231919163
In [5]:
from IPython.display import YouTubeVideo
YouTubeVideo('3ZnhGLGVjk')
Out[5]:
In [43]:
!perl /Volumes/web/cnidarian/david_excelfix.pl /Volumes/web/cnidarian/chart_B1049AF0BD891379525818063.txt > /Volumes/web/cnidarian/mod_chart_B1049AF0BD891379525818063.txt
In [44]:
!head /Volumes/web/cnidarian/mod_chart_B1049AF0BD891379525818063.txt
Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_BP_FAT GO:0045216~cell-cell junction organization 4 4.49438202247191 0.003215314108718092 Q01989 70 5 959 10.96 0.9080385953446322 0.9080385953446322 4.7758884843944305 GOTERM_BP_FAT GO:0045216~cell-cell junction organization 4 4.49438202247191 0.003215314108718092 Q94887 70 5 959 10.96 0.9080385953446322 0.9080385953446322 4.7758884843944305 GOTERM_BP_FAT GO:0045216~cell-cell junction organization 4 4.49438202247191 0.003215314108718092 Q9Y490 70 5 959 10.96 0.9080385953446322 0.9080385953446322 4.7758884843944305 GOTERM_BP_FAT GO:0045216~cell-cell junction organization 4 4.49438202247191 0.003215314108718092 Q9VN14 70 5 959 10.96 0.9080385953446322 0.9080385953446322 4.7758884843944305 GOTERM_BP_FAT GO:0034330~cell junction organization 4 4.49438202247191 0.006099557106516472 Q01989 70 6 959 9.133333333333333 0.9892581980951891 0.8963573354992699 8.877881028273594 GOTERM_BP_FAT GO:0034330~cell junction organization 4 4.49438202247191 0.006099557106516472 Q94887 70 6 959 9.133333333333333 0.9892581980951891 0.8963573354992699 8.877881028273594 GOTERM_BP_FAT GO:0034330~cell junction organization 4 4.49438202247191 0.006099557106516472 Q9Y490 70 6 959 9.133333333333333 0.9892581980951891 0.8963573354992699 8.877881028273594 GOTERM_BP_FAT GO:0034330~cell junction organization 4 4.49438202247191 0.006099557106516472 Q9VN14 70 6 959 9.133333333333333 0.9892581980951891 0.8963573354992699 8.877881028273594 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.3707865168539324 0.02785621995858589 Q94887 70 4 959 10.275 0.9999999991903415 0.9990679612912189 34.903309791067706
In [ ]:
#join in SQLShare
In [45]:
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d OA_enrich2 /Volumes/web/cnidarian/mod_chart_B1049AF0BD891379525818063.txt
processing chunk line 0 to 51 (0.00206899642944 s elapsed) pushing /Volumes/web/cnidarian/mod_chart_B1049AF0BD891379525818063.txt... parsing C8BE2D77... finished OA_enrich2
In [46]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[OA_enrich2]g left join [sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]sp on g.Genes = sp.SPID" -o /Volumes/web/cnidarian/OA_enrich2_join_SPID.csv
In [47]:
!head /Volumes/web/cnidarian/OA_enrich2_join_SPID.csv
In [48]:
!head /Volumes/web/cnidarian/OA_enrich2_join_SPID_sig.csv
Category,Term,Count,%,PValue,Genes,List Total,Pop Hits,Pop Total,Fold Enrichment,Bonferroni,Benjamini,FDR,CGI_ID,SPID,evalue,Description GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9Y490,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10000332,Q9Y490,3.00E-38,Talin-1 GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9Y490,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10002289,Q9Y490,6.00E-69,Talin-1 GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9VN14,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10002691,Q9VN14,9.00E-56,Contactin GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9VN14,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10003022,Q9VN14,6.00E-13,Contactin GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9VN14,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10006138,Q9VN14,1.00E-145,Contactin GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9Y490,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10011964,Q9Y490,0,Talin-1 GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9Y490,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10012794,Q9Y490,7.00E-37,Talin-1 GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q01989,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10013678,Q01989,0,Myosin heavy chain 95F GOTERM_BP_FAT,GO:0045216~cell-cell junction organization,4,4.494382022,0.003215314,Q9VN14,70,5,959,10.96,0.908038595,0.908038595,4.775888484,CGI_10014483,Q9VN14,3.00E-106,Contactin
Comparing mechanical stress at 400uatm¶
No enriched processes
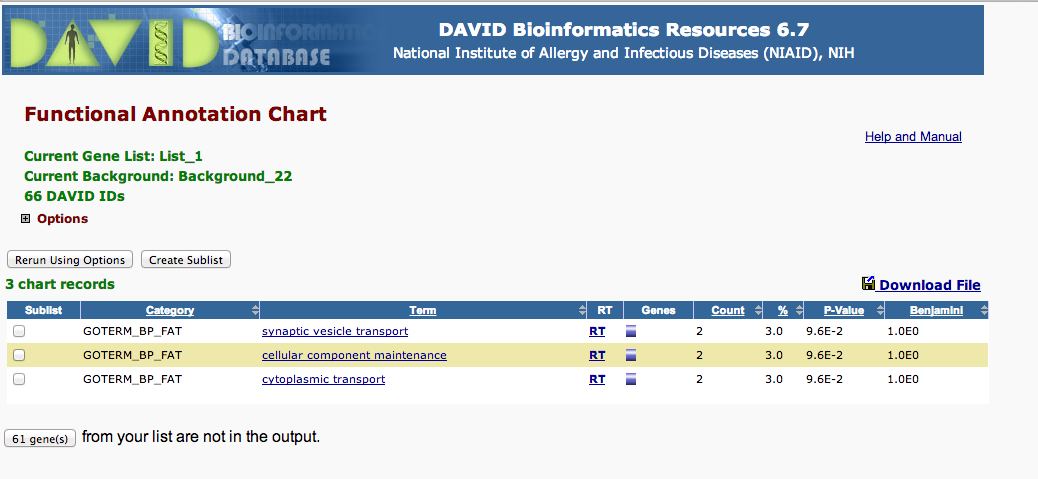
2800 mechanincal Stress¶
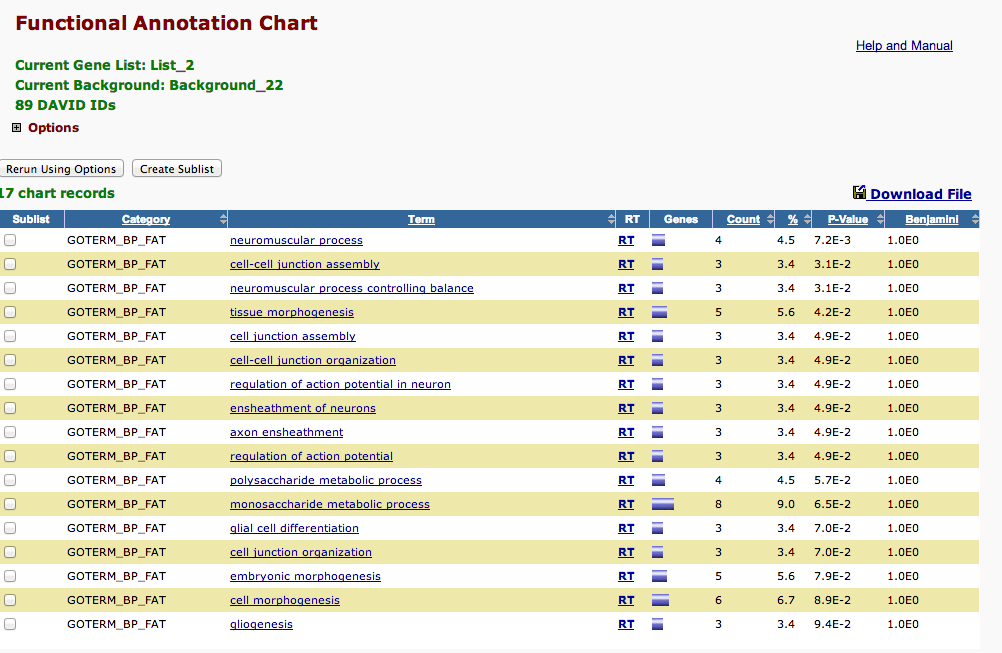
In [3]:
!head /Volumes/web/cnidarian/chart_976F753C30211379944509986.txt
Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.49438202247191 0.007169536737378732 Q9MYM4, P58365, P07686, P26232 74 6 959 8.63963963963964 0.9964256357671437 0.9964256357671437 10.428890558189362 GOTERM_BP_FAT GO:0050885~neuromuscular process controlling balance 3 3.3707865168539324 0.031023648581194604 Q9MYM4, P58365, P07686 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.3707865168539324 0.031023648581194604 Q94887, Q9Y490, Q9VN14 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0048729~tissue morphogenesis 5 5.617977528089887 0.04175586476702081 Q9MYM4, P46531, Q94887, Q5XIH7, P08630 74 18 959 3.59984984984985 0.9999999999999969 0.9999853687865333 47.94517596118537 GOTERM_BP_FAT GO:0008366~axon ensheathment 3 3.3707865168539324 0.0491782737898529 Q94887, Q9VN14, P07686 74 5 959 7.775675675675677 1.0 0.9999483704675248 53.786476584957185 GOTERM_BP_FAT GO:0034329~cell junction assembly 3 3.3707865168539324 0.0491782737898529 Q94887, Q9Y490, Q9VN14 74 5 959 7.775675675675677 1.0 0.9999483704675248 53.786476584957185 GOTERM_BP_FAT GO:0019228~regulation of action potential in neuron 3 3.3707865168539324 0.0491782737898529 Q94887, Q9VN14, P07686 74 5 959 7.775675675675677 1.0 0.9999483704675248 53.786476584957185 GOTERM_BP_FAT GO:0001508~regulation of action potential 3 3.3707865168539324 0.0491782737898529 Q94887, Q9VN14, P07686 74 5 959 7.775675675675677 1.0 0.9999483704675248 53.786476584957185 GOTERM_BP_FAT GO:0007272~ensheathment of neurons 3 3.3707865168539324 0.0491782737898529 Q94887, Q9VN14, P07686 74 5 959 7.775675675675677 1.0 0.9999483704675248 53.786476584957185
</Volumes/web/cnidarian/chart_976F753C30211379944509986.txt>
In [4]:
!perl /Volumes/web/cnidarian/david_excelfix.pl /Volumes/web/cnidarian/chart_976F753C30211379944509986.txt > /Volumes/web/cnidarian/mod_chart_976F753C30211379944509986.txt
In [5]:
!head /Volumes/web/cnidarian/mod_chart_976F753C30211379944509986.txt
Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.49438202247191 0.007169536737378732 Q9MYM4 74 6 959 8.63963963963964 0.9964256357671437 0.9964256357671437 10.428890558189362 GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.49438202247191 0.007169536737378732 P58365 74 6 959 8.63963963963964 0.9964256357671437 0.9964256357671437 10.428890558189362 GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.49438202247191 0.007169536737378732 P07686 74 6 959 8.63963963963964 0.9964256357671437 0.9964256357671437 10.428890558189362 GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.49438202247191 0.007169536737378732 P26232 74 6 959 8.63963963963964 0.9964256357671437 0.9964256357671437 10.428890558189362 GOTERM_BP_FAT GO:0050885~neuromuscular process controlling balance 3 3.3707865168539324 0.031023648581194604 Q9MYM4 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0050885~neuromuscular process controlling balance 3 3.3707865168539324 0.031023648581194604 P58365 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0050885~neuromuscular process controlling balance 3 3.3707865168539324 0.031023648581194604 P07686 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.3707865168539324 0.031023648581194604 Q94887 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.3707865168539324 0.031023648581194604 Q9Y490 74 4 959 9.719594594594595 0.9999999999808037 0.999995618637879 38.269514414920444
In [6]:
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d 2800m_enrich2 /Volumes/web/cnidarian/mod_chart_976F753C30211379944509986.txt
processing chunk line 0 to 66 (0.000998973846436 s elapsed) pushing /Volumes/web/cnidarian/mod_chart_976F753C30211379944509986.txt... parsing 8723DBCB... finished 2800m_enrich2
In [7]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[2800m_enrich2]g left join [sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]sp on g.Genes = sp.SPID" -o /Volumes/web/cnidarian/2800m_enrich2_join_SPID.csv
In [8]:
!head /Volumes/web/cnidarian/2800m_enrich2_join_SPID.csv
.07 pvalue thrshold http://eagle.fish.washington.edu/cnidarian/2800m_enrich2_join_SPID_sig.xls
In [ ]:
Research
P07686
Q9MYM4
Q9VFC8
P13280
C3YWU0
Q9CQ60
P10155
Q5XIG6
In [10]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/P07686 width=700 height=350></iframe>')
Out[10]:
In [11]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/Q9MYM4 width=700 height=350></iframe>')
Out[11]:
In [12]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/Q9VFC8 width=700 height=350></iframe>')
Out[12]:
In [14]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/P13280 width=700 height=350></iframe>')
Out[14]:
In [15]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/C3YWU0 width=700 height=350></iframe>')
Out[15]:
In [16]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/Q9CQ60 width=700 height=350></iframe>')
Out[16]:
In [17]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/P10155 width=700 height=350></iframe>')
Out[17]:
In [18]:
from IPython.display import HTML
HTML('<iframe src=http://www.uniprot.org/uniprot/Q5XIG6 width=700 height=350></iframe>')
Out[18]:
redux lets add qvalue proteins not 5 fold into david P22695 Q00651¶
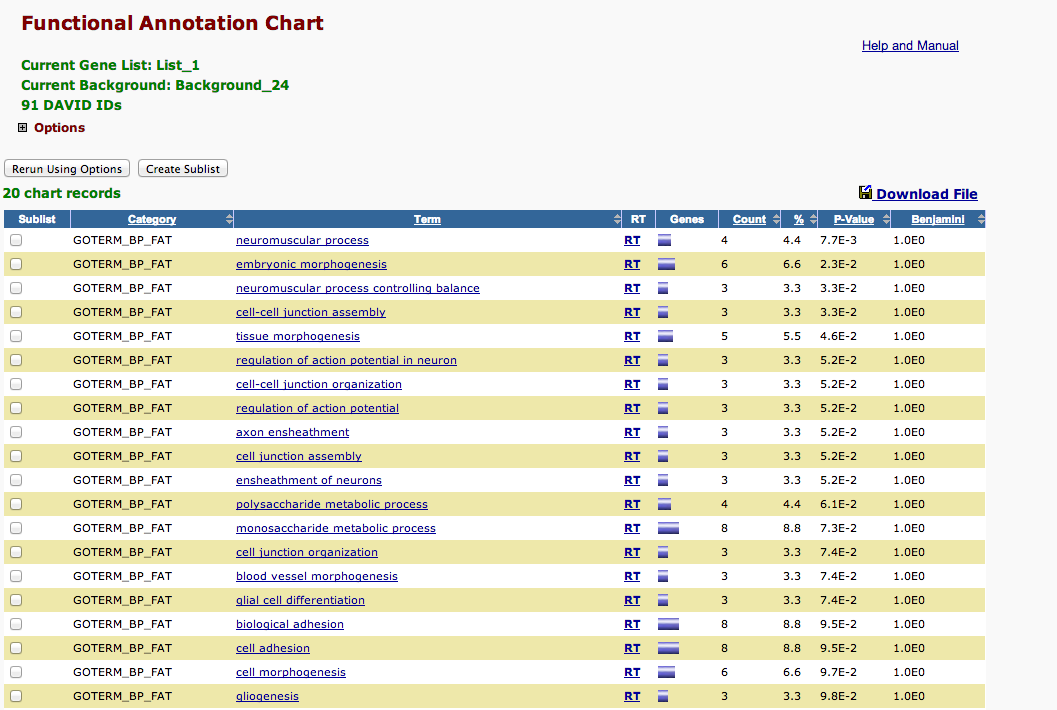
In [20]:
!head /Volumes/web/cnidarian/chart_307AE9BAF7BE1379947826112.txt
Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_BP_FAT GO:0050905~neuromuscular process 4 4.395604395604396 0.007745415012843447 Q9MYM4, P58365, P07686, P26232 76 6 959 8.412280701754385 0.9979488423930916 0.9979488423930916 11.243775404230384 GOTERM_BP_FAT GO:0048598~embryonic morphogenesis 6 6.593406593406594 0.023480583159572355 P46531, Q94887, Q00651, P58365, P11046, P08630 76 22 959 3.441387559808612 0.9999999938908196 0.9999218387540135 30.544682816797387 GOTERM_BP_FAT GO:0050885~neuromuscular process controlling balance 3 3.296703296703297 0.0326636560370076 Q9MYM4, P58365, P07686 76 4 959 9.463815789473683 0.999999999996691 0.9998509845372993 39.915842133228416 GOTERM_BP_FAT GO:0007043~cell-cell junction assembly 3 3.296703296703297 0.0326636560370076 Q94887, Q9Y490, Q9VN14 76 4 959 9.463815789473683 0.999999999996691 0.9998509845372993 39.915842133228416 GOTERM_BP_FAT GO:0048729~tissue morphogenesis 5 5.4945054945054945 0.04551929059176884 Q9MYM4, P46531, Q94887, Q5XIH7, P08630 76 18 959 3.505116959064327 0.9999999999999999 0.9999058838959778 51.063867887999635 GOTERM_BP_FAT GO:0019228~regulation of action potential in neuron 3 3.296703296703297 0.05170410612678131 Q94887, Q9VN14, P07686 76 5 959 7.571052631578947 1.0 0.9997864674362187 55.708473599859396 GOTERM_BP_FAT GO:0001508~regulation of action potential 3 3.296703296703297 0.05170410612678131 Q94887, Q9VN14, P07686 76 5 959 7.571052631578947 1.0 0.9997864674362187 55.708473599859396 GOTERM_BP_FAT GO:0045216~cell-cell junction organization 3 3.296703296703297 0.05170410612678131 Q94887, Q9Y490, Q9VN14 76 5 959 7.571052631578947 1.0 0.9997864674362187 55.708473599859396 GOTERM_BP_FAT GO:0007272~ensheathment of neurons 3 3.296703296703297 0.05170410612678131 Q94887, Q9VN14, P07686 76 5 959 7.571052631578947 1.0 0.9997864674362187 55.708473599859396
In [ ]:
!perl /Volumes/web/cnidarian/david_excelfix.pl /Volumes/web/cnidarian/chart_307AE9BAF7BE1379947826112.txt > /Volumes/web/cnidarian/mod_chart_307AE9BAF7BE1379947826112.txt
In [22]:
!python /Users/sr320/sqlshare-pythonclient/tools/singleupload.py -d 2800m_enrich3 /Volumes/web/cnidarian/mod_chart_307AE9BAF7BE1379947826112.txt
processing chunk line 0 to 86 (0.000899076461792 s elapsed) pushing /Volumes/web/cnidarian/mod_chart_307AE9BAF7BE1379947826112.txt... parsing 77722FAD... finished 2800m_enrich3
In [23]:
!python /Users/sr320/sqlshare-pythonclient/tools/fetchdata.py -s "SELECT * FROM [sr320@washington.edu].[2800m_enrich3]g left join [sr320@washington.edu].[qDOD Cgigas Gene Descriptions (Swiss-prot)]sp on g.Genes = sp.SPID" -o /Volumes/web/cnidarian/2800m_enrich3_join_SPID.csv
In [24]:
!head /Volumes/web/cnidarian/2800m_enrich3_join_SPID.csv
In [ ]: