A practical introduction to sensitivity analysis¶
Leif Rune Hellevik, Department of Structural Engineering, NTNU
Date: Jul 13, 2018
# ipython magic
%matplotlib notebook
%load_ext autoreload
%autoreload 2
# plot configuration
import matplotlib
import matplotlib.pyplot as plt
plt.style.use("ggplot")
# import seaborn as sns # sets another style
matplotlib.rcParams['lines.linewidth'] = 3
fig_width, fig_height = (7.0,5.0)
matplotlib.rcParams['figure.figsize'] = (fig_width, fig_height)
# import modules
import numpy as np
from numpy import linalg as LA
import chaospy as cp
from sensitivity_examples_nonlinear import generate_distributions
from monte_carlo import generate_sample_matrices_mc
from monte_carlo import calculate_sensitivity_indices_mc
import pandas as pd
from operator import index
Introduction¶
This practical introduction to sensitivity analysis is based on the presentation and examples found in [saltelli_global_2008]. To give the reader an even better hands on experience of the topic, we have integrated the computations in a python notebook format.
Many sensitivity analyses reported in the literature are based on derivatives at set point or point of interest. Indeed such approaches are based on the fact that the derivative of $\partial Y_i/\partial X_j$ of quantity of interest $Y_i$ as a function of an input variable $X_j$ can be thought of as the mathematical definition of the sensitivity of $Y_i$ versus $X_j$.
However, what is important to keep in mind is that local derivatives are only informative at the set point in the parameter space at which they are computed, and do not provide information for the rest of the parameter space. Naturally, such a linearisation will matter little for linear models, but for general, nonlinear models, care must be taken. In particular this is important in situations when the input parameters are uncertain.
Local versus global sensitivity analysis¶
Motivation and useful purposes of sensitivity analysis
Parameter prioritization of parameters of high sensitivity (importance)
Parameter fixation of parameters of low sensitivity (importance)
Reveal surprising relations/properties of the model
Indentify critical regions in the input parameter space
Local approaches based on derivatives¶
Many sensitivity analyses found in the scientific literature are based on derivatives. This fact has naturally a rational basis as the partial derivative $\partial y/\partial Z_i$ of a model predicion $y$ with respect to an input $Z_i$, can be understood as the mathematical representation of the sensitivity of $y$ with respect to $Z_i$.
Even though a local, partial derivative approach is computationallyXS inexpensive it has in general limited usage for nonlinear models. The derivatives are linearizations of the model sensitivities around the point in the parameter space at which they are evaluated, and may only be extrapolated to provide information on the sensitivity in other regions of the parameter space in the case of a linear model.
As a first motivation you may consider the ratio areas of circle with diameter $D$ to the corresponding square with sides $D$, as an optimistic approximation of the portion of the parameter space you might represent with a one factor at the time (OAT) approach:
Area ratio of circle to a square.
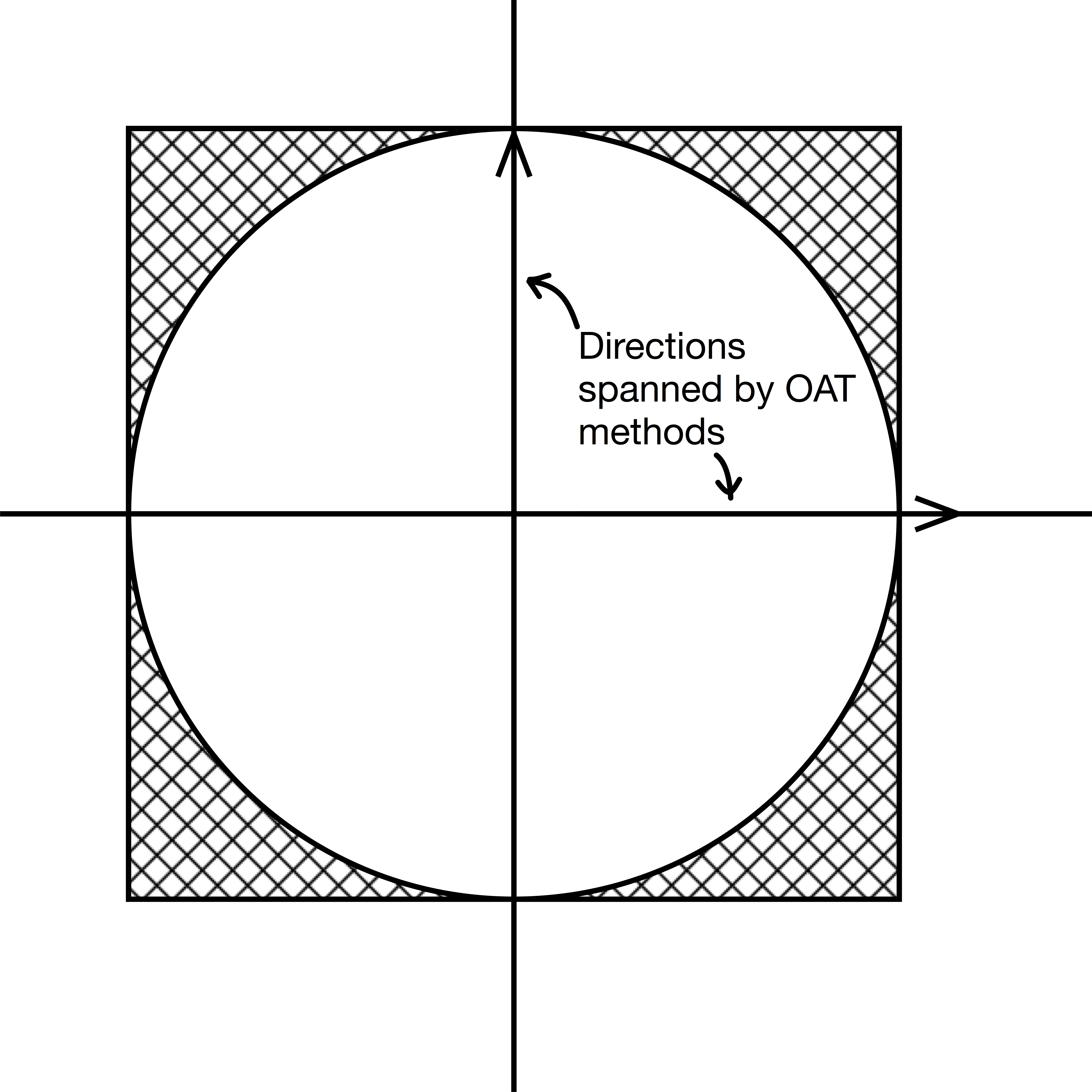
You may compute the corresonding ratio for a sphere and a cube yourself. For higher dimensions we provide a code snippet below which calculates the ratio of a hypersphere to a hypercube. This is you motivate why you should avoid "one factor at the time" analyses for sensitivity analysis in higher dimensions with multiple parameters.
# See https://en.wikipedia.org/wiki/N-sphere#Recurrences
%matplotlib nbagg
import ipywidgets as widgets
import matplotlib.pyplot as plt
import interactive_pie
from interactive_pie import update_pie
plt.suptitle('Why you should avoid "one factor at the time" in higher dimensions.')
w_dim = widgets.IntSlider(min=1, max=20, value=2, description='Dimensions')
widgets.interactive(update_pie, Ndim=w_dim)
Based on the brief motivation above we will present of methods based on the exploration of the input parameter space by judiciously selecting samples in that space. Such approaches result in more robust and informative sensitivity measures, than what would be the result from a local derivative app roach at the center of the parameter space.
To introduce the methods of sensitivity analysis, we shall start from derivatives and illustrate them on a very simple linear model.
A simple linear model¶
As an simple linear model example consider:
where the input factors are $\mathbf{X} = (\Omega_1, \Omega_2, \ldots, \Omega_r, Z_1, Z_2, \ldots, Z_r)$. For simplicity we assume that the model output $Y$ of (2) is a single variable and that the $\Omega s$ are fixed coefficients or weights.
Consequently, the true factors of (2) are just $(Z_1, Z_2, \ldots, Z_r)$. The individual variables $Z_i$ are taken to normally distributed with mean zero
As the predicted value $Y$ of the model in (2) is linear combination of normally distributed factors, it is easy to verify (see exercices in [saltelli_global_2008]) that $Y$ also will be normally distributed with:
Furthermore, we order the factors from the most certain to the less certain, i.e.:
# start the linear model
def linear_model(w, z):
return np.sum(w*z, axis=1)
To hold the mean and the standard deviation of all the input factors we use a numpy-array of size $r\times 2$, with one row per factor, where the first column holds the mean whereas the second column holds the standard deviation. The weights $\Omega_{1\ldots r}$ are stored in a numpy-vector.
# Set mean (column 0) and standard deviations (column 1) for each factor z. Nrv=nr. rows
Nrv = 4 # number of random variables
zm = np.array([[0., i] for i in range(1, Nrv + 1)])
# The above "list comprehension" is equivalent to the next for lines
# zm = np.zeros((Nrv, 2))
# zm[0, 1] = 1
# zm[1, 1] = 2
# zm[2, 1] = 3
# zm[3, 1] = 4
# Set the weight
c = 2
w = np.ones(Nrv) * c
We may now perform a Monte Carlo experiment on our model by generating $N$ samples from the distributions of each factor and an input sample is thus produced:
We may the compute a value of $Y$ from (2) for each column in (7) to produce a solution vector $\mathbf{Y}$. Having sampled $N$ values from each input factor we may produce $r$ scatter plots, by projecting in turn the $N$ values of $\mathbf{Y}$ against the $N$ values of each of the $r$ input factors.
# Generate distributions for each element in z and sample
Ns = 500
# jpdf = generate_distributions(zm)
pdfs = []
for i, z in enumerate(zm):
pdfs.append(cp.Normal(z[0], z[1]))
jpdf = cp.J(*pdfs)
# generate Z
Z = jpdf.sample(Ns)
# evaluate the model
Y = linear_model(w, Z.transpose())
print(np.var(Y))
# Scatter plots of data for visual inspection of sensitivity
fig=plt.figure()
for k in range(Nrv):
plt.subplot(2, 2, k + 1)
plt.plot(Z[k, :], Y[:], '.')
xlbl = 'Z' + str(k)
plt.xlabel(xlbl)
fig.tight_layout() # adjust subplot(s) to the figure area.
Note that the assumption of independent factors $Z_i$ allows us to sample
each $Z_i$ independently from its own marginal distribution. We store
all the samples for all the factors $Z_i$ in the the numpy array
Z[i,:], where $i$ corresponds to $Z_i$ as:
pdf.append(cp.Normal(z[0],z[1]))
Z[i,:]=pdf[i].sample(N)
From the scatterplots generated by the python code above we intuitively get the impression that $Y$ is more sensitive to $Z_4$ than to $Z_3$, and that $Y$ is more sensitive to $Z_3$ than to $Z_3$, and that we may order the factors my influence on $Y$ as:
Our intuitive notion of influence is based on that there is more shape (or better pattern) in the plot for $Z_4$ than for $Z_3$ and likewise.
For our simple linear model in (2) we are in the fortunate situation that we may compute the local derivatives analyticaly:
In our code example we set all the $\Omega_i=2$ for $i=1,\ldots,4$, and according to the local sensitivity meansure $S_{Z_i}^{p}$ in (10) all the input factors $Z_i$s are equally important and independent of the variation of each factor. This measure is clearly at odds with the ranking of influence based on the scatterplots in (9) and is an indication of the usefullness of scatterplots in sensititivy analysis. However, the bidimensional scatterplots may in some cases be deceiving and lead to type II errors (i.e. failure to identify influential parameters). ref to Saltelli 2004...
Most sensitivity measures aim to preserve the rich information provided by the scatterplots in a condensed format. The challenge is how to rank the factors rapidly and automatically without having to inspect many scatterplots in situations with many input factors. Another challenge with scatterplots is that sensitivities for sets cannot be visualized, while luckily compact sensitivity measures may be defined in such cases.
Normalized derivatives¶
A simple way to improve the derivative sensitivity measure $S_{Z_i}^{p}$ in (10) is to scale the input-output variables with their standard deviations:
Based on the linearity of our model we previously found (5) which also yields:
The normalized derivative measure of sensitivity in (11) is more convincing than (10): first, as it involves both the weights $\Omega_i$ and the factors $Z_i$ in (2); second as the measures are properly scaled and summarizes to one, which allows for an easy interpretation of the output sensitivity with respect to each of the input factors.
# Theoretical sensitivity indices
std_y = np.sqrt(np.sum((w * zm[:, 1])**2))
s = w * zm[:,1]/std_y
print("\nTheoretical sensitivity indices\n")
row_labels= ['S_'+str(idx) for idx in range(1,Nrv+1)]
print(pd.DataFrame(s**2, columns=['S analytic'],index=row_labels).round(3))
Based on samples of the random input variables and subsequent model evaluations, we may estimate the standard deviation of $\mathbf{Y}$ and compute the relative error with respect to the theoretical value. You may change the number of sample above, i.e. $N$, and see how $N$ influence the estimates.
# Expectation and variance from sampled values
print("Expectation and std from sampled values\n")
print('std(Y)={:2.3f} and relative error={:2.3f}'.format(np.std(Y, 0), (np.std(Y, 0) - std_y) / std_y))
print('mean(Y)={:2.3f} and E(Y)={:2.3}'.format(np.mean(Y, 0), np.sum(zm[:,0]*w)))
Note that Ns is the size of our Monte Carlo experiment, corresponding
to the number of times we have evaluated our simple linear model
(2). The evaluation of the model is normally the
most computationally expensive part of the analysis, and for that
reasons Ns is referred to as the cost of the analysis.
Conditional variances¶
As noted previously, the importance of a factor $Z_i$ is manifested
the existence of a shape or pattern in the model outputs
$Y$. Conversely, a uniform cloud of output points $Y$ as a function of
$Z_i$ is a symptom, albeit not a proof, indicating that $Z_i$ is a
noninfluential factor. In this section we seek to demonstrate that
conditional variances is a usefull means to quantify the shape or
pattern in the outputs.
The shape in the outputs $Y$ for a given $Z_i$, may be seen in the scatterplot as of $Y$ versus $Z_i$. In particular, we may cut the $Z_i$-axis into slices and assess how the distribution of the outputs $Y$ changes from slice to slice. This is illustrated in the code snippet below, where the slices are identified with vertical dashed lines at equidistant locations on each $Z_i$-axis, $i=1, \ldots,4$.
# # Scatter plots of data, z-slices, and linear model
fig=plt.figure()
Ndz = 10 # Number of slices of the Z-axes
Zslice = np.zeros((Nrv, Ndz)) # array for mean-values in the slices
ZBndry = np.zeros((Nrv, Ndz + 1)) # array for boundaries of the slices
dz = np.zeros(Nrv)
for k in range(Nrv):
plt.subplot(2, 2, k + 1)
zmin = np.min(Z[k, :])
zmax = np.max(Z[k, :]) # each Z[k,:] may have different extremas
dz[k] = (zmax - zmin) / Ndz
ZBndry[k, :] = np.linspace(zmin, zmax, Ndz + 1) # slice Zk into Ndz slices
Zslice[k, :] = np.linspace(zmin + dz[k] / 2., zmax - dz[k] / 2., Ndz) # Midpoint in the slice
# Plot the the vertical slices with axvline
for i in range(Ndz):
plt.axvline(ZBndry[k, i], np.amin(Y), np.amax(Y), linestyle='--', color='.75')
# Plot the data
plt.plot(Z[k, :], Y[:], '.')
xlbl = 'Z' + str(k)
plt.xlabel(xlbl)
plt.ylabel('Y')
Ymodel = w[k] * Zslice[k, :] # Produce the straight line
plt.plot(Zslice[k, :], Ymodel)
ymin = np.amin(Y); ymax = np.amax(Y)
plt.ylim([ymin, ymax])
fig.tight_layout() # adjust subplot(s) to the figure area.
Note, that average value of $Y$ in a very thin slice, corresponds to keeping $Z_i$ fixed while averaging over all output values of $Y$ due to all-but $Z_i$, which corresponds to the conditional expected value:
For convenience we let $Z_{\sim i}$ denote all-but $Z_i$. Naturally,
a measure of how much $E_{Z_{\sim i}} (Y\;|\;Z_i)$ varies in the range
of $Z_i$ is given by the conditional variance:
Further, the variance the output $Y$ may be decomposed into:
A large $\text{V}_{Z_i}(E_{Z_{\sim i}} (Y\;|\;Z_i))$ will imply that
$Z_i$ is an important factor and is therefore coined the first-order
effect of $Z_i$ on $Y$, and its fraction of the total variation of $Y$ is expressed by $S_i$, the first-order sensitivity index of $Z_i$ on $Y$:
By (17), $S_i$ is number always in the range $[0,1]$, and a high value implies an important factor.
# # Scatter plots of averaged y-values per slice, with averaged data
Zsorted = np.zeros_like(Z)
Ysorted = np.zeros_like(Z)
YsliceMean = np.zeros((Nrv, Ndz))
fig=plt.figure()
for k in range(Nrv):
plt.subplot(2, 2, k + 1)
# sort values for Zk,
sidx = np.argsort(Z[k, :]) #sidx holds the indexes for the sorted values of Zk
Zsorted[k, :] = Z[k, sidx].copy()
Ysorted[k, :] = Y[sidx].copy() # Ysorted is Y for the sorted Zk
for i in range(Ndz):
plt.axvline(ZBndry[k, i], np.amin(Y), np.amax(Y), linestyle='--', color='.75')
# find indexes of z-values in the current slice
zidx_range = np.logical_and(Zsorted[k, :] >= ZBndry[k, i], Zsorted[k, :] < ZBndry[k, i + 1])
if np.any(zidx_range): # check if range has elements
YsliceMean[k, i] = np.mean(Ysorted[k, zidx_range])
else: # set value to None if noe elements in z-slice
YsliceMean[k, i] = None
plt.plot(Zslice[k, :], YsliceMean[k, :], '.')
# # Plot linear model
Nmodel = 3
zmin = np.min(Zslice[k, :])
zmax = np.max(Zslice[k, :])
zvals = np.linspace(zmin, zmax, Nmodel)
#linear_model
Ymodel = w[k] * zvals
plt.plot(zvals, Ymodel)
xlbl = 'Z' + str(k)
plt.xlabel(xlbl)
plt.ylim(ymin, ymax)
fig.tight_layout() # adjust subplot(s) to the figure area.
SpoorMan=[np.nanvar(YsliceMean[k,:],axis=0)/np.var(Y) for k in range(4)]
print(SpoorMan)
How to compute the sensitivity indices¶
Below we will demostrate how the Sobol sensitivity indices may be computed with two approaches; the Monte Carlo method and the polynomial chaos expansion method.
Monte Carlo¶
Below some code snippets are provided to illustrate how we may compute the Soboil indices with the MCM. For the interested reader we have also writen a seperate and more detailed notebook A brief introduction to UQ and SA with the Monte Carlo method.
# calculate sens indices of non additive model
def mc_sensitivity_linear(Ns, jpdf, w, sample_method='R'):
Nrv = len(jpdf)
# 1. Generate sample matrices
A, B, C = generate_sample_matrices_mc(Ns, Nrv, jpdf, sample_method)
# 2. Evaluate the model
Y_A, Y_B, Y_C = evaluate_linear_model(A, B, C, w)
# 3. Approximate the sensitivity indices
S, ST = calculate_sensitivity_indices_mc(Y_A, Y_B, Y_C)
return A, B, C, Y_A, Y_B, Y_C, S, ST
# end calculate sens indices of non additive model
# model evaluation
def evaluate_linear_model(A, B, C, w):
number_of_parameters = A.shape[1]
number_of_sampless = A.shape[0]
# 1. evaluate sample matrices A
Y_A = linear_model(w, A)
# 2. evaluate sample matrices B
Y_B = linear_model(w, B)
# 3. evaluate sample matrices C
Y_C = np.empty((number_of_sampless, number_of_parameters))
for i in range(number_of_parameters):
z = C[i, :, :]
Y_C[:, i] = linear_model(w, z)
return Y_A, Y_B, Y_C
# Monte Carlo
# get joint distributions
jpdf = generate_distributions(zm)
Ns_mc = 1000000
# calculate sensitivity indices
A_s, B_s, C_s, f_A, f_B, f_C, S_mc, ST_mc = mc_sensitivity_linear(Ns_mc, jpdf, w)
Sensitivities=np.column_stack((S_mc,s**2))
row_labels= ['S_'+str(idx) for idx in range(1,Nrv+1)]
print("First Order Indices")
print(pd.DataFrame(Sensitivities,columns=['Smc','Sa'],index=row_labels).round(3))
Polynomial chaos expansion¶
As for the MCM some code snippets are provided to illustrate how we may compute
the Soboil indices with the polynomial chaos expansions using chaospy. A more in dept treatment of chaospy and its usage is provided in the separate notebook A practical introduction to polynomial chaos with the chaospy package.
# Polychaos computations
Ns_pc = 80
samples_pc = jpdf.sample(Ns_pc)
polynomial_order = 4
poly = cp.orth_ttr(polynomial_order, jpdf)
Y_pc = linear_model(w, samples_pc.T)
approx = cp.fit_regression(poly, samples_pc, Y_pc, rule="T")
exp_pc = cp.E(approx, jpdf)
std_pc = cp.Std(approx, jpdf)
print("Statistics polynomial chaos\n")
print('\n E(Y) | std(Y) \n')
print('pc : {:2.5f} | {:2.5f}'.format(float(exp_pc), std_pc))
S_pc = cp.Sens_m(approx, jpdf)
Sensitivities=np.column_stack((S_mc,S_pc, s**2))
print("\nFirst Order Indices")
print(pd.DataFrame(Sensitivities,columns=['Smc','Spc','Sa'],index=row_labels).round(3))
# print("\nRelative errors")
# rel_errors=np.column_stack(((S_mc - s**2)/s**2,(S_pc - s**2)/s**2))
# print(pd.DataFrame(rel_errors,columns=['Error Smc','Error Spc'],index=row_labels).round(3))
# Polychaos convergence
Npc_list = np.logspace(1, 3, 10).astype(int)
error = []
for i, Npc in enumerate(Npc_list):
Zpc = jpdf.sample(Npc)
Ypc = linear_model(w, Zpc.T)
Npol = 4
poly = cp.orth_chol(Npol, jpdf)
approx = cp.fit_regression(poly, Zpc, Ypc, rule="T")
s_pc = cp.Sens_m(approx, jpdf)
error.append(LA.norm((s_pc - s**2)/s**2))
plt.figure()
plt.semilogy(Npc_list, error)
_=plt.xlabel('Nr Z')
_=plt.ylabel('L2-norm of error in Sobol indices')
References¶
- **A. Saltelli**. *Global Sensitivity Analysis : the Primer*, John Wiley, 2008.